Abstract
Objective
Ovarian cancer is the seventh most common cancer in women, and it causes many deaths in women worldwide. Patients with ovarian cancer have a poor prognosis and low survival rate. This study aimed to explore the role of the SRC-3/TRAF4/PI3K/AKT pathway in ovarian cancer development.
Methods
SRC-3 and TRAF4 expression in ovarian cancer cell lines were assessed using qRT-PCR and western-blotting. The expression of SRC-3 and TRAF4 in ovarian cancer cells was downregulated by transient transfection with sh-RNAs. An MTT assay was performed to evaluate cell proliferation. Cell migration and invasion were measured using a Transwell assay. Cell stemness was detected using a cell spheroidization assay and western blotting. The expression levels of stem cell factors and PI3K/AKT pathway proteins were determined by qRT-PCR and western blot analysis.
Results
SRC-3 and TRAF4 were upregulated in ovarian cancer cell lines. TRAF4 is a downstream factor of SRC-3, and the protein level of TRAF4 was regulated by SRC-3. SRC-3 knockdown reduced TRAF4 expression. Silencing SRC-3 or TRAF4 inhibited cell proliferation, migration, and invasion, as well as the expression of stem cell factors. Furthermore, sh-TRAF4 as well as treatment with LY294002, the PI3K/Akt inhibitor, inhibited the phosphorylation of Akt and PI3K, thus repressing the activation of PI3K/AKT signaling pathway in ovarian cancer cell lines. However, TRAF4 overexpression reversed the effect of SRC-3 silencing on cell proliferation, migration, invasion, and stemness.
Conclusion
Our study demonstrated that SRC-3/TRAF4 promotes ovarian cancer cell growth, migration, invasion, and stemness by activating the PI3K/AKT pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ovarian cancer is the seventh most common cancer and is one the leading cause of malignancy-related mortality in women [1]. Most ovarian cancers are derived from serous tubal intraepithelial carcinoma, while others develop from serous endometrial intraepithelial carcinoma [2]. Owing to the lack of obvious symptoms and effective detection methods, 70 percent of ovarian cancers are diagnosed at stage III or IV with extensive metastases in the abdominal cavity [3, 4]. Debulking surgery and chemotherapy are regarded as the most common treatment options for ovarian cancer [2, 5]. According to ovarian cancer surgery guidelines (ESGO 2017), surgical resection of macroscopic tumors is the first choice for patients in their early stages [6]. The chemotherapeutic modality, platinum/taxane treatment, for instance, would be administered to the patients [7]. However, for patients with advanced stages of tumors (III and IV), neoadjuvant chemotherapy is administered before surgery. Bevacizumab and paclitaxel are the first-line treatment options [7]. However, the risk of recurrence for advanced ovarian cancer is very high (~ 70%) [2, 8,9,10]. Chemotherapy is the principal treatment for recurrent ovarian cancer, and the combination of platinum and other drugs will be useful [7, 11]. However, once ovarian cancer relapses, the tumors develop rapidly with therapy resistance to some extent [2]. It causes many deaths in women worldwide, and patients with ovarian cancer have a poor prognosis and low survival rate [12].
Previous studies have shown that ovarian cancer stem cell transformation could be one of the reasons for ovarian cancer development [13]. Cancer stem cells then give rise to chemoresistant recurrent tumors at the metastatic sites. It has been reported that ovarian cancer stem cells are related to survival after conventional chemotherapy [7]. Numerous studies have been conducted to elucidate the biological mechanisms that regulate ovarian cancer stemness. In regard to ovarian cancer, various specific markers of stemness, such as Nanog, SOX2, and OCT4, have been used to isolate and characterize ovarian cancer stem cells. Moreover, these stem cell factors have been found to regulate ovarian cancer development [14,15,16]. PBX1, a stem cell reprogramming factor, mediates chemotherapy resistance in ovarian carcinomas [17]. The Wnt signaling pathway is involved in ovarian cancer development by regulating the balance between stemness and differentiation of ovarian cancer stem cells [18]. However, the underlying mechanisms of cancer stem cells in ovarian cancer development remain unclear, and a deeper understanding of cancer cell migration, invasion, and their interaction with stem cell factors is needed to facilitate studies on ovarian cancer prevention and the development of novel therapeutic approaches.
