Abstract
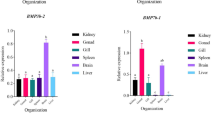
We systematically analyzed BMP gene family in H. sapiens to elucidate genetic structure, phylogenetic relationships, adaptive evolution and tissue-specific expression pattern. Total of 13 BMPs genes were identified in the H. sapiens genome. Bone morphogenetic proteins (BMPs) are composed of a variable number of exons ranging from 2 to 21. They exhibit a molecular weight ranging from 31,081.81 to 82,899.61 Da. These proteins possess hydrophilic characteristics, display thermostability, and exhibit a pH range from acidic to basic. We identified four segmental and two tandem duplication events in BMP gene family of H. sapiens. All of the vertebrate species that were studied show the presence of BMPs 1, 2, 3, 4, 5, 6, 7, 8A, and 15, however only Homo sapiens demonstrated the presence of BMP9 and BMP11. The pathway and process enrichment analysis of BMPs genes showed that these were considerably enriched in positive regulation of pathway-restricted SMAD protein phosphorylation (92%) and cartilage development (77%) biological processes. These genes exhibited positive selection signals that were shown to be conserved across vertebrate lineages. The results showed that BMP2/3/5/6/8a/15 proteins underwent adaptive selection at many amino acid locations and increased positive selection was detected in TGF-β propeptide and TGF-β super family domains which were involved in dorso-ventral patterning, limb bud development. More over the expression pattern of BMP genes revealed that BMP1 and BMP5; BMP4 and BMP6 exhibited substantially identical expression patterns in all tissues while BMP10, BMP15, and BMP3 showed tissue-specific expression.









Similar content being viewed by others
Data Availability
All the data is included in the manuscript or as a supplementary material. All figures are generated using different bioinformatics tools/software.
References
Abrams, K. L., Xu, J., Nativelle-Serpentini, C., Dabirshahsahebi, S., & Rogers, M. B. (2004). An evolutionary and molecular analysis of Bmp2 expression. Journal of Biological Chemistry, 279(16), 15916–15928. https://doi.org/10.1074/jbc.M313531200
Adiba, M., Das, T., Paul, A., Das, A., Chakraborty, S., Hosen, M. I., & Nabi, A. H. M. N. (2021). In silico characterization of coding and non-coding SNPs of the androgen receptor gene. Informatics in Medicine Unlocked, 24, 100556. https://doi.org/10.1016/j.imu.2021.100556
Ahmad, H. I., Zhou, J., Ahmad, M. J., Afzal, G., Jiang, H., Zhang, X., Elokil, A. A., Khan, M. A., Li, L., Li, H., **, L., & Chen, J. (2020). Adaptive selection in the evolution of programmed cell death-1 and its ligands in vertebrates. Aging, 12(4), 3516–3557. https://doi.org/10.18632/aging.102827
Al-Amrani, S., Al-Jabri, Z., Al-Zaabi, A., Alshekaili, J., & Al-Khabori, M. (2021). Proteomics: Concepts and applications in human medicine. World Journal of Biological Chemistry, 12(5), 57–69. https://doi.org/10.4331/wjbc.v12.i5.57
Asharani, P. V., Keupp, K., Semler, O., Wang, W., Li, Y., Thiele, H., Yigit, G., Pohl, E., Becker, J., Frommolt, P., Sonntag, C., Altmüller, J., Zimmermann, K., Greenspan, D. S., Akarsu, N. A., Netzer, C., Schönau, E., Wirth, R., Hammerschmidt, M., …, Carney, T. J. (2012). Attenuated BMP1 function compromises osteogenesis, leading to bone fragility in humans and zebrafish. The American Journal of Human Genetics, 90(4), 661–674. https://doi.org/10.1016/j.ajhg.2012.02.026
Ashkenazy, H., Abadi, S., Martz, E., Chay, O., Mayrose, I., Pupko, T., & Ben-Tal, N. (2016). ConSurf 2016: An improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Research, 44(W1), W344-350. https://doi.org/10.1093/nar/gkw408
Bailey, T. L., Johnson, J., Grant, C. E., & Noble, W. S. (2015). The MEME suite. Nucleic Acids Research, 43(W1), W39-49. https://doi.org/10.1093/nar/gkv416
Baker, F. N., & Porollo, A. (2016). CoeViz: A web-based tool for coevolution analysis of protein residues. BMC Bioinformatics, 17(1), 119. https://doi.org/10.1186/s12859-016-0975-z
Baker, F. N., & Porollo, A. (2018). CoeViz: A web-based integrative platform for interactive visualization of large similarity and distance matrices. Data, 3(1), 4.
