Abstract
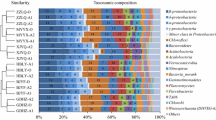
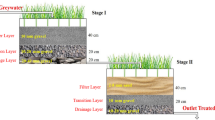
The vertical flow constructed wetlands (VFCW) for the treatment of domestic wastewater has become a conventional and cost effective treatment system with one of the major disadvantage of elevated nitrate concentrations. The present study makes an effort in providing a new design of anaerobic denitrification unit termed as anaerobic chamber (AC) which was introduced after two-stage VFCW to remove nitrates from the treated wastewater (WW). The AC provided all the essential conditions of effective denitrification such as anaerobic environment with enough carbon and nitrogen source. To understand the pollutant removal mechanism in AC, microbial diversity and functional annotation was studied by metagenomic analysis of sequences obtained from biofilm formed in AC. The efficiency of AC was measured with respect to physicochemical wastewater quality parameters. The removal efficiencies were 88, 65, 43 and 27% for total nitrogen, nitrate (NO3), ammoniacal-nitrogen (NH4) and ortho-phosphate respectively. The microbial flora was much more diverse and unique pertaining to anaerobic microbes in AC compared to WW with total of 954 and 1191 genuses respectively with minimum abundance of 10 hits. The metagenomes exhibited 188% more Archaea in the AC than WW where Crenarchaeota, Euryarchaeota, Korarchaeota, Nanoarchaeota and Thaumarchaeota were major phyla with 60 genuses. The nitrogen metabolism was reported in terms of assimilatory nitrate reductase. As the class, Proteobacteria, Actinobacteria were prominent in WW, whereas Proteobacteria, Chloroflexi in AC were abundant. From functional annotation of sequences, the microbial flora in AC has the potential of removal of pollutants present in the form of carbon, nitrogen, and phosphorus.
Graphical abstract










Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and supplementary material provided.
References
AWWA American Public Health Association (1995) Standard methods for the examination of water and wastewater, Standard methods for the examination of water and wastewater. 1–1000
Bai Y, Liang J, Liu R, Hu C, Qu J (2014) Metagenomic analysis reveals microbial diversity and function in the rhizosphere soil of a constructed wetland. Environ Technol 35:2521–2527. https://doi.org/10.1080/09593330.2014.911361
Balcom IN, Driscoll H, Vincent J, Leduc M (2016) Metagenomic analysis of an ecological wastewater treatment plant’s microbial communities and their potential to metabolize pharmaceuticals. F1000Research. https://doi.org/10.12688/f1000research.9157.1
Bhatti AA, Haq S, Bhat RA (2017) Actinomycetes benefaction role in soil and plant health. Microb Pathog 111:458–467. https://doi.org/10.1016/j.micpath.2017.09.036
Central Pollution Control Board (CPCB) (2000) Schedule – V, “General standards for discharge of environmental. Pollutants Part-A : Effluents”, Environmental Standards
Chen Y, Wen Y, Cheng J, Xue C, Yang D, Zhou Q (2011) Effects of dissolved oxygen on extracellular enzymes activities and transformation of carbon sources from plant biomass: implications for denitrification in constructed wetlands. Bioresour Technol 102:2433–2440. https://doi.org/10.1016/j.biortech.2010.10.122
Chiu Y-C, Chung M-S (2003) Determination of optimal COD/nitrate ratio for biological denitrification. Int Biodeterior Biodegrad 51:43–49. https://doi.org/10.1016/S0964-8305(02)00074-4
Delforno T, Lacerda G Jr, Sierra-Garcia I, Okada D, Macedo T, Varesche MBA, Oliveira V (2017) Metagenomic analysis of the microbiome in three different bioreactor configurations applied to commercial laundry wastewater treatment. Sci Total Environ 587:389–398. https://doi.org/10.1016/j.scitotenv.2017.02.170
Deng Q, Wan L, Li X, Cao X, Zhou Y, Song C (2020) Metagenomic evidence reveals denitrifying community diversity rather than abundance drives nitrate removal in stormwater biofilters amended with different organic and inorganic electron donors. Chemosphere 257:134–142. https://doi.org/10.1016/j.chemosphere.2020.127269
Díaz AI, Oulego P, Laca A, González JM, Díaz M (2019) Metagenomic analysis of bacterial communities from a nitrification–denitrification treatment of landfill leachates. Clean-Soil Air Water 47:11–33. https://doi.org/10.1002/clen.201900156
Ding X, Lan W, Wu J, Hong Y, Li Y, Ge Q, Urzi C, Katayama Y, Gu J-D (2020) Correction to: Microbiome and nitrate removal processes by microorganisms on the ancient Preah Vihear temple of Cambodia revealed by metagenomics and N-15 isotope analyses. Appl Microbiol Biotechnol 104:9823–9837. https://doi.org/10.1007/s00253-020-10956-7