Steroid receptor coactivator 3 (SRC-3) is an oncogenic nuclear receptor coactivator belonging to the p160 family of coactivators [19]. Numerous studies have shown that SRC-3 is overexpressed in ovarian carcinomas and other human malignant diseases [19,20,21]. SRC-3 has been regarded as a marker of aggressive diseases because it plays an important role in drug resistance and activation of multiple pathways, such as E2F and AKT signaling [21,22,23]. In addition, SRC-3 directly inhibits the function of the tumor suppressor p53 to exert its oncogenic role [24]. Increased SRC-3 expression in ovarian cancer predicts chemoresistance and poor prognosis [21]. SRC-3 interacts with nuclear receptors such as estrogen receptor (ER) to enhance target gene transcription and promote cancer cell proliferation [19, 25]. It has been reported that SRC-3 can control metastasis of ovarian cancer cells [19]. Downregulation of SRC-3 expression in ovarian cancer cells significantly inhibits their migration and spread [21]. However, the mechanism by which SRC-3 is involved in ovarian cancer development remains unclear.
TRAF4 [Tumor necrosis factor (TNF) receptor-associated factor 4], a member of the TRAF protein family, is a critical downstream player of SRC-3 [24, 26]. TRAF4 is expressed at basal levels in normal tissues and cells [26]. However, it is abundantly expressed in various human malignancies [27]. Although it shares a common TRAF domain at the C-terminus with other TRAF family members, unlike other TRAF family members, TRAF4 does not interact with TNF and interleukin receptors [26]. High TRAF4 expression has been detected in different cancers, such as hepatocellular carcinoma, endometrial cancer, and breast cancer [28,29,30]. Emerging evidence indicates that TRAF4 plays an essential role as an oncogene in cancer development [31]. TRAF4 modulates cell proliferation, apoptosis, migration, and invasion in breast cancer and other cancer types [32, 33]. Previous studies have shown that breast cancer patients with high TRAF4 expression have poor prognosis [34]. Furthermore, overexpression of TRAF4 eliminates the inhibition of cancer cell migration via DR6 knockdown [35]. Liu et al. demonstrated that TRAF4 facilitates HCC cell migration and invasion by activating the PI3K/Akt signaling pathway [28]. Furthermore, it has been reported that Oct4 is a downstream gene of the PI3K/AKT pathway. TRAF4 increased the expression of Oct4 in endometrial cancer cells. TRAF4 promotes cell proliferation and migration by activating the PI3K/AKT/Oct4 pathway, which regulates the progression of endometrial cancer [29].
As a downstream factor of SRC-3, the protein level of TRAF4 is regulated by SRC-3 in cells and breast tumors. With the help of AP-1 transcription factor, SRC-3 directly regulates TRAF4 transcription [24]. The expression of TRAF4 is positively correlated with SRC-3 expression in breast tumors [24]. Breast cancer cells with high levels of TRAF4 are more resistant to nitric oxide-induced cell death, similar to cells with high levels of SRC-3 [36]. A previous study reported that TRAF4 was upregulated in endometrial cancer tissues [29]. TRAF4 promotes the development of endometrial cancer by activating the PI3K/AKT pathway and enhancing the phosphorylation of AKT and PI3K [29]. However, it is unclear whether SRC-3 employs a similar TRAF4/PI3K/AKT pathway to facilitate ovarian cancer development.
In this study, we tested the growth, migration, invasion, and stemness of ovarian cancer cells by knocking down expression of SRC-3 or TRAF4. In addition, we verified that the specific binding of SRC-3 and TRAF4 in ovarian cancer cells promoted the development of ovarian cancer by activating the PI3K/AKT pathway. A better understanding of the molecular mechanisms of the SRC-3/TRAF4 axis in the development of ovarian cancer will improve our understanding of carcinogenesis and facilitate additional therapeutic avenues for targeting ovarian cancers.
