Abstract
Background
Ferroptosis and lncRNAs both play crucial roles in cancers. But the roles of ferroptosis-related lncRNAs (FRLncs) in HBV-related HCC (HBV-HCC) remain ambiguous.
Methods
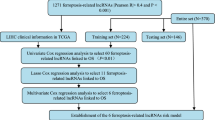
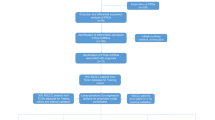
The gene expression profile and clinical data were originated from the Cancer Genome Atlas (TCGA) database. The risk signature was constructed by FRLncs based on the Cox regression analysis. The survival curve, Cox regression analysis, and time-dependent receiver operating characteristic (ROC) curve were adopted to verify the independence and reliability of the signature. A nomogram was established. Immune-infiltrating cells, immune functions, and checkpoints were analyzed.
Results
A risk signature composed of 7 FRLncs (LINC00942, AC131009.1, POLH-AS1, AC090772.3, MKLN1-AS, AC009403.1, AL031985.3) was constructed and divided HBV-HCC patients into high- and low-risk groups. Patients in the high-risk group showed a poor prognosis. The area under curves (AUC) of the signature for 1-, 3-, and 5-year was satisfactory. A nomogram composed of gender, stage, age, grade, and risk signature was established. The risk signature and nomogram displayed appreciable independence and reliability in HBV-HCC patients. The T-cell CD8 + , monocyte, and macrophage M1 were expressed differently significantly in HCC patients, while macrophage M2 showed an obvious difference in the HBV-HCC patients between the different risk groups. PDCD1 and CTL4 were expressed higher in the high-risk group of HCC patients.
Conclusion
A 7-lncRNA signature was identified as a potential prognostic predictor for HBV-HCC patients. Immune therapy may be a promising strategy for HCC patients, especially HBV-HCC patients.







Similar content being viewed by others
Data Availability
The RNA sequencing data with clinical information were acquired from The Cancer Genome Atlas (TCGA) (https://portal.gdc.cancer.gov/). The ferroptosis-related genes were acquired from the FerrDb V2 (http://www.zhounan.org/ferrdb/current/). The Infiltration Estimation for all TCGA tumors was downloaded from TIMER2.0 (http://timer.cistrome.org/). The authors confirm that the data supporting the findings of this study are available within the article.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA Cancer J Clin. 2022;72:7–33.
Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Wei X, Yi X, Zhu XH, Jiang DS. Posttranslational modifications in ferroptosis. Oxid Med Cell Longev. 2020;2020:8832043.
Ma S, Henson ES, Chen Y, Gibson SB. Ferroptosis is induced following siramesine and lapatinib treatment of breast cancer cells. Cell Death Dis. 2016;7:e2307.
Ghoochani A, Hsu EC, Aslan M, et al. Ferroptosis inducers are a novel therapeutic approach for advanced prostate cancer. Can Res. 2021;81:1583–94.
Yao F, Deng Y, Zhao Y, et al. A targetable LIFR-NF-kappaB-LCN2 axis controls liver tumorigenesis and vulnerability to ferroptosis. Nat Commun. 2021;12:7333.
Gao R, Kalathur RKR, Coto-Llerena M, et al. YAP/TAZ and ATF4 drive resistance to sorafenib in hepatocellular carcinoma by preventing ferroptosis. EMBO Mol Med. 2021;13:e14351.
Zhang H, Deng T, Liu R, et al. CAF secreted miR-522 suppresses ferroptosis and promotes acquired chemo-resistance in gastric cancer. Mol Cancer. 2020;19:43.
Tsagakis I, Douka K, Birds I, Aspden JL. Long non-coding RNAs in development and disease: conservation to mechanisms. J Pathol. 2020;250:480–95.
Schmitz SU, Grote P, Herrmann BG. Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci. 2016;73:2491–509.
Huang Z, Zhou JK, Peng Y, He W, Huang C. The role of long noncoding RNAs in hepatocellular carcinoma. Mol Cancer. 2020;19:77.
Nia A, Dhanasekaran R. Genomic landscape of HCC. Curr Hepatol Rep. 2020;19:448–61.
Woo Y, Lee HJ, Jung YM, Jung YJ. Regulated necrotic cell death in alternative tumor therapeutic strategies. Cells. 2020;9.