Eberly JO, Indest KJ, Hancock DE, Jung CM, Crocker FH (2016) Metagenomic analysis of denitrifying wastewater enrichment cultures able to transform the explosive, 3-nitro-1, 2, 4-triazol-5-one (NTO). J Ind Microbiol Biotechnol 43:795–805. https://doi.org/10.1007/s10295-016-1755-5
Emmanuel SA, Sul WJ, Seong HJ, Rhee C, Ekpheghere KI, Kim I-S, Kim H-G, Koh S-C (2019) Metagenomic analysis of relationships between the denitrification process and carbon metabolism in a bioaugmented full-scale tannery wastewater treatment plant. World J Microbiol Biotechnol 35:1–12. https://doi.org/10.1007/s11274-019-2716-8
Frostegård Ã, Vick SH, Lim NY, Bakken LR, Shapleigh JP (2022) Linking meta-omics to the kinetics of denitrification intermediates reveals pH-dependent causes of N2O emissions and nitrite accumulation in soil. ISME J 16:26–37. https://doi.org/10.1038/s41396-021-01045-2
Gao H, Mao Y, Zhao X, Liu W-T, Zhang T, Wells G (2019) Genome-centric metagenomics resolves microbial diversity and prevalent truncated denitrification pathways in a denitrifying PAO-enriched bioprocess. Water Res 155:275–287. https://doi.org/10.1016/j.watres.2019.02.020
Ginige MP, Bowyer JC, Foley L, Keller J, Yuan Z (2009) A comparative study of methanol as a supplementary carbon source for enhancing denitrification in primary and secondary anoxic zones. Biodegradation 20:221–234. https://doi.org/10.1007/s10532-008-9215-1
Goh H, Zakaria N, Lau T, Foo K, Chang C, Leow C (2017) Mesocosm study of enhanced bioretention media in treating nutrient rich stormwater for mixed development area. Urban Water 14:134–142. https://doi.org/10.1080/1573062X.2015.1076861
Huang Y, Yang H, Li K, Meng Q, Wang S, Wang Y, Zhu P, Niu Q, Yan H, Li X (2022) Red mud conserved compost nitrogen by enhancing nitrogen fixation and inhibiting denitrification revealed via metagenomic analysis. Bioresour Technol. https://doi.org/10.1016/j.biortech.2021.126654
Itoh T (2014) The Family Thermoproteaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The Prokaryotes: other major lineages of Bacteria and the Archaea. Springer, Berlin, Heidelberg, pp 389–401. https://doi.org/10.1007/978-3-642-38954-2_330
Jadeja NB, More RP, Purohit HJ, Kapley A (2014) Metagenomic analysis of oxygenases from activated sludge. Bioresour Technol 165:250–256. https://doi.org/10.1016/j.biortech.2014.02.045
Li W, Zhuang J-l, Zhou Y-y, Meng F-g, Kang D, Zheng P, Shapleigh JP (2020) Metagenomics reveals microbial community differences lead to differential nitrate production in anammox reactors with differing nitrogen loading rates. Water Res 169:1–8. https://doi.org/10.1016/j.watres.2019.115279
Liu S, Chen Y, **ao L (2021) Metagenomic insights into mixotrophic denitrification facilitated nitrogen removal in a full-scale A2/O wastewater treatment plant. PLoS ONE 16:250–283. https://doi.org/10.1371/journal.pone.0250283
Lv X-M, Shao M-F, Li J, Li C-L (2015) Metagenomic analysis of the sludge microbial community in a lab-scale denitrifying phosphorus removal reactor. Appl Biochem Biotechnol 175:3258–3270. https://doi.org/10.1007/s12010-015-1491-8
Ma R-C, Chu Y-X, Wang J, Wang C, Leigh MB, Chen Y, He R (2021) Stable-isotopic and metagenomic analyses reveal metabolic and microbial link of aerobic methane oxidation coupled to denitrification at different O2 levels. Sci Total Environ 764:142901. https://doi.org/10.1016/j.scitotenv.2020.142901
Martin HG, Ivanova N, Kunin V, Warnecke F, Barry KW, McHardy AC, Yeates C, He S, Salamov AA, Szeto E (2006) Metagenomic analysis of two enhanced biological phosphorus removal (EBPR) sludge communities. Nat Biotechnol 24:1263–1269. https://doi.org/10.1038/nbt1247
Massoud MA, Tarhini A, Nasr JA (2009) Decentralized approaches to wastewater treatment and management: applicability in develo** countries. J Environ Manag 90:652–659. https://doi.org/10.1016/j.jenvman.2008.07.001
Meyer F, Paarmann D, D’Souza M, Olson R, Glass EM, Kubal M, Paczian T, Rodriguez A, Stevens R, Wilke A (2008) The metagenomics RAST server–a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinform 9:1–8. https://doi.org/10.1186/1471-2105-9-386
Miao Y, Liao R, Zhang X-X, Liu B, Li Y, Wu B, Li A (2015) Metagenomic insights into salinity effect on diversity and abundance of denitrifying bacteria and genes in an expanded granular sludge bed reactor treating high-nitrate wastewater. Chem Eng J 277:116–123. https://doi.org/10.1016/j.cej.2015.04.125
Nadeau SA, Roco CA, Debenport SJ, Anderson TR, Hofmeister KL, Walter MT, Shapleigh JP (2019) Metagenomic analysis reveals distinct patterns of denitrification gene abundance across soil moisture, nitrate gradients. Environ Microbiol 21:1255–1266. https://doi.org/10.1111/1462-2920.14587