Materials and methods
Plasmid constructs
cDNAs encoding the full-length TRAF4 was amplified from ovarian cancer cell via reverse transcription-PCR and cloned into pcDNA3.1 (Invitrogen). Short-hairpin RNA (shRNA) targeting SRC-3 and TRAF4 were constructed by annealing two pairs of primers. The sense stranded oligos and their respective anti-sense stranded oligos were annealed and cloned into pLKO.1 vector (Invitrogen). A negative control was constructed by the same method with another pair of primers. All constructs were verified by DNA sequencing.
Cell cultures and transfection
The Anglne cells were obtained from Procell (Wuhan, China). The Other human ovarian cancer cell lines and normal ovarian epithelial cells were obtained from American Type Culture Collection (ATCC, USA) and were maintained in Dulbecco's modified Eagle’s medium (DMEM; Gibco, USA) containing 10% fetal bovine serum (FBS) (Gibco, USA) and 1% penicillin–streptomycin solution (Gibco, USA) at 37 °C.
CAOV-3 and SKOV3 cells were seeded at a density of 1 × 106 cells/well in 6-well plates and transfected with sh-SRC-3 and sh-TRAF4 alone or together with pcTRAF4 using Lipofectamine 2000 reagent (Invitrogen) in serum-free OPTI-MEM (Invitrogen), according to the manufacturer's instructions. CAOV-3 and SKOV3 cells were incubated in a fresh medium containing 20 μM LY294002. The cells were harvested at specific time points, and protein expression was analyzed.
MTT assay
Cell proliferation was determined using the 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) colorimetric assay (Sigma Aldrich) according to the manufacturer's instructions. Cells transfected with sh-SRC-3 or sh-TRAF4 alone, or together with pc-TRAF4, were seeded at a density of 1 × 104 cells/well in 96-well plates. After 1, 2, and 3 d, the cells were washed twice with PBS. MTT (5 mg/mL) was added into the cells and incubated with the cells at 37 °C. After 3 h, 100 μL of DMSO was added to dissolve the formazan crystals. The density of each well was measured at 570 nm wavelength using a microplate reader (Bio-Rad Laboratories, USA).
Transwell assay for migration
Transwell migration assays were performed using 8.0-μm pore inserts (BD Biosciences, USA). Cells (5 × 104) were suspended in 200 μL of DMEM and loaded into the upper wells, and 600 μL of DMEM with 5% FBS was added to the lower chambers. Cells were incubated at 37 °C for 2 d. The cells were then fixed with 4% paraformaldehyde and stained with 0.1% crystal violet solution for 4 h. The number of migratory cells was determined by counting five random areas of constant size per well.
Transwell assay for invasion
Transwell invasion analysis was performed using 100 μL Matrigel (1:5)-coated inserts (BD Biosciences, USA). Then, 5 × 104 cells were added to each insert. DMEM containing 5% FBS was added to the lower chamber. The cells and medium were incubated at 37 °C for 2 d. After incubation, the medium was removed and the cells were washed twice with PBS. The cells were then fixed with 4% paraformaldehyde and stained with 0.1% crystal violet solution for 4 h, and the number of invading cells was confirmed by counting five random areas of constant size per well.
qRT-PCR analysis
Total RNA was extracted from cells using TRIzol reagent (Invitrogen, USA), according to the manufacturer's instructions. cDNA was obtained by reverse transcription using a reverse transcription kit (M1701; Promega, USA). The expression of SRC-3, TRAF4, CD44, CD133, SOX2, OCT3/4, and NANOG was detected by qRT-PCR using the SYBR Green Master Mix (Takara). GAPDH mRNA was used as an internal control and relative expression changes were calculated using the 2−ΔΔCt method.