Bal, Z., & Kushioka, J. (2020). BMP and TGFβ use and release in bone regeneration. Turkish Journal of Medical Sciences, 50(2), 1707–1722. https://doi.org/10.3906/sag-2003-127
Canty-Laird, E., Carré, G.-A., Mandon-Pépin, B., Kadler, K. E., & Fabre, S. (2010). First evidence of bone morphogenetic protein 1 expression and activity in sheep ovarian follicles1. Biology of Reproduction, 83(1), 138–146. https://doi.org/10.1095/biolreprod.109.082115
Cao, X., & Chen, D. (2005). The BMP signaling and in vivo bone formation. Gene, 357(1), 1–8. https://doi.org/10.1016/j.gene.2005.06.017
Cecchi, S., Bennet, S. J., & Arora, M. (2016). Bone morphogenetic protein-7: Review of signalling and efficacy in fracture healing. Journal of Orthopaedic Translation, 4, 28–34. https://doi.org/10.1016/j.jot.2015.08.001
Chatzou, M., Magis, C., Chang, J.-M., Kemena, C., Bussotti, G., Erb, I., & Notredame, C. (2015). Multiple sequence alignment modeling: Methods and applications. Briefings in Bioinformatics, 17(6), 1009–1023. https://doi.org/10.1093/bib/bbv099
Chen, C., Chen, H., Zhang, Y., Thomas, H. R., Frank, M. H., He, Y., & **a, R. (2020). TBtools: An integrative toolkit developed for interactive analyses of big biological data. Molecular Plant, 13(8), 1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chen, G., Xu, H., Yao, Y., Xu, T., Yuan, M., Zhang, X., Lv, Z., & Wu, M. (2020). BMP signaling in the development and regeneration of cranium bones and maintenance of calvarial stem cells. Frontiers in Cell and Developmental Biology, 8, 135. https://doi.org/10.3389/fcell.2020.00135
Corcoran, D., Maltbie, N., Sudalairaj, S., Baker, F. N., Hirschfeld, J., & Porollo, A. (2021). CoeViz 2: Protein graphs derived from amino acid covariance. Frontiers in Bioinformatics. https://doi.org/10.3389/fbinf.2021.653681
Cui, G., Feng, S., Yan, Y., Wang, L., He, X., Li, X., Duan, Y., Chen, J., Tang, K., Zheng, P., Tam, P. P. L., Si, W., **g, N., & Peng, G. (2022). Spatial and molecular anatomy of germ layers in the gastrulating Cynomolgus monkey embryo. bioRxiv, 2022.2001.2026.474719. https://doi.org/10.1101/2022.01.26.474719
Dathe, K., Kjaer, K. W., Brehm, A., Meinecke, P., Nürnberg, P., Neto, J. C., Brunoni, D., Tommerup, N., Ott, C. E., Klopocki, E., Seemann, P., & Mundlos, S. (2009). Duplications involving a conserved regulatory element downstream of BMP2 are associated with brachydactyly type A2. The American Journal of Human Genetics, 84(4), 483–492. https://doi.org/10.1016/j.ajhg.2009.03.001
Del Amparo, R., Branco, C., Arenas, J., Vicens, A., & Arenas, M. (2021). Analysis of selection in protein-coding sequences accounting for common biases. Briefings in Bioinformatics. https://doi.org/10.1093/bib/bbaa431