Pan Y, Ni B-J, Bond PL, Ye L, Yuan Z (2013) Electron competition among nitrogen oxides reduction during methanol-utilizing denitrification in wastewater treatment. Water Res 47:3273–3281. https://doi.org/10.1016/j.watres.2013.02.054
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Pessi IS, Viitamäki S, Virkkala A-M, Eronen-Rasimus E, Delmont TO, Marushchak ME, Hultman J (2020) Truncated denitrifiers dominate the denitrification pathway in tundra soil metagenomes. bioRxiv. https://doi.org/10.1101/2020.12.21.419267
Pessi IS, Viitamäki S, Virkkala A-M, Eronen-Rasimus E, Delmont TO, Marushchak ME, Luoto M, Hultman J (2022) In-depth characterization of denitrifier communities across different soil ecosystems in the tundra. bioRxiv. https://doi.org/10.1186/s40793-022-00424-2
Qian W, Ma B, Li X, Zhang Q, Peng Y (2019) Long-term effect of pH on denitrification: high pH benefits achieving partial-denitrification. Bioresour Technol 278:444–449. https://doi.org/10.1016/j.biortech.2019.01.105
Sánchez O (2017) Constructed wetlands revisited: microbial diversity in the–omics era. Microb Ecol 73:722–733. https://doi.org/10.1007/s00248-016-0881-y
Vymazal J (2005) Constructed wetlands for wastewater treatment. Ecol Eng 25:475–477. https://doi.org/10.1016/j.ecoleng.2005.07.002
Wang J, Long Y, Yu G, Wang G, Zhou Z, Li P, Zhang Y, Yang K, Wang S (2022) A review on microorganisms in constructed wetlands for typical pollutant removal: species, function, and diversity. Front Microbiol 13:845725. https://doi.org/10.3389/fmicb.2022.845725
Xu J, Liu X, Huang J, Huang M, Wang T, Bao S, Tang W, Fang T (2020) The contributions and mechanisms of iron-microbes-biochar in constructed wetlands for nitrate removal from low carbon/nitrogen ratio wastewater. RSC Adv 10:23212–23220. https://doi.org/10.1039/D0RA03609A
Xue Z, Wang C, Cao J, Luo J, Feng Q, Fang F, Li C, Zhang Q (2018) An alternative carbon source withdrawn from anaerobic fermentation of soybean wastewater to improve the deep denitrification of tail water. Biochem Eng 132:217–224. https://doi.org/10.1016/j.bej.2018.01.025
Yadav TC, Pal RR, Shastri S, Jadeja NB, Kapley A (2015) Comparative metagenomics demonstrating different degradative capacity of activated biomass treating hydrocarbon contaminated wastewater. Bioresour Technol 188:24–32. https://doi.org/10.1016/j.biortech.2015.01.141
Yang Z, Zhou Q, Sun H, Jia L, Zhao L, Wu W (2021) Metagenomic analyses of microbial structure and metabolic pathway in solid-phase denitrification systems for advanced nitrogen removal of wastewater treatment plant effluent: a pilot-scale study. Water Res 196:1–9. https://doi.org/10.1016/j.watres.2021.117067
Yaragal R, Mutnuri S (2022) Nitrates removal using ion exchange resin: batch, continuous column and pilot-scale studies. Int J Environ Sci Technol 20:1–16. https://doi.org/10.1007/s13762-021-03836-8
Yaragal RR, Kumar D, Mutnuri S (2020) Development of UPLCMS/MS method for analyzing phorate: application to wastewater treatment. J Iran Chem Soc 17:2923–2931. https://doi.org/10.1007/s13738-020-01971-6
Acknowledgements
The authors are thankful to BITS Pilani, K.K. Birla Goa Campus, for providing an opportunity to conduct the experiments and allowed to construct the VFCW and in the sewage treatment plant at the campus.
Funding
The present study is not funded.
Author information
Authors and Affiliations
Contributions
All the authors have made substantial contributions to the present paper. Author SM was involved in the conception and design of the treatment units. Author RY was involved in acquisition of data, analysis and interpretation of data. Both the authors have participated in drafting the article and revising it critically for important intellectual content and have given final approval of the version to be submitted.
Corresponding author
Ethics declarations
Competing interests
All the authors declare that there are no competing interests.
Ethical approval
Not applicable as the present study did not involve any testing/study on human or animal subjects.
Consent to participate
Not applicable.
Consent to Publish
All the authors provide consent be published in the above Journal and Article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yaragal, R.R., Mutnuri, S. Diversity and functional annotation of microorganisms in anaerobic chamber treating nitrate-rich wastewater. World J Microbiol Biotechnol 39, 311 (2023). https://doi.org/10.1007/s11274-023-03750-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03750-w