Western blot
The proteins used for western blotting were extracted from the tissues and cells using RIPA lysis buffer. Five microliters of the lysates were removed for protein concentration measurement (Bio-Rad Protein Assay Dye Reagent Concentrate). Total protein (30 μg) was subjected to sodium dodecyl sulfate–polyacrylamide gel electrophoresis. Proteins were then transferred to nitrocellulose membrane filters (Millipore, USA). After blocking with 5% milk in PBST at room temperature for 30 min, the membranes were incubated with primary antibodies at room temperature for 60 min. The primary antibodies used in this study were as follows: anti-SRC-3 (mouse mAb #2115, 1:1000), anti-CD44 (mouse mAb #3570, 1:1000), anti-CD133 (rabbit mAb #86781, 1:1000), anti-SOX2 (mouse mAb #4900, 1:1000), anti-NANOG(mouse mAb #4893, 1:1000), anti-TRAF4 (rabbit mAb #18527, 1:1000), anti-p-AKT (rabbit mAb #4058, 1:1000), anti-AKT (rabbit mAb #75692, 1:1000), anti-p-PI3K (rabbit mAb #17366, 1:1000), and anti-PI3K (antibody #4255, 1:1000) were purchased from Cell Signaling Technology; anti-GAPDH (mouse mAb, AC002, 1:5000) and anti-OCT3/4 (rabbit pAb, A7920, 1:1000) were purchased from ABclonal. After washing three times with PBST buffer, the membranes were incubated with HRP-conjugated secondary antibodies (Sigma-Aldrich) for 1 h at room temperature. Images were visualized using a Licor Odyssey imager (LiCor Inc., Lincoln, NE, USA). The relative density of the bands was analyzed and quantified using Image Quant software from the Licor Odyssey imager.
Colony formation assay
About 200 cells were inoculated in a low-adhesive 6-well plate containing stem cell culture medium (DMEM/F12 + 20 ng/mL EGF, 10 ng/mL βFGF, and 10 μL/mL B27) (Gibco, USA) and incubated at 37 °C without serum. After two weeks, an inverted microscope was used to observe clones with diameters greater than 50 μm.
Statistical analysis
All assays were repeated at least three times to ensure accuracy. The results of multiple experiments are presented as mean ± SD. Statistical analyses were performed using the GraphPad software (version 5.0; GraphPad Software, USA). P-values were calculated using one-way analysis of variance (ANOVA). A P-value of < 0.05 was considered statistically significant.
Results
SRC-3 and TRAF4 were upregulated in ovarian cancer cells
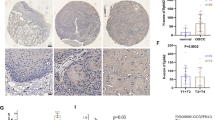
We assessed the mRNA levels of SRC-3 and TRAF4 in multiple ovarian cancer cell lines using qRT-PCR. As shown in Fig. 1A, the mRNA levels of SRC-3 and TRAF4 were increased in ovarian cancer cell lines, such as Anglne, CAOV-3, IGROV1, SW626, and SKOV3, compared to normal ovarian epithelial cells, IOSE80, with the mRNA levels of SRC3 and TRAF4 in CAOV3 and SKOV3 cells showing the highest change (Fig. 1A). In addition, SRC3 and TRAF4 protein expression was detected in IOSE80, CAOV3, and SKOV3 cells. Western blotting results showed that the protein expression of SRC3 and TRAF4 was significantly higher in CAOV3 and SKOV3 cells than in IOSE80 cells (Fig. 1B). Because SRC-3 and TRAF4 were more highly expressed in CAOV-3 and SKOV3 cells than in other ovarian cancer cell lines, we speculated that these two cell lines might be most significantly regulated by the expression of SRC3 and TRAF4. Therefore, these two cell lines were selected for subsequent experiments. Taken together, these results strongly support that SRC-3 and TRAF4 are upregulated in ovarian cancer cells.
