Abstract
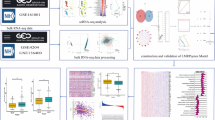
Multiple sclerosis (MS) is a common chronic autoimmune disorder of the central nervous system that predominantly affects young adults. Mounting evidence indicates that deregulation of microRNAs (miRNAs) in cerebrospinal fluid (CSF) has been implicated in MS as a potential biomarker. However, comprehensive assessments of CSF miRNAs and their target genes are lacking. Here, aberrantly expressed CSF miRNAs of MS patients were obtained from numerous studies by manual search. With detailed information on these miRNAs, we utilized online databases to screen out immune-related target genes and further performed Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses. To identify MS high-risk pathways and pivotal genes, pathway crosstalk and pathway-gene networks were constructed, followed by the establishment of a protein–protein interaction (PPI) network. The datasets collected from ArrayExpress were used to assess pivotal genes. Overall, 21 MS-related CSF miRNAs were included in this study. Subsequently, we identified 469 MS-related genes and 14 high-risk pathways. In the pathway-gene network, 27 critical MS-related genes participated in at least half of the high-risk pathways, and these genes were used to identify pivotal genes. Finally, miR-150, miR-328, and miR-34c-5p were determined to be risk miRNAs via the regulation of the pivotal risk genes MAPK1, AKT1, and VEGFA. Among them, VEGFA was validated to be significantly decreased in the CSF cells of MS patients by transcriptomic datasets. These findings may provide potential biomarkers or therapeutic targets and help elucidate the molecular mechanisms underlying the pathogenesis of MS.







Similar content being viewed by others
Data Availability
The data and other items supporting the results of the study will be made available upon reasonable request.
References
Afrasiabi A, Fewings NL, Schibeci SD, Keane JT, Booth DR, Parnell GP, Swaminathan S (2021) The interaction of human and Epstein-Barr virus miRNAs with multiple sclerosis risk loci. Int J Mol Sci. https://doi.org/10.3390/ijms22062927
Ahlbrecht J, Martino F, Pul R et al (2016) Deregulation of microRNA-181c in cerebrospinal fluid of patients with clinically isolated syndrome is associated with early conversion to relapsing-remitting multiple sclerosis. Mult Scler 22:1202–1214. https://doi.org/10.1177/1352458515613641
Amaral AJ, Andrade J, Foxall RB et al (2017) miRNA profiling of human naive CD4 T cells links miR-34c-5p to cell activation and HIV replication. EMBO J 36:346–360. https://doi.org/10.15252/embj.201694335
Bergman P, Piket E, Khademi M, James T, Brundin L, Olsson T, Piehl F, Jagodic M (2016) Circulating miR-150 in CSF is a novel candidate biomarker for multiple sclerosis. Neurol Neuroimmunol Neuroinflamm 3:e219. https://doi.org/10.1212/NXI.0000000000000219
Chen X, Liang H, Zhang J, Zen K, Zhang CY (2012) Secreted microRNAs: a new form of intercellular communication. Trends Cell Biol 22:125–132. https://doi.org/10.1016/j.tcb.2011.12.001
Cheng Z, Qiu S, Jiang L, Zhang A, Bao W, Liu P, Liu J (2012) MiR-320a is downregulated in patients with myasthenia gravis and modulates inflammatory cytokines production by targeting mitogen-activated protein kinase 1. J Clin Immunol 33:567–576. https://doi.org/10.1007/s10875-012-9834-5
Di Palo A, Siniscalchi C, Salerno M, Russo A, Gravholt CH, Potenza N (2020) What microRNAs could tell us about the human X chromosome. Cell Mol Life Sci 77:4069–4080. https://doi.org/10.1007/s00018-020-03526-7
Fitzner B, Hecker M, Zettl UK (2015) Molecular biomarkers in cerebrospinal fluid of multiple sclerosis patients. Autoimmun Rev 14:903–913. https://doi.org/10.1016/j.autrev.2015.06.001
Gandhi R, Healy B, Gholipour T et al (2013) Circulating microRNAs as biomarkers for disease staging in multiple sclerosis. Ann Neurol 73:729–740. https://doi.org/10.1002/ana.23880
Guo X, Su B, Zhou Z, Sha J (2009) Rapid evolution of mammalian X-linked testis microRNAs. BMC Genomics. https://doi.org/10.1186/1471-2164-10-97
Haghikia A, Haghikia A, Hellwig K, Baraniskin A, Holzmann A, Décard B, Thum T, Gold R (2012) Regulated microRNAs in the CSF of patients with multiple sclerosis a case-control study. Neurology 79:2166–2170
Hu Z, Cui Y, Qiao X et al (2018) Silencing miR-150 ameliorates experimental autoimmune encephalomyelitis. Front Neurosci. https://doi.org/10.3389/fnins.2018.00465