Delolme, F., Anastasi, C., Alcaraz, L. B., Mendoza, V., Vadon-Le Goff, S., Talantikite, M., Capomaccio, R., Mevaere, J., Fortin, L., Mazzocut, D., Damour, O., Zanella-Cléon, I., Hulmes, D. J. S., Overall, C. M., Valcourt, U., Lopez-Casillas, F., & Moali, C. (2015). Proteolytic control of TGF-β co-receptor activity by BMP-1/tolloid-like proteases revealed by quantitative iTRAQ proteomics. Cellular and Molecular Life Sciences, 72(5), 1009–1027. https://doi.org/10.1007/s00018-014-1733-x
Delport, W., Poon, A. F. Y., Frost, S. D. W., & Kosakovsky Pond, S. L. (2010). Datamonkey 2010: A suite of phylogenetic analysis tools for evolutionary biology. Bioinformatics, 26(19), 2455–2457. https://doi.org/10.1093/bioinformatics/btq429
Deng, Z. H., Li, Y. S., Gao, X., Lei, G. H., & Huard, J. (2018). Bone morphogenetic proteins for articular cartilage regeneration. Osteoarthritis and Cartilage, 26(9), 1153–1161. https://doi.org/10.1016/j.joca.2018.03.007
Ducy, P., & Karsenty, G. (2000). The family of bone morphogenetic proteins. Kidney International, 57(6), 2207–2214. https://doi.org/10.1046/j.1523-1755.2000.00081.x
Ehata, S., & Miyazono, K. (2022). Bone morphogenetic protein signaling in cancer; some topics in the recent 10 years. Frontiers in Cell and Developmental Biology, 10, 883523. https://doi.org/10.3389/fcell.2022.883523
Gasteiger, E., Gattiker, A., Hoogland, C., Ivanyi, I., Appel, R. D., & Bairoch, A. (2003). ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Research, 31(13), 3784–3788. https://doi.org/10.1093/nar/gkg563
Gasteiger, E., Hoogland, C., Gattiker, A., Duvaud, S. E., Wilkins, M. R., Appel, R. D., & Bairoch, A. (2005). Protein identification and analysis tools on the ExPASy server. In J. M. Walker (Ed.), The proteomics protocols handbook (pp. 571–607). Humana Press.
Ge, G., & Greenspan, D. S. (2006). BMP1 controls TGFbeta1 activation via cleavage of latent TGFbeta-binding protein. Journal of Cell Biology, 175(1), 111–120. https://doi.org/10.1083/jcb.200606058
Gipson, G. R., Goebel, E. J., Hart, K. N., Kappes, E. C., Kattamuri, C., McCoy, J. C., & Thompson, T. B. (2020). Structural perspective of BMP ligands and signaling. Bone, 140, 115549. https://doi.org/10.1016/j.bone.2020.115549
Graham, S. J. L., Wicher, K. B., Jedrusik, A., Guo, G., Herath, W., Robson, P., & Zernicka-Goetz, M. (2014). BMP signalling regulates the pre-implantation development of extra-embryonic cell lineages in the mouse embryo. Nature Communications, 5(1), 5667. https://doi.org/10.1038/ncomms6667
Guha, U., Gomes, W. A., Kobayashi, T., Pestell, R. G., & Kessler, J. A. (2002). In vivo evidence that BMP signaling is necessary for apoptosis in the mouse limb. Developmental Biology, 249(1), 108–120. https://doi.org/10.1006/dbio.2002.0752
Guo, A. Y., Zhu, Q. H., Chen, X., & Luo, J. C. (2007). GSDS: A gene structure display server. Yi Chuan, 29(8), 1023–1026.
Hall, B. G. (2013). Building phylogenetic trees from molecular data with MEGA. Molecular Biology and Evolution, 30(5), 1229–1235. https://doi.org/10.1093/molbev/mst012
Hoffmann, A., & Gross, G. (2001). BMP signaling pathways in cartilage and bone formation. Critical Reviews in Eukaryotic Gene Expression, 11(1–3), 23–45.