SRC-3 and TRAF4 were upregulated in ovarian cancer cells. A The mRNA level of SRC-3 and TRAF4 were detected by qRT-PCR in ovarian cancer cells and normal ovarian epithelial cells. B The protein levels of SRC-3 and TRAF4 were detected by western blot in CAOV-3 and SKOV3 cells and normal ovarian epithelial cells. n = 3. *P < 0.05, **P < 0.01, ***P < 0.001
Knockdown of SRC-3 repressed the growth, migration, invasion, and stemness of ovarian cancer cells
We further investigated the role of SRC-3 in ovarian cancer cell growth, migration, invasion, and stemness. First, we tried to downregulate SRC-3 expression using sh-RNA. The mRNA level of SRC-3 was dramatically decreased by shRNA-targeting SRC-3 compared to that in the control and sh-NC groups (Fig. 2A). Furthermore, the protein level of SRC-3 was detected by western blot in SRC-3 knocked down CAOV-3 and SKOV3 cells. Consistent with our mRNA results, the protein level of SRC-3 was significantly decreased when sh-SRC-3 was transfected into CAOV-3 and SKOV3 cells (Fig. 2A). The expression level of TRAF4 was detected in SRC-3 knocked down CAOV-3 and SKOV3 cells. qRT-PCR and western blotting results showed that SRC-3 silencing dramatically reduced the mRNA and protein levels of TRAF4 in CAOV-3 and SKOV3 cells (Fig. 2B). Cell proliferation was determined using the MTT assay. As shown in Fig. 2C, SRC-3 silencing significantly decreased the viability of CAOV-3 and SKOV3 cells in a time-dependent manner (Fig. 2C). The migration and invasion of ovarian cancer cells were detected using a transwell assay. Compared to the control group, SRC-3 knockdown resulted in a significant decrease in cell migration (Fig. 2D). In accordance with the migration results, inhibition of SRC-3 expression substantially decreased the relative invasion of CAOV-3 and SKOV3 cells compared to the control group (Fig. 2E). Furthermore, the effect of SRC-3 silencing on cellular stemness was detected using spheroidization analysis. Similarly, compared to the control group, the sh-SRC-3 group showed significantly less cell spheroidization (Fig. 2F). Additionally, the expression levels of stem cell factors were detected by qRT-PCR and western blot assays. As shown in Fig. 2G, knockdown of SRC-3 profoundly decreased the mRNA levels of CD44, CD133, SOX2, OCT3/4, and NANOG in CAOV-3 and SKOV3 cells compared to the control group (Fig. 2G). Similarly, the protein levels of CD44, CD133, SOX2, OCT3/4, and NANOG dramatically decreased when SRC-3 was silenced (Fig. 2H). These results suggested that SRC-3 silencing inhibited the growth, migration, invasion, and stemness of ovarian cancer cells.
Knocking down SRC-3 repressed the growth, migration, invasion and stemness of ovarian cancer cells. A The expression of SRC-3 was decreased in sh-SRC-3 transfected CAOV-3 and SKOV3 cells by qRT-PCR and western blot assay. B The expression of TRAF4 was decreased in sh-SRC-3 transfected CAOV-3 and SKOV3 cells by qRT-PCR and western blot assay. C Cell proliferation was detected by MTT assay after being transfected with sh-SRC-3. D and E SRC-3 silence decreased cell migration and invasion by transwell assay. F The cell spheroidization test detected the number of CAOV-3 and SKOV3 cells into spheroids after knocking down SRC-3. G and H The expression of stem cell factors after silencing SRC-3 was analyzed by qRT-PCR and western blot. n = 3. *P < 0.05, **P < 0.01, ***P < 0.001
TRAF4 silencing decreased ovarian cancer cell growth and development
Previous studies have suggested that TRAF4 is critical for breast cancer development and acts downstream of SRC-3 [24]. To further explore the role of TRAF4 in ovarian cancer cell growth and development, TRAF4 was knocked down using shRNA. The mRNA and protein expression of TRAF4 was significantly decreased by sh-TRAF4 (Fig. 3A, B). The MTT assay showed that cell proliferation was sharply decreased when TRAF4 was silenced by sh-TRAF4 in a time-dependent manner compared with that in the control group (Fig. 3C). Cell migration was suppressed when TRAF4 was knocked down (Fig. 3D). Similarly, cell invasion was inhibited when TRAF4 was silenced by shRNA compared to the control group (Fig. 3E). Spheroidization analysis showed that cell spheroidization decreased dramatically when TRAF4 was silenced (Fig. 3F). In addition, qRT-PCR and western blot assays showed that knockdown of TRAF4 substantially downregulated the mRNA and protein expression of CD44, CD133, SOX2, OCT3/4, and NANOG in CAOV-3 and SKOV3 cells compared to that in the control group (Fig. 3G, H). These results demonstrated that TRAF4 plays an important role in ovarian cancer cell growth and development.