Zhu Y, Zhou B, Hu X, et al. lncRNA LINC00942 promotes chemoresistance in gastric cancer by suppressing MSI2 degradation to enhance c-Myc mRNA stability. Clin Transl Med. 2022;12:e703.
Wang R, Wang X, Zhang J, Liu Y. LINC00942 Promotes tumor proliferation and metastasis in lung adenocarcinoma via FZD1 upregulation. Technol Cancer Res Treat. 2021;20:1533033820977526. https://doi.org/10.1177/1533033820977526.
Wang W, Ye Y, Zhang X, Ye X, Liu C, Bao L. Construction of a necroptosis-associated long non-coding RNA signature to predict prognosis and immune response in hepatocellular carcinoma. Front Mol Biosci. 2022;9:937979.
Tang Y, Zhang H, Chen L, Zhang T, Xu N, Huang Z. Identification of hypoxia-related prognostic signature and competing endogenous RNA regulatory axes in hepatocellular carcinoma. Int J Mol Sci. 2022;23.
Guo C, Zhou S, Yi W, et al. SOX9/MKLN1-AS axis induces hepatocellular carcinoma proliferation and epithelial-mesenchymal transition. Biochem Genet. 2022;60:1914–33.
Guo C, Zhou S, Yi W, et al. Long non-coding RNA muskelin 1 antisense RNA (MKLN1-AS) is a potential diagnostic and prognostic biomarker and therapeutic target for hepatocellular carcinoma. Exp Mol Pathol. 2021;120:104638. https://doi.org/10.1016/j.yexmp.2021.104638. Epub 102021 Apr 104618.
Pan G, Zhang J, You F, et al. ETS proto-oncogene 1-activated muskelin 1 antisense RNA drives the malignant progression of hepatocellular carcinoma by targeting miR-22-3p to upregulate ETS proto-oncogene 1. Bioengineered. 2022;13:1346–58. https://doi.org/10.1080/21655979.2021.2017565.
Wu ZH, Li ZW, Yang DL, Liu J. Development and validation of a pyroptosis-related long non-coding RNA signature for hepatocellular carcinoma. Front Cell Dev Biol. 2021;9:713925.
Wei H, Xu Z, Chen L, et al. Long non-coding RNA PAARH promotes hepatocellular carcinoma progression and angiogenesis via upregulating HOTTIP and activating HIF-1н╠/VEGF signaling. Cell Death Dis. 2022;13:102. https://doi.org/10.1038/s41419-022-04505-5.
Chen H, Nio K, Tang H, et al. BMP9-ID1 signaling activates HIF-1н╠ and VEGFA expression to promote tumor angiogenesis in hepatocellular carcinoma. Int J Mol Sci. 2022;23:1475. https://doi.org/10.3390/ijms23031475.
Dixon SJ, Patel DN, Welsch M, et al. Pharmacological inhibition of cystine-glutamate exchange induces endoplasmic reticulum stress and ferroptosis. Elife. 2014;3:e02523. https://doi.org/10.7554/eLife.02523.
Hu Z, Li L, Li M, et al. miR-21–5p inhibits ferroptosis in hepatocellular carcinoma cells by regulating the AKT/mTOR signaling pathway through MELK. J Immunol Res. 2023;2023:8929525. https://doi.org/10.1155/2023/8929525.eCollection8922023.
Abdolmohammadi MH, Roozbehani M, Hamzeloo-Moghadam M, Heidari F, Fallahian F. Targeting PPARнЁ/ NF-н╨B signaling pathway by Britannin, a sesquiterpene lactone from Inula aucheriana DC., in Gastric Cancer. Anticancer Agents Med Chem. 2023;18:1.871520623666231e+24.
Liu X, Qian D, Liu H, et al. Genetic variants of the peroxisome proliferator-activated receptor (PPAR) signaling pathway genes and risk of pancreatic cancer. Mol Carcinog. 2020;59:930–9. https://doi.org/10.1002/mc.23208. Epub 22020 May 23205.
Yang X, Yang R, Zhang Y, et al. **anlinglianxiafang Inhibited the growth and metastasis of triple-negative breast cancer via activating PPARн Ё/AMPK signaling pathway. Biomed Pharmacother. 2023;165:115164. https://doi.org/10.1016/j.biopha.2023.115164. Epub 112023 Jul 115119.
