Abstract
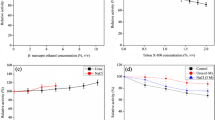
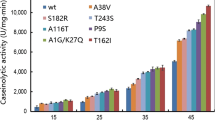
An alkaline protease (DHAP) from Bacillus pumilus has shown great potential in hide dehairing. To get better insights on its catalytic properties for application, the substrate specificity and thermostability were investigated using five natural proteins and nine synthetic peptides. The results showed that DHAP could hydrolyze five proteins tested here in different specificity. Collagen, a component of animal skin, was more resistant to hydrolysis than casein, fibrin, and gelatin. Among the synthetic peptides, the enzyme showed activity mainly with tetrapeptide substrates with the catalytic efficiency in order of Phe>Leu>Ala at P1 site, although k m value for AAVA-pN is much lower than that for AAPL-pN and AAPF-pN. With tripeptide substrates, smaller side-chain group (Gly) at P1 site was not hydrolyzed by DHAP. The enzyme showed good thermostability below 60 °C, and lost activity so quickly above 70 °C. The thermostability was largely dependent on metal ion, especially Ca2+, although other ions, like Mg2+, Mn2+, and Co2+, could sustain stability at certain extent within limited time. Cu2+, Fe2+, as well as Al3+, did not support the enzyme to retain activity at 60 °C even in 5 min. In addition, the selected metal ions could coordinate calcium in improvement or destruction of thermostability for DHAP.






Similar content being viewed by others
References
Rao, M. B., Tanksale, A. M., Ghatge, M. S., & Deshpande, V. V. (1998). Microbiology and Molecular Biology Reviews, 62, 597–635.
Bryan, P. N. (2000). Biochimica et Biophysica Acta, 1543, 203–222.
Thanilaivelan, P., Rao, J. R., Nair, B. U., & Ramasami, T. (2004). Trends in Biotechnology, 22, 181–188. doi:10.1016/j.tibtech.2004.02.008.
Nilegaonkar, S. S., Zambare, V. P., Kanekar, P. P., Dhakephalkar, P. K., & Sarnaik, S. S. (2007). Bioresource Technology, 98, 1238–1245. doi:10.1016/j.biortech.2006.05.003.
Macedo, A. J., Beys da Silva, W. O., Gava, R., Driemeier, D., Henriques, J. A. P., & Termignoni, C. (2005). Applied and Environmental Microbiology, 71, 594–596. doi:10.1128/AEM.71.1.594-596.2005.
Wang, H. Y., Liu, D. M., Liu, Y., Cheng, C. F., Ma, Q. Y., Huang, Q., et al. (2007). Letters in Applied Microbiology, 44, 1–6. doi:10.1111/j.1472-765X.2006.02039.x.
Huang, Q., Peng, Y., Li, X., Wang, H. F., & Zhang, Y. Z. (2003). Current Microbiology, 46, 169–173. doi:10.1007/s00284-002-3850-2.
Pan, J., Huang, Q., & Zhang, Y. Z. (2004). Current Microbiology, 49, 165–169. doi:10.1007/s00284-004-4305-8.
Kumar, C. G. (2002). Letters in Applied Microbiology, 34, 13–17. doi:10.1046/j.1472-765x.2002.01044.x.
Han, X. Q., & Damodaran, S. (1998). Journal of Agricultural and Food Chemistry, 46, 3596–3603. doi:10.1021/jf980361m.
Takahashi, M., Sekine, T., Kuba, N., Nakamori, S., Yasuda, M., & Takagi, H. (2004). Journal of Biochemistry, 136, 549–556. doi:10.1093/jb/mvh158.
Yasuda, M., Aoyama, M., Sakaguchi, M., Nakachi, K., & Kobamoto, N. (1999). Applied Microbiology and Biotechnology, 51, 474–479. doi:10.1007/s002530051419.
Aoyama, M., Yasuda, M., Nakachi, K., Kobamoto, N., Oku, H., & Kato, F. (2000). Applied Microbiology and Biotechnology, 53, 390–395. doi:10.1007/s002530051631.
Miyaji, T., Otta, Y., Makagawa, T., Watanabe, T., Niimura, Y., & Tomizuka, N. (2006). Letters in Applied Microbiology, 42, 242–247. doi:10.1111/j.1472-765X.2005.01851.x.
Jaouadi, B., Ellouz-Chaabouni, S., Rhimi, M., & Bejar, S. (2008). Biochimie, 90, 1291–1305. doi:10.1016/j.biochi.2008.03.004.
Gupta, R., Beg, Q. K., & Lorenz, P. (2002). Applied Microbiology and Biotechnology, 59, 15–32. doi:10.1007/s00253-002-0975-y.
Bakhtiar, S., Estiveira, R. J., & Hatti-Kaul, R. (2005). Enzyme and Microbial Technology, 37, 534–540. doi:10.1016/j.enzmictec.2005.04.003.
Perona, J. J., & Craik, C. S. (1995). Protein Science, 4, 337–360.
Strausberg, S. L., Ruan, B., Fisher, K. E., Alexander, P. A., & Bryan, P. N. (2005). Biochemistry, 44, 3272–3279. doi:10.1021/bi047806m.
Alexander, P. A., Ruan, B., & Bryan, P. N. (2001). Biochemistry, 40, 10634–10639. doi:10.1021/bi010797m.
Ghorbel, B., Sellami-kamoun, A., & Nasri, M. (2003). Enzyme and Microbial Technology, 32, 513–518. doi:10.1016/S0141-0229(03)00004-8.
Alexander, P. A., Ruan, B., Strausberg, S. L., & Bryan, P. N. (2001). Biochemistry, 40, 10640–10644. doi:10.1021/bi010798e.
Kidd, R. D., Yennawar, H. P., Sears, P., Wong, C.-H., & Farber, G. K. (1996). Journal of the American Chemical Society, 118, 1645–1650. doi:10.1021/ja953443p.
Lee, S., & Jang, D. J. (2001). Biophysical Journal, 81, 2972–2978.
Podsiadlo, P., Komiyama, T., Fuller, R. S., & Blum, O. (2004). The Journal of Biological Chemistry, 279, 36219–36227. doi:10.1074/jbc.M400338200.
Acknowledgment
This work is supported by the “863” High Tech Program (2006AA02Z221) from the Department of Science and Technology of China to H. F.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wan, MY., Wang, HY., Zhang, YZ. et al. Substrate Specificity and Thermostability of the Dehairing Alkaline Protease from Bacillus pumilus . Appl Biochem Biotechnol 159, 394–403 (2009). https://doi.org/10.1007/s12010-008-8497-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-008-8497-4