TRAF4 silence decreased ovarian cancer cell growth and development. A and B The expression of TRAF4 in CAOV-3 and SKOV3 cells after knocking down TRAF4 was analyzed by qRT-PCR and western blot. C The effect of silencing TRAF4 on cell proliferation was detected by MTT assay. D and E Transwell assay detected the effect of TRAF4 knockdown on cell migration and invasion. F The cell spheroidization experiment detected the number of cells spheroids when TRAF4 was knocked down. G and H The expression of knockdown TRAF4 stem cell factor was analyzed by qRT-PCR and western blot. n = 3. *P < 0.05, **P < 0.01, ***P < 0.001
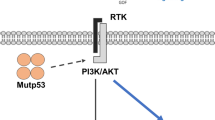
SRC-3/TRAF4 promoted ovarian cancer cell growth and development by activating the PI3K/AKT pathway
Previous studies have reported that TRAF4 activates the PI3K/AKT pathway to promote endometrial cancer development [9,10]. Currently, immunotherapies are considered for treating ovarian cancer, even though the success rate is very low [45]. Currently, no FDA approval for immunotherapy is available for ovarian cancer. The scientific community was motivated by encouraging results in other closely related tumor types [45]. As the SRC-3/TRAF4/PI3K/AKT pathway plays a vital role in ovarian cancer development, the SRC-3/TRAF4 pathway may be a promising therapeutic target for ovarian cancer treatment.
Data availability
All data generated or analyzed during this study are included in this article. The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Tew WP. Ovarian cancer in the older woman. J Geriatr Oncol. 2016;7(5):354–61.
Ottevanger PB. Ovarian cancer stem cells more questions than answers. Semin Cancer Biol. 2017;44:67–71.
Lee J, et al. Musashi-2 is a novel regulator of paclitaxel sensitivity in ovarian cancer cells. Int J Oncol. 2016;49(5):1945–52.
Schorge JO, McCann C, Del Carmen MG. Surgical debulking of ovarian cancer: what difference does it make? Rev Obstet Gynecol. 2010;3(3):111–7.
Cheng X, et al. Solute carrier family 39 member 6 gene promotes aggressiveness of esophageal carcinoma cells by increasing intracellular levels of zinc, activating phosphatidylinositol 3-kinase signaling, and up-regulating genes that regulate metastasis. Gastroenterology. 2017;152(8):1985-97.e12.
Cortez AJ, et al. Advances in ovarian cancer therapy. Cancer Chemother Pharmacol. 2018;81(1):17–38.
Keyvani V, et al. Ovarian cancer stem cells and targeted therapy. J Ovarian Res. 2019;12(1):120.
Wang J, et al. Low BCL9 expression inhibited ovarian epithelial malignant tumor progression by decreasing proliferation, migration, and increasing apoptosis to cancer cells. Cancer Cell Int. 2019;19:330.
Kotsopoulos J, Narod SA. Prophylactic sal**ectomy for the prevention of ovarian cancer: who should we target? Int J Cancer. 2020;147(5):1245–51.
Pfisterer J, et al. Abagovomab for ovarian cancer. Expert Opin Biol Ther. 2011;11(3):395–403.
Lopez-Guerrero JA, Romero I, Poveda A. Trabectedin therapy as an emerging treatment strategy for recurrent platinum-sensitive ovarian cancer. Chin J Cancer. 2015;34(1):41–9.
Sfakianos GP, Havrilesky LJ. A review of cost-effectiveness studies in ovarian cancer. Cancer Control. 2011;18(1):59–64.
Bapat SA, et al. Stem and progenitor-like cells contribute to the aggressive behavior of human epithelial ovarian cancer. Cancer Res. 2005;65(8):3025–9.
Gawlik-Rzemieniewska N, Bednarek I. The role of NANOG transcriptional factor in the development of malignant phenotype of cancer cells. Cancer Biol Ther. 2016;17(1):1–10.
Wang X, et al. SOX2 enhances the migration and invasion of ovarian cancer cells via Src kinase. PLoS ONE. 2014;9(6): e99594.
Parte SC, et al. Ovarian Cancer Stem Cells: Unraveling a Germline Connection. Stem Cells Dev. 2017;26(24):1781–803.
Thiaville MM, et al. Identification of PBX1 target genes in cancer cells by global map** of PBX1 binding sites. PLoS ONE. 2012;7(5): e36054.