Hopkins, D. R., Keles, S., & Greenspan, D. S. (2007). The bone morphogenetic protein 1/Tolloid-like metalloproteinases. Matrix Biology, 26(7), 508–523. https://doi.org/10.1016/j.matbio.2007.05.004
Hu, B., **, J., Guo, A. Y., Zhang, H., Luo, J., & Gao, G. (2015). GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics, 31(8), 1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Huang, Y., & Umulis, D. M. (2019). Scale invariance of BMP signaling gradients in zebrafish. Scientific Reports, 9(1), 5440. https://doi.org/10.1038/s41598-019-41840-8
Jha, K., & Saha, S. (2020). Amalgamation of 3D structure and sequence information for protein–protein interaction prediction. Scientific Reports, 10(1), 19171. https://doi.org/10.1038/s41598-020-75467-x
John, M., Kim, K.-J., Bae, S. D. W., Qiao, L., & George, J. (2019). Role of BMP-9 in human liver disease. Gut, 68(11), 2097. https://doi.org/10.1136/gutjnl-2018-317543
Katagiri, T., & Watabe, T. (2016). Bone morphogenetic proteins. Cold Spring Harbor Perspectives in Biology, 8(6), a021899. https://doi.org/10.1101/cshperspect.a021899
Kokabu, S., & Rosen, V. (2018). BMP3 expression by osteoblast lineage cells is regulated by canonical Wnt signaling. FEBS Open Bio, 8(2), 168–176. https://doi.org/10.1002/2211-5463.12347
Lademann, F., Hofbauer, L. C., & Rauner, M. (2020). The bone morphogenetic protein pathway: The osteoclastic perspective. Frontiers in Cell and Developmental Biology, 8, 586031. https://doi.org/10.3389/fcell.2020.586031
Lei, L., Zhu, J., Chen, C., Wang, Y., Wu, C., Qi, M., Wang, Y., Liu, X., Hong, X., Yu, L., Chen, H., Wei, C., Liu, Y., Li, W., & Zhu, X. (2023). Genome-wide identification, evolution and expression analysis of bone morphogenetic protein (BMP) gene family in Chinese soft-shell turtle (Pelodiscus sinensis). Frontires in Genetics. https://doi.org/10.3389/fgene.2023.1109478
Letunic, I., & Bork, P. (2021). Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Research, 49(W1), W293-w296. https://doi.org/10.1093/nar/gkab301
Li, Y., Lv, X., Wang, S., Cao, X., Yuan, Z., Getachew, T., Mwacharo, J. M., Haile, A., & Sun, W. (2022). BMP7 functions to regulate proliferation of dermal papilla cells in Hu sheep. Genes, 13(2), 201.
Liu, M., Goldman, G., MacDougall, M., & Chen, S. (2022). BMP signaling pathway in dentin development and diseases. Cells. https://doi.org/10.3390/cells11142216
Liu, F., Hata, A., Baker, J. C., Doody, J., Cárcamo, J., Harland, R. M., & Massagué, J. (1996). A human Mad protein acting as a BMP-regulated transcriptional activator. Nature, 381(6583), 620–623. https://doi.org/10.1038/381620a0
Liu, X., Liu, F., Xu, H., Yang, Y., Wang, Y., Hong, X., Li, W., Yu, L., Chen, C., Xu, H., & Zhu, X. (2022). Characterization of the in vitro cultured ovarian cells in the Asian Yellow Pond Turtle (Mauremys mutica). Biology (Basel), 11(10), 1404. https://doi.org/10.3390/biology11101404
Lochab, A. K., & Extavour, C. G. (2017). Bone morphogenetic protein (BMP) signaling in animal reproductive system development and function. Developmental Biology, 427(2), 258–269. https://doi.org/10.1016/j.ydbio.2017.03.002
Ma, Y., Liu, Y., & Cheng, J. (2018). Protein secondary structure prediction based on data partition and semi-random subspace method. Scientific Reports, 8(1), 9856. https://doi.org/10.1038/s41598-018-28084-8