Lu P, Li X, Li B, et al. The mitochondrial-derived peptide MOTS-c suppresses ferroptosis and alleviates acute lung injury induced by myocardial ischemia reperfusion via PPARн Ё signaling pathway. Eur J Pharmacol. 2023;953:175835. https://doi.org/10.1016/j.ejphar.2023.175835. Epub 172023 Jun 175837.
Liao P, Wang W, Wang W, et al. CD8(+) Tб═cells and fatty acids orchestrate tumor ferroptosis and immunity via ACSL4. Cancer Cell. 2022;40:365-378.e366. https://doi.org/10.1016/j.ccell.2022.02.003. Epub 2022 Feb 1024.
Wang W, Green M, Choi JE, et al. CD8(+) T cells regulate tumour ferroptosis during cancer immunotherapy. Nature. 2019;569:270–4. https://doi.org/10.1038/s41586-019-1170-y. Epub 42019 May 41581.
Tao L, Li D, Mu S, Tian G, Yan G. lncRNA MAPKAPK5_AS1 facilitates cell proliferation in hepatitis B virus -related hepatocellular carcinoma. Lab Invest. 2022;102:494–504. https://doi.org/10.1038/s41374-022-00731-9. Epub 42022 Mar 41379.
Jiang Z, Lim SO, Yan M, et al. TYRO3 induces anti-PD-1/PD-L1 therapy resistance by limiting innate immunity and tumoral ferroptosis. J Clin Invest. 2021;131:e139434. https://doi.org/10.1172/JCI139434.
Hao X, Zheng Z, Liu H, et al. Inhibition of APOC1 promotes the transformation of M2 into M1 macrophages via the ferroptosis pathway and enhances anti-PD1 immunotherapy in hepatocellular carcinoma based on single-cell RNA sequencing. Redox Biol. 2022;56:102463. https://doi.org/10.1016/j.redox.2022.102463. Epub 102022 Sep 102462.
Shi L, Liu Y, Li M, Luo Z. Emerging roles of ferroptosis in the tumor immune landscape: from danger signals to anti-tumor immunity. FEBS J. 2022;289:3655–65. https://doi.org/10.1111/febs.16034. Epub 12021 Jun 16011.
Funding
This work was supported by the Natural Foundation of Shandong Province (No. ZR2020QH035 and No. ZR2021QH195) and the Medical Health Science and Technology Development Plan Project of Shandong Province (No. 202103030765).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Wenwen Wang, Lifen Wang, Tong mu, Chunxia Song and **hua Hu. The first draft of the manuscript was written by Wenwen Wang and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical Statement
The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key findings
1. Prognostic signature constructed of seven ferroptosis-related lncRNAs displayed appreciable independence and reliability both in HBV and HBV-HCC patients.
2. The risk signature stratified patients into high- and low-risk groups which could help guide the personalized therapy in the future.
What is known and what is new?
• Both the lncRNAs and ferroptosis play important roles in the multiple cancers.
• Ferroptosis-related lncRNAs composed a risk signature which was considered to be an independent prognostic factor for HBV-HCC patients and had good predictive accuracy for long-term survival probability.
What is the implication, and what should change now?
• Ferroptosis combined with immune therapy may be a promising strategy for all HCC patients, especially HBV-HCC patients. Further, in vitro and in vivo studies were needed to verify.
• the relevant mechanisms.
Supplementary Information
Below is the link to the electronic supplementary material.
12029_2023_977_MOESM1_ESM.zip
Supplemental Table 1: 484 FRGs downloaded from the FerrDb V2 database. Supplemental Table 2: 1317 lncRNAs were identified as FRLncs through the co-expression analysis. Supplemental Table 3: 141 differently expressed ferroptosis related genes were recognized based on the differential expression analysis. Supplementary Table 4: 784 differentially expressed ferroptosis related lncRNAs were discerned according to the differential expression analysis. Supplementary Table 5: The median value of each FRLncs calculated by the SPSS software. Supplementary Table 6: The risk signature constructed based on the LASSO regression. Supplementary Figure 1: (A) 12 FRLncs were screened in the LASSO regression model. (B) regression coefficient profiles of 12 FRLncs in the LASSO model. (C) The ROC curve of the risk signature constructed based on the LASSO regression. (ZIP 506 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, W., Wang, L., Song, C. et al. Prognostic Signature Constructed of Seven Ferroptosis-Related lncRNAs Predicts the Prognosis of HBV-Related HCC. J Gastrointest Canc 55, 444–456 (2024). https://doi.org/10.1007/s12029-023-00977-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12029-023-00977-6