Teeuwssen M, Fodde R. Wnt signaling in ovarian cancer stemness EMT, and Therapy Resistance. J Clin Med. 2019;8:1658.
Ma G, et al. SRC-3 has a role in cancer other than as a nuclear receptor coactivator. Int J Biol Sci. 2011;7(5):664–72.
Gojis O, et al. The role of steroid receptor coactivator-3 (SRC-3) in human malignant disease. Eur J Surg Oncol. 2010;36(3):224–9.
Palmieri C, et al. Expression of steroid receptor coactivator 3 in ovarian epithelial cancer is a poor prognostic factor and a marker for platinum resistance. Br J Cancer. 2013;108(10):2039–44.
Yan J, Tsai SY, Tsai MJ. SRC-3/AIB1: transcriptional coactivator in oncogenesis. Acta Pharmacol Sin. 2006;27(4):387–94.
Yan J, et al. Steroid receptor coactivator-3 and activator protein-1 coordinately regulate the transcription of components of the insulin-like growth factor/AKT signaling pathway. Cancer Res. 2006;66(22):11039–46.
Yi P, et al. SRC-3 coactivator regulates cell resistance to cytotoxic stress via TRAF4-mediated p53 destabilization. Genes Dev. 2013;27(3):274–87.
Zheng FF, et al. Rapid estrogen-induced phosphorylation of the SRC-3 coactivator occurs in an extranuclear complex containing estrogen receptor. Mol Cell Biol. 2005;25(18):8273–84.
Kedinger V, Rio MC. TRAF4, the unique family member. Adv Exp Med Biol. 2007;597:60–71.
Camilleri-Broet S, et al. TRAF4 overexpression is a common characteristic of human carcinomas. Oncogene. 2007;26(1):142–7.
Liu K, et al. TRAF4 regulates migration, invasion, and epithelial-mesenchymal transition via PI3K/AKT signaling in hepatocellular carcinoma. Oncol Res. 2017;25(8):1329–40.
**e P, et al. TRAF4 promotes endometrial cancer cell growth and migration by activation of PI3K/AKT/Oct4 signaling. Exp Mol Pathol. 2019;108:9–16.
Zhang L, et al. TRAF4 promotes TGF-β receptor signaling and drives breast cancer metastasis. Mol Cell. 2013;51(5):559–72.
Rousseau A, Rio MC, Alpy F. TRAF4, at the crossroad between morphogenesis and cancer. Cancers (Basel). 2011;3(2):2734–49.
Ren H, et al. TRAF4, a new substrate of SIAH1, participates in chemotherapy resistance of breast cancer cell by counteracting SIAH1-mediated downregulation of β-catenin. Breast Cancer Res Treat. 2020;183(2):275–89.
Zhu L, et al. Down-regulation of TRAF4 targeting RSK4 inhibits proliferation, invasion and metastasis in breast cancer xenografts. Biochem Biophys Res Commun. 2018;500(3):810–6.
Zhou J, et al. High expression of TRAF4 predicts poor prognosis in tamoxifen-treated breast cancer and promotes tamoxifen resistance. Anticancer Drugs. 2020;31(6):558–66.
Shi B, et al. Death receptor 6 promotes ovarian cancer cell migration through KIF11. FEBS Open Bio. 2018;8(9):1497–507.
De Nadai C, et al. Nitric oxide inhibits tumor necrosis factor-alpha-induced apoptosis by reducing the generation of ceramide. Proc Natl Acad Sci USA. 2000;97(10):5480–5.
Plaks V, Kong N, Werb Z. The cancer stem cell niche: how essential is the niche in regulating stemness of tumor cells? Cell Stem Cell. 2015;16(3):225–38.
Li L, Deng CX, Chen Q. SRC-3, a steroid receptor coactivator: implication in cancer. Int J Mol Sci. 2021;22(9):4760.
Rohira AD, et al. Targeting SRC coactivators blocks the tumor-initiating capacity of cancer stem-like cells. Cancer Res. 2017;77(16):4293–304.
Dasgupta S, et al. Metabolic enzyme PFKFB4 activates transcriptional coactivator SRC-3 to drive breast cancer. Nature. 2018;556(7700):249–54.
