Abstract
Under salt stress, the plants need to maintain a low Na+ concentration and Na+/K+ ratio in the cell cytoplasm to keep normal growth and development. High-affinity K+ transporter (HKT) genes are known to play an important role in regulating the transportation of Na+ and K+ in higher plants. However, reports on its potential role in conferring stress tolerance in cotton are rare. In a previous study, we isolated a potassium transporter SbHKT1 from halophyte Salicornia bigelovii. With the intention to assess whether the SbHKT1 gene would improve salt tolerance in cotton, cotton plants overexpressing SbHKT1 were generated by Agrobacterium-mediated transgenic technology. Overexpression of SbHKT1 in cotton increased germination rate and biomass as well as root systems compared with wild-type plants. Transgenic cotton had significantly higher K+ content, lower Na+ content, and lower Na+/K+ ratio than wild-type plants in leaves, stems and roots under salt stress. Moreover, there were significant higher activity of antioxidant enzymes including SOD, POD, and CAT and lower malondialdehyde content which means better cell membrane integrity in transgenic cotton compared to control plants. These results indicated that overexpressing SbHKT1 in cotton improved salt tolerance by increasing the capacity of K+ uptake, K+/Na+ homeostasis, and the scavenging of reactive oxygen species.





Similar content being viewed by others
References
Ali Z, Park HC, Ali A, Oh DH, Aman R, Kropornicka A et al (2012) TsHKT1;2, a HKT1 homolog from the extremophile Arabidopsis relative Thellungiella salsuginea, shows K+ specificity in the presence of NaCl. Plant Physiol 158:1463–1474
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285:1256–1258
Ardie SW, **e L, Takahashi R, Liu S, Takano T (2009) Cloning of a high-affinity K+ transporter gene PutHKT2;1 from Puccinellia tenuiflora and its functional comparison with OsHKT2;1 from rice in yeast and Arabidopsis. J Exp Bot 60:3491–3502
Ashraf M (2002) Salt tolerance of cotton: some new advances. Crit Rev Plant Sci 21:1–30
Fricke W, Akhiyarova G, Veselov D, Kudoyarova G (2004) Rapid and tissue-specific changes in ABA and in growth rate in response to salinity in barley leaves. J Exp Bot 55:1115–1123
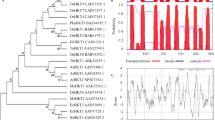
Gao YY, Zhang BL, Yang YW, Shen XL, Ni WC (2010) Cloning, expression pattern and bioinformatic analysis of high-affinity potassium transporter gene SbHKT1 from Halophyte Salicornia bigelovi. Genom Appl Biol 29:646–652 (In Chinese)
Inan G, Zhang Q, Li P, Wang Z, Cao Z, Zhang H et al (2004) Salt cress. A halophyte and cryophyte Arabidopsis relative model system and its applicability to molecular genetic analyses of growth and development of extremophiles. Plant Physiol 135:1718–1737
James RA, Blake C, Byrt CS, Munns R (2011) Major genes for Na+ exclusion, Nax1 and Nax2 (wheat HKT1;4 and HKT1;5), decrease Na+ accumulation in bread wheat leaves under saline and waterlogged conditions. J Exp Bot 62:2939–2947
Jiang Z, Song G, Shan X, Wei Z, Liu Y, Jiang C et al (2018) Association analysis and identification of ZmHKT1;5 variation with salt-stress tolerance. Front Plant Sci 9:1485
Li F, Wu S, Lü F, Chen T, Ju M, Wang H et al (2009) Modified fiber qualities of the transgenic cotton expressing a silkworm fibroin gene. Chin Sci Bull 54:1210–1216
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C et al (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30:360
Nawaz I, Iqbal M, Bliek M, Schat H (2017) Salt and heavy metal tolerance and expression levels of candidate tolerance genes among four extremophile Cochlearia species with contrasting habitat preferences. Sci Total Environ 584–585:731–741
Qiu L, Wu D, Ali S, Cai S, Dai F, ** X et al (2011) Evaluation of salinity tolerance and analysis of allelic function of HvHKT1 and HvHKT2 in Tibetan wild barley. Theor Appl Genet 122:695–703
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY et al (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146
Rus A, Yokoi S, Sharkhuu A, Reddy M, Lee B-H, Matsumoto TK et al (2001) AtHKT1 is a salt tolerance determinant that controls Na+ entry into plant roots. Proc Natl Acad Sci 98:14150
Shi H, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477
Taji T, Seki M, Satou M, Sakurai T, Kobayashi M, Ishiyama K et al (2004) Comparative genomics in salt tolerance between Arabidopsis and Arabidopsis-related halophyte salt cress using Arabidopsis microarray. Plant Physiol 135:1697–1709
Volkov V, Wang B, Dominy PJ, Fricke W, Amtmann A (2004) Thellungiella halophila, a salt-tolerant relative of Arabidopsis thaliana, possesses effective mechanisms to discriminate between potassium and sodium. Plant Cell Environ 27:1–14
Wang S, Zhao G, Gao Y, Tang Z, Zhang C (2005) Puccinellia tenuiflora exhibits stronger selectivity for K+ over Na+ than wheat. J Plant Nutr 27:1841–1857
Wang TT, Ren ZJ, Liu ZQ, Feng X, Guo RQ, Li BG et al (2014) SbHKT1;4, a member of the high-affinity potassium transporter gene family from Sorghum bicolor, functions to maintain optimal Na+/K+ balance under Na+ stress. J Integr Plant Biol 56:315–332
Wu SJ, Wang HH, Li FF, Chen TZ, Zhang J, Jiang YJ et al (2008) Enhanced Agrobacterium-mediated transformation of embryogenic calli of upland cotton via efficient selection and timely subculture of somatic embryos. Plant Mol Biol Rep 26:174–185
Zhang M, Cao Y, Wang Z, Wang ZQ, Shi J, Liang X et al (2018a) A retrotransposon in an HKT1 family sodium transporter causes variation of leaf Na+ exclusion and salt tolerance in maize. New Phytol 217:1161–1176
Zhang Y, Fang J, Wu X, Dong L (2018b) Na+/K+ balance and transport regulatory mechanisms in weedy and cultivated rice (Oryza sativa L.) under salt stress. BMC Plant Biol 18:375
Zhu M, Shabala L, Cuin TA, Huang X, Zhou M, Munns R et al (2016) Nax loci affect SOS1-like Na+/H+ exchanger expression and activity in wheat. J Exp Bot 67:835–844
Funding
This work was supported by Grants from the Major Project of the National Transgene (2018ZX0800915B), State Key Laboratory of Cotton Biology (CB2018A12), Jiangsu Planned Projects for Postdoctoral Research Funds (2018K227C) and Jiangsu Collaborative Innovation Center for Modern Crop Production.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by H. Peng.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Guo, Q., Meng, S., Tao, S. et al. Overexpression of a samphire high-affinity potassium transporter gene SbHKT1 enhances salt tolerance in transgenic cotton. Acta Physiol Plant 42, 36 (2020). https://doi.org/10.1007/s11738-020-3027-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-020-3027-2