Ma, Y., **ao, Y., **ao, Z., Wu, Y., Zhao, H., Gao, G., Wu, L., Wang, T., Zhao, N., & Li, J. (2022). Genome-wide identification, characterization and expression analysis of the BMP family associated with beak-like teeth in Oplegnathus. Front Genet, 13, 938473. https://doi.org/10.3389/fgene.2022.938473
Madende, M., & Osthoff, G. (2019). Comparative genomics of casein genes. Journal of Dairy Research, 86(3), 323–330. https://doi.org/10.1017/s0022029919000414
Manzari-Tavakoli, A., Babajani, A., Farjoo, M. H., Ha**asrollah, M., Bahrami, S., & Niknejad, H. (2022). The cross-talks among bone morphogenetic protein (BMP) signaling and other prominent pathways involved in neural differentiation. Frontiers in Molecular Neuroscience, 15, 827275. https://doi.org/10.3389/fnmol.2022.827275
Mao, L., Yano, M., Kawao, N., Tamura, Y., Okada, K., & Kaji, H. (2013). Role of matrix metalloproteinase-10 in the BMP-2 inducing osteoblastic differentiation. Endocrine Journal, 60(12), 1309–1319. https://doi.org/10.1507/endocrj.ej13-0270
Marchler-Bauer, A., Derbyshire, M. K., Gonzales, N. R., Lu, S., Chitsaz, F., Geer, L. Y., Geer, R. C., He, J., Gwadz, M., Hurwitz, D. I., Lanczycki, C. J., Lu, F., Marchler, G. H., Song, J. S., Thanki, N., Wang, Z., Yamashita, R. A., Zhang, D., Zheng, C., & Bryant, S. H. (2015). CDD: NCBI’s conserved domain database. Nucleic Acids Research, 43(Database issue), D222–D226. https://doi.org/10.1093/nar/gku1221
Morgan, C. C., & Hart, M. W. (2019). Molecular evolution of mammalian genes with epistatic interactions in fertilization. BMC Evolutionary Biology, 19(1), 154. https://doi.org/10.1186/s12862-019-1480-6
Asharani, P. V., Keupp, K., Semler, O., Wang, W., Li, Y., Thiele, H., Yigit, G., Pohl, E., Becker, J., Frommolt, P., Sonntag, C., Altmüller, J., Zimmermann, K., Greenspan, D. S., Akarsu, N. A., Netzer, C., Schönau, E., Wirth, R., Hammerschmidt, M., …, Carney, T. J. (2012). Attenuated BMP1 function compromises osteogenesis, leading to bone fragility in humans and zebrafish. The American Journal of Human Genetics, 90, 661–674. doi: https://doi.org/10.1016/j.ajhg.2012.02.026
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T., & Salzberg, S. L. (2016). Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nature Protocols, 11(9), 1650–1667. https://doi.org/10.1038/nprot.2016.095
Pham, H. G., Mukherjee, S., Choi, M. J., & Yun, J. W. (2021). BMP11 regulates thermogenesis in white and brown adipocytes. Cell Biochemistry and Function, 39(4), 496–510. https://doi.org/10.1002/cbf.3615
Qu, X., Liu, Y., & Cao, D. (2019). BMP10 preserves cardiac function through its dual activation of SMAD-mediated and STAT3-mediated pathways. Journal of Biological Chemistry, 294(52), 19877–19888. https://doi.org/10.1074/jbc.RA119.010943
Sangsin, A., Kuptanon, C., Srichomthong, C., Pongpanich, M., Suphapeetiporn, K., & Shotelersuk, V. (2017). Two novel compound heterozygous BMP1 mutations in a patient with osteogenesis imperfecta: A case report. BMC Medical Genetics, 18(1), 25. https://doi.org/10.1186/s12881-017-0384-9
Shah, T. A., Zhu, Y., Shaikh, N. N., Harris, M. A., Harris, S. E., & Rogers, M. B. (2017). Characterization of new bone morphogenetic protein (Bmp)-2 regulatory alleles. Genesis, 55(7), e23035. https://doi.org/10.1002/dvg.23035