Huang XL, Zhang L, Li JP, Wang YJ, Duan Y, Wang J (2015) MicroRNA-150: A potential regulator in pathogens infection and autoimmune diseases. Autoimmunity 48:503–510. https://doi.org/10.3109/08916934.2015.1072518
Iacobaeus E, Amoudruz P, Strom M et al (2011) The expression of VEGF-A is down regulated in peripheral blood mononuclear cells of patients with secondary progressive multiple sclerosis. PLoS ONE 6:e19138. https://doi.org/10.1371/journal.pone.0019138
International Multiple Sclerosis Genetics Consortium (2019) Multiple sclerosis genomic map implicates peripheral immune cells and microglia in susceptibility. Science. https://doi.org/10.1126/science.aav7188
International Multiple Sclerosis Genetics Consortium, Beecham AH, Patsopoulos NA et al (2013) Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat Genet 45:1353–1360. https://doi.org/10.1038/ng.2770
Khalifa O, Pers YM, Ferreira R, Senechal A, Jorgensen C, Apparailly F, Duroux-Richard I (2016) X-Linked miRNAs associated with gender differences in rheumatoid arthritis. Int J Mol Sci. https://doi.org/10.3390/ijms17111852
Kramer S, Haghikia A, Bang C et al (2019) Elevated levels of miR-181c and miR-633 in the CSF of patients with MS: a validation study. Neurol Neuroimmunol Neuroinflamm 6:e623. https://doi.org/10.1212/NXI.0000000000000623
Li Z, Liu Y, Jia A, Cui Y, Feng J (2021) Cerebrospinal fluid cells immune landscape in multiple sclerosis. J Transl Med. https://doi.org/10.1186/s12967-021-02804-7
Lovato L, Willis SN, Rodig SJ et al (2011) Related B cell clones populate the meninges and parenchyma of patients with multiple sclerosis. Brain 134:534–541. https://doi.org/10.1093/brain/awq350
Luo D, Fu J (2018) Identifying characteristic miRNAs-genes and risk pathways of multiple sclerosis based on bioinformatics analysis. Oncotarget 9:5287–5300
Luo D, Wang J, Zhang X, Rang X, Xu C, Fu J (2020) Identification and functional analysis of specific MS risk miRNAs and their target genes. Mult Scler Relat Disord. https://doi.org/10.1016/j.msard.2020.102044
Majd M, Hosseini A, Ghaedi K et al (2018) MiR-9–5p and miR-106a-5p dysregulated in CD4(+) T-cells of multiple sclerosis patients and targeted essential factors of T helper17/regulatory T-cells differentiation. Iran J Basic Med Sci 21:277–283. https://doi.org/10.22038/ijbms.2018.25382.6275
Mandolesi G, De Vito F, Musella A et al (2017) miR-142-3p is a key regulator of IL-1beta-dependent synaptopathy in neuroinflammation. J Neurosci 37:546–561. https://doi.org/10.1523/JNEUROSCI.0851-16.2016
Mohammed EM (2020) Environmental influencers, MicroRNA, and multiple sclerosis. J Cent Nerv Syst Dis. https://doi.org/10.1177/1179573519894955
Ouyang S, Zeng Q, Tang N et al (2019) Akt-1 and Akt-2 differentially regulate the development of experimental autoimmune encephalomyelitis by controlling proliferation of thymus-derived regulatory T cells. J Immunol 202:1441–1452. https://doi.org/10.4049/jimmunol.1701204
Padgett KA, Lan RY, Leung PC et al (2009) Primary biliary cirrhosis is associated with altered hepatic microRNA expression. J Autoimmun 32:246–253. https://doi.org/10.1016/j.jaut.2009.02.022
Pappalardo J, Zhang L, Pecsok M et al (2020) Transcriptomic and clonal characterization of T cells in the human central nervous system. Sci Immunol 5:eabb8786
Perdaens O, Dang HA, D’Auria L, van Pesch V (2020) CSF microRNAs discriminate MS activity and share similarity to other neuroinflammatory disorders. Neurol Neuroimmunol Neuroinflamm. https://doi.org/10.1212/NXI.0000000000000673
Quintana E, Ortega FJ, Robles-Cedeno R et al (2017) miRNAs in cerebrospinal fluid identify patients with MS and specifically those with lipid-specific oligoclonal IgM bands. Mult Scler 23:1716–1726. https://doi.org/10.1177/1352458516684213
Rahimirad S, Navaderi M, Alaei S, Sanati MH (2021) Identification of hsa-miR-106a-5p as an impact agent on promotion of multiple sclerosis using multi-step data analysis. Neurol Sci 42:3791–3799. https://doi.org/10.1007/s10072-020-04979-1
Regev K, Paul A, Healy B et al (2016) Comprehensive evaluation of serum microRNAs as biomarkers in multiple sclerosis. Neurol Neuroimmunol Neuroinflamm 3:e267. https://doi.org/10.1212/NXI.0000000000000267
Reich DS, Lucchinetti CF, Calabresi PA (2018) Multiple sclerosis. N Engl J Med 378:169–180. https://doi.org/10.1056/NEJMra1401483