Rousseau A, et al. TRAF4 is a novel phosphoinositide-binding protein modulating tight junctions and favoring cell migration. PLoS Biol. 2013;11(12): e1001726.
Yao W, et al. TRAF4 enhances osteosarcoma cell proliferation and invasion by Akt signaling pathway. Oncol Res. 2014;22(1):21–8.
Zhou F, et al. TRAF4 mediates activation of TGF-β signaling and is a biomarker for oncogenesis in breast cancer. Sci China Life Sci. 2014;57(12):1172–6.
Aletti GD, et al. Multidisciplinary approach in the management of advanced ovarian cancer patients: a personalized approach. Results from a specialized ovarian cancer unit. Gynecol Oncol. 2017;144(3):468–73.
Chandra A, et al. Ovarian cancer: current status and strategies for improving therapeutic outcomes. Cancer Med. 2019;8(16):7018–31.
Parmar MK, et al. Paclitaxel plus platinum-based chemotherapy versus conventional platinum-based chemotherapy in women with relapsed ovarian cancer: the ICON4/AGO-OVAR-2.2 trial. Lancet. 2003;361(9375):2099–106.
Högberg T, Glimelius B, Nygren P. A systematic overview of chemotherapy effects in ovarian cancer. Acta Oncol. 2001;40(2–3):340–60.
Pfisterer J, et al. Gemcitabine plus carboplatin compared with carboplatin in patients with platinum-sensitive recurrent ovarian cancer: an intergroup trial of the AGO-OVAR, the NCIC CTG, and the EORTC GCG. J Clin Oncol. 2006;24(29):4699–707.
Sehouli J, et al. Pegylated liposomal doxorubicin (CAELYX) in patients with advanced ovarian cancer: results of a German multicenter observational study. Cancer Chemother Pharmacol. 2009;64(3):585–91.
Ferrandina G, et al. Phase III trial of gemcitabine compared with pegylated liposomal doxorubicin in progressive or recurrent ovarian cancer. J Clin Oncol. 2008;26(6):890–6.
Acknowledgements
We express our sincere gratitude to the reviewers for their constructive comments.
Funding
This work was supported by the Scientific Research Project of Hunan Provincial Health Commission (No. 202105011313), Hunan Key Laboratory of Cancer Metabolism (No. 2020TP1018), National Natural Science Foundation of China (No. 82003050), China Postdoctoral Science Foundation (No. 2022M711126) and the Natural Science Foundation of Hunan Province (No. 2020JJ5338).
Author information
Authors and Affiliations
Contributions
YW: Conceptualization; Funding acquisition; Writing-original draft; XL: Data curation; Resources; NYYW: Methodology; Formal analysis; QJL: Investigation; Software; Visualization; JW: Project administration; Supervision; Validation; Writing-review & editing. All authors have read and approved the final version of this manuscript to be published.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethical approval
The protocol of this study was approved by the ethics committee of The Affiliated Cancer Hospital of **angya School of Medicine, Central South University. The specimen collection procedure has gotten patients’ consent and the approval of the Medical Ethics Committee.
Consent for publication
The informed consent was obtained from study participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12032_2022_1944_MOESM1_ESM.tif
Supplementary file1 (TIF 3577 kb)—Supplementary Fig. 1 SRC-3/TRAF4 promoted ovarian cancer cell growth and development by activating the PI3K/AKT pathway. A The effect of LY294002 on the phosphorylation of AKT and PI3K was detected by western blot. B After LY294002 treatment, the proliferation of CAOV-3 and SKOV3 cells was detected by MTT method. C and D The cell migration and invasion after treatment with LY294002 were measured by transwell. E Detected the number of CAOV-3 and SKOV3 cells into spheroids after LY294002 treatment by cell spheronization test. F and G The expression of stem cell factors in LY294002-treated cells was analyzed by qRT-PCR and western blot. n=3. *P < 0.05, **P < 0.01, ***P < 0.001
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Wang, Y., Luo, X., Wu, N. et al. SRC-3/TRAF4 facilitates ovarian cancer development by activating the PI3K/AKT signaling pathway. Med Oncol 40, 76 (2023). https://doi.org/10.1007/s12032-022-01944-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-022-01944-0