Smith, M. D., Wertheim, J. O., Weaver, S., Murrell, B., Scheffler, K., & Kosakovsky Pond, S. L. (2015). Less is more: An adaptive branch-site random effects model for efficient detection of episodic diversifying selection. Molecular Biology and Evolution, 32(5), 1342–1353. https://doi.org/10.1093/molbev/msv022
Su, Z., He, L., Shang, H., Dai, T., Xu, F., & Zhao, J. (2020). Overexpression of bone morphogenetic protein-1 promotes osteogenesis of bone marrow mesenchymal stem cells in vitro. Medical Science Monitor, 26, e920122. https://doi.org/10.12659/msm.920122
Szklarczyk, D., Kirsch, R., Koutrouli, M., Nastou, K., Mehryary, F., Hachilif, R., Gable, A. L., Fang, T., Doncheva, N. T., Pyysalo, S., Bork, P., Jensen, L. J., & von Mering, C. (2022). The STRING database in 2023: Protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Research, 51(D1), D638–D646. https://doi.org/10.1093/nar/gkac1000
Tamura, K., Stecher, G., & Kumar, S. (2021). MEGA11: Molecular evolutionary genetics analysis version 11. Molecular Biology and Evolution, 38(7), 3022–3027. https://doi.org/10.1093/molbev/msab120
Thakur, V., & Kumar, P. (2018). Analysis of hepatitis E virus (HEV) X-domain structural model. Bioinformation, 14, 398–403. https://doi.org/10.6026/97320630014398
von Bubnoff, A., Peiffer, D. A., Blitz, I. L., Hayata, T., Ogata, S., Zeng, Q., Trunnell, M., & Cho, K. W. Y. (2005). Phylogenetic footprinting and genome scanning identify vertebrate BMP response elements and new target genes. Developmental Biology, 281(2), 210–226. https://doi.org/10.1016/j.ydbio.2005.02.014
Wang, R. N., Green, J., Wang, Z., Deng, Y., Qiao, M., Peabody, M., Zhang, Q., Ye, J., Yan, Z., Denduluri, S., Idowu, O., Li, M., Shen, C., Hu, A., Haydon, R. C., Kang, R., Mok, J., Lee, M. J., Luu, H. L., & Shi, L. L. (2014). Bone morphogenetic protein (BMP) signaling in development and human diseases. Genes & Diseases, 1(1), 87–105. https://doi.org/10.1016/j.gendis.2014.07.005
Weaver, S., Shank, S., Spielman, S., Li, M., Muse, S., & Pond, S. (2018). Datamonkey 2.0: A modern web application for characterizing selective and other evolutionary processes. Molecular Biology and Evolution, 35, 773–777. https://doi.org/10.1093/molbev/msx335
Wei, Z., Salmon, R. M., Upton, P. D., Morrell, N. W., & Li, W. (2014). Regulation of bone morphogenetic protein 9 (BMP9) by redox-dependent proteolysis. Journal of Biological Chemistry, 289(45), 31150–31159. https://doi.org/10.1074/jbc.M114.579771
Wu, X., Gong, Q., Chen, Y., Liu, Y., Song, M., Li, F., Li, P., & Lai, J. (2022). Full-length transcriptome and analysis of bmp-related genes in Platypharodon extremus. Heliyon, 8(10), e10783. https://doi.org/10.1016/j.heliyon.2022.e10783
Xu, J., & Rogers, M. B. (2007). Modulation of bone morphogenetic protein (BMP) 2 gene expression by Sp1 transcription factors. Gene, 392(1–2), 221–229. https://doi.org/10.1016/j.gene.2006.12.032
Xu, S., Zhang, S., Zhang, W., Liu, H., Wang, M., Zhong, L., Bian, W., & Chen, X. (2022). Genome-wide identification, phylogeny, and expression profile of the Dmrt (doublesex and Mab-3 related transcription factor) gene family in channel catfish (Ictalurus punctatus). Frontiers in Genetics, 13, 891204. https://doi.org/10.3389/fgene.2022.891204