Romania P, Cifaldi L, Pignoloni B et al (2017) Identification of a genetic variation in ERAP1 aminopeptidase that prevents human cytomegalovirus miR-UL112-5p-mediated immunoevasion. Cell Rep 20:846–853. https://doi.org/10.1016/j.celrep.2017.06.084
Schafflick D, Xu CA, Hartlehnert M et al (2020) Integrated single cell analysis of blood and cerebrospinal fluid leukocytes in multiple sclerosis. Nat Commun. https://doi.org/10.1038/s41467-019-14118-w
Skulina C, Schmidt S, Dornmair K et al (2004) Multiple sclerosis brain-infiltrating CD8+ T cells persist as clonal expansions in the cerebrospinal fluid and blood. PNAS 101:2428–2433
Stanojlovic M, Pang X, Lin Y, Stone S, Cvetanovic M, Lin W (2016) Inhibition of vascular endothelial growth factor receptor 2 exacerbates loss of lower motor neurons and axons during experimental autoimmune encephalomyelitis. PLoS ONE 11:e0160158. https://doi.org/10.1371/journal.pone.0160158
Stoicea N, Du A, Lakis DC, Tipton C, Arias-Morales CE, Bergese SD (2016) The MiRNA journey from theory to practice as a CNS biomarker. Front Genet. https://doi.org/10.3389/fgene.2016.00011
Szklarczyk D, Morris JH, Cook H et al (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45:D362–D368. https://doi.org/10.1093/nar/gkw937
Tham E, Gielen AW, Khademi M, Martin C, Piehl F (2006) Decreased expression of VEGF-A in rat experimental autoimmune encephalomyelitis and in cerebrospinal fluid mononuclear cells from patients with multiple sclerosis. Scand J Immunol 64:609–622. https://doi.org/10.1111/j.1365-3083.2006.01851.x
Thamilarasan M, Koczan D, Hecker M, Paap B, Zettl UK (2012) MicroRNAs in multiple sclerosis and experimental autoimmune encephalomyelitis. Autoimmun Rev 11:174–179. https://doi.org/10.1016/j.autrev.2011.05.009
Thompson AJ, Baranzini SE, Geurts J, Hemmer B, Ciccarelli O (2018) Multiple sclerosis. Lancet 391:1622–1636. https://doi.org/10.1016/s0140-6736(18)30481-1
Tu Y, Hu Y (2021) MiRNA-34c-5p protects against cerebral ischemia/reperfusion injury: involvement of anti-apoptotic and anti-inflammatory activities. Metab Brain Dis 36:1341–1351. https://doi.org/10.1007/s11011-021-00724-5
Vistbakka J, Elovaara I, Lehtimaki T, Hagman S (2017) Circulating microRNAs as biomarkers in progressive multiple sclerosis. Mult Scler 23:403–412. https://doi.org/10.1177/1352458516651141
Wu R, He Q, Chen H et al (2017) MicroRNA-448 promotes multiple sclerosis development through induction of Th17 response through targeting protein tyrosine phosphatase non-receptor type 2 (PTPN2). Biochem Biophys Res Commun 486:759–766. https://doi.org/10.1016/j.bbrc.2017.03.115
Zeitelhofer M, Adzemovic MZ, Gomez-Cabrero D et al (2017) Functional genomics analysis of vitamin D effects on CD4+ T cells in vivo in experimental autoimmune encephalomyelitis. Proc Natl Acad Sci U S A 114:E1678–E1687. https://doi.org/10.1073/pnas.1615783114
Zhang Z, Xue Z, Liu Y et al (2018) MicroRNA-181c promotes Th17 cell differentiation and mediates experimental autoimmune encephalomyelitis. Brain Behav Immun 70:305–314. https://doi.org/10.1016/j.bbi.2018.03.011
Zhao X, Li S, Wang Z, Feng Y (2021) miR-101-3p negatively regulates inflammation in systemic lupus erythematosus via MAPK1 targeting and inhibition of the NF-κB pathway. Mol Med Rep. https://doi.org/10.3892/mmr.2021.11998
Zhou B, Wang S, Mayr C, Lodish H (2008) miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. PNAS 104:7080–7085
Acknowledgements
Thanks to Zhiqiang Zhang for suggestions and English editing on the manuscript.
Funding
This work was supported by a grant from the Provincial Natural Science Foundation of Heilongjiang province (No. ZD2020H004).
Author information
Authors and Affiliations
Contributions
YS and ZL designed research, analyzed the data, created the figures and tables, and wrote the manuscript. XR and YW analyzed the data and revised the original manuscript. JF designed research and provided final critical revisions. All authors critically read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Su, Y., Li, Z., Rang, X. et al. Integrated Analysis and Identification of CSF-Derived Risk miRNAs and Pivotal Genes in Multiple Sclerosis. J Mol Neurosci 72, 1916–1928 (2022). https://doi.org/10.1007/s12031-022-02007-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-022-02007-9