Xu, X., Lv, F., Song, Y., Li, L., Asan, Wei, X., Zhao, X., Jiang, Y., Wang, O., **ng, X., **a, W., & Li, M. (2019). Novel mutations in BMP1 induce a rare type of osteogenesis imperfecta. Clinica Chimica Acta, 489, 21–28. https://doi.org/10.1016/j.cca.2018.11.004
Yan, Y., & Wang, Q. (2021). BMP signaling: Lighting up the way for embryonic dorsoventral patterning. Frontiers in Cell and Developmental Biology, 9, 799772. https://doi.org/10.3389/fcell.2021.799772
Yang, D., Yang, X., Dai, F., Wang, Y., Yang, Y., Hu, M., & Cheng, Y. (2021). The role of bone morphogenetic protein 4 in ovarian function and diseases. Reproductive Sciences, 28(12), 3316–3330. https://doi.org/10.1007/s43032-021-00600-8
You, J., Wang, W., Chang, H.-M., Yi, Y., Zhao, H., Zhu, H., Sun, Y., Tang, M., Wang, C., Sang, Y., Feng, G., Cheng, S., Leung, P. C. K., & Zhu, Y.-M. (2021). The BMP2 signaling axis promotes invasive differentiation of human trophoblasts. Frontiers in Cell and Developmental Biology. https://doi.org/10.3389/fcell.2021.607332
Yu, H., Wang, Y., Wang, M., Liu, Y., Cheng, J., & Zhang, Q. (2020). Growth differentiation factor 9 (gdf9) and bone morphogenetic protein 15 (bmp15) are potential intraovarian regulators of steroidogenesis in Japanese flounder (Paralichthys olivaceus). General and Comparative Endocrinology, 297, 113547. https://doi.org/10.1016/j.ygcen.2020.113547
Zhang, C., Wang, W., & Wang, D. (2022). Genome-wide identification and characterization of the WRKY gene family in Scutellaria baicalensis Georgi under diverse abiotic stress. International Journal of Molecular Sciences, 23(8), 4225. https://doi.org/10.3390/ijms23084225
Zhang, C., Xu, X., Xu, X., Li, Y., Zhao, P., Chen, X., Shen, X., Zhang, Z., Chen, Y., Liu, S., Xu, X., Yuling, L., Lai, Z., & Lai, Z. (2022). Genome-wide identification, evolution analysis of cytochrome P450 monooxygenase multigene family and their expression patterns during the early somatic embryogenesis in Dimocarpus longan Lour. Gene, 826, 146453. https://doi.org/10.1016/j.gene.2022.146453
Zhang, H., Zhang, W., Bai, G., Gao, L., & Li, K. (2021). Bone morphogenetic protein-7 (BMP-7) promotes neuronal differentiation of bone marrow mesenchymal stem cells (BMSCs) in vitro. BioMed Research International, 2021, 7239783. https://doi.org/10.1155/2021/7239783
Acknowledgements
None.
Author information
Authors and Affiliations
Contributions
ZR: Conceptualization, Data Analysis, original draft writing; MH: Investigation, original draft writing; SP: Investigation, Supervision, Review and editing; MS: Investigation, original draft writing; SS: Original draft writing, Review and editing; UI: Original draft writing, ZF: Original draft writing, MT: Original draft writing; Investigation.
Corresponding author
Ethics declarations
Conflict of interest
Authors have declared no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Riaz, Z., Hussain, M., Parveen, S. et al. In Silico Analysis: Genome-Wide Identification, Characterization and Evolutionary Adaptations of Bone Morphogenetic Protein (BMP) Gene Family in Homo sapiens. Mol Biotechnol (2023). https://doi.org/10.1007/s12033-023-00944-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-023-00944-3