Abstract
Phenazines are heteroaromatic compounds consisting of a central pyrazine ring fused with two benzenes. Different functional groups attached to the dibenzopyrasin core cause differences in the chemical, physical, and biological properties of phenazines. Interest in these compounds has not diminished for decades. New biological activities and practical applications discovered in recent years force researchers to investigate all aspects of the synthesis, degradation, and mechanisms of action of phenazines. In this study, we have demonstrated the involvement of the coxA gene product (cytochrome c oxidase, su I) in the production of phenazines in P. chlororaphis subsp. aurantiaca. Overlap PCR was used to knock out the coxA gene and the resulting mutants were screened for their ability to grow on rich and minimal culture media and for phenazine production. The reintroduction of the full-length coxA gene into the B-162/coxA strains was used to further confirm the role of this gene product in the ability to produce phenazines. We were able to show that the product of the coxA gene is necessary for phenazine production in rich growth media. At the same time, the CoxA protein does not seem to have any effect on phenazine production in M9 minimal salt medium. We could show that knocking down even one subunit of the cytochrome c oxidase complex leads to a significant reduction (to trace concentrations) or complete suppression of phenazine antibiotic production on rich PCA medium in P. chlororaphis subsp. aurantiaca.
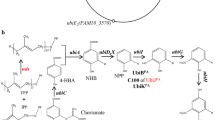
Graphical abstract







Similar content being viewed by others
Data availability
Additional data available on request.
References
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (1993) Current protocols in molecular biology. John Wiley & Sons Inc., New Jersey
Bakker PAHM, Pieterse CMJ, van Loon LC (2007) Induced systemic resistance by fluorescent Pseudomonas spp. Phytopathology 97: 239–243. https://doi.org/10.1094/PHYTO-97-2-0239
Branden G, Gennis RB, Brzezinski P (2006) Transmembrane proton translocation by cytochrome c oxidase. Biochim Biophys Acta 1757(8):1052–1063. https://doi.org/10.1016/j.bbabio.2006.05.020
Bratton M, Millsm D, Castleden CK, Hosler J, Meunier B (2003) Disease-related mutations in cytochrome c oxidase studied in yeast and bacterial models. Eur J Biochem 270(6): 1222–1230. https://doi.org/10.1046/j.1432-1033.2003.03482.x
Calhoun MW, Thomas JW, Gennis RB (1994) The cytochrome oxidase superfamily of redox-driven proton pumps. Trends in Biochem Sci 19(8):325–330. https://doi.org/10.1016/0968-0004(94)90071-X
Chen LH, Tsai HC, Yu PL, Chung KR (2017) A major facilitator superfamily transporter-mediated resistance to oxidative stress and fungicides requires Yap1, Skn7, and MAP kinases in the citrus fungal pathogen Alternaria alternata. PLoS ONE 12(1): e0169103. https://doi.org/10.1371/journal.pone.0169103
Chin-A-Woeng TFC, Bloemberg GV, Lugtenberg BJJ (2003) Phenazines and their role in biocontrol by Pseudomonas bacteria. New Phytol 157(3):503–523. https://doi.org/10.1046/j.1469-8137.2003.00686.x
Costa KC, Bergkessel M, Saunders S, Korlach J, Newman DK (2015) Enzymatic degradation of phenazines can generate energy and protect sensitive organisms from toxicity. mBio 6(6): 1–20. https://doi.org/10.1128/mBio.01520-15
Dahlstrom KM, Newman DK (2021) Soil bacteria protect fungi from phenazines by acting as toxin sponges. Curr Biol 32:1–14. https://doi.org/10.1016/j.cub.2021.11.002
Dar D, Thomashow LS, Weller DM, Newman DK (2020) Global landscape of phenazine biosynthesis and biodegradation reveals species-specific colonization patterns in agricultural soils and crop microbiomes. eLife 9: e59726. https://doi.org/10.7554/elife.59726
French SW, Palmer DS, Sim WA (1973) Phenazine methosulfate uptake fey rat liver mitochondria. Can J Biochem 51(3):235–240. https://doi.org/10.1139/o73-030
Garcia-Horsman JA, Barquera B, Rumbley J, Ma J, Gennis RB (1994) The superfamily of heme-copper respiratory oxidases. J Bacteriol 176(18):5587–5600. https://doi.org/10.1128/jb.176.18.5587-5600.1994
Girard G, van Rij ET, Lugtenberg BJJ, Bloemberg GV (2006) Regulatory roles of psrA and rpoS in phenazine-1-carboxamide synthesis by Pseudomonas chlororaphis PCL1391. Microbiology 152(1): 43–58. https://doi.org/10.1099/mic.0.28284-0
Hmelo L, Borlee B, Almblad H et al (2015) Precision-engineering the Pseudomonas aeruginosa genome with two-step allelic exchange. Nat Protoc 10:1820–1841. https://doi.org/10.1038/nprot.2015.115
Huang H, Sunm L, Bi K, Zhong G, Hu M (2016) The effect of phenazine-1-carboxylic acid on the morphological, physiological, and molecular characteristics of Phellinus noxius. Molecules 21(5): 613–623. https://doi.org/10.3390/molecules21050613
Jo J, Cortez KL, Cornell WC, Price-Whelan A, Dietrich LEP (2017) An orphan cbb3-type cytochrome oxidase subunit supports Pseudomonas aeruginosa biofilm growth and virulence. eLIFE 6: e30205. https://doi.org/10.7554/eLife.30205.001
Kadenbach B (2018) Regulation of mitochondrial respiration and ATP synthesis via cytochrome c oxidase. Rend Fis Acc Lincei 29:421–435. https://doi.org/10.1007/s12210-018-0710-y
Kawakami T, Kuroki M, Ishii M, Igarashi Y, Arai H (2010) Differential expression of multiple terminal oxidases for aerobic respiration in Pseudomonas aeruginosa. Environ Microbiol 12(6):1399–1412. https://doi.org/10.1111/j.1462-2920.2009.02109.x
Laursen JB, Nielsen J (2004) Phenazine natural products: biosynthesis, synthetic analogues and biological activity. Chem Rev 104:1663–1686. https://doi.org/10.1021/cr020473j
Levitch ME, Stadtman ER (1964) A study of the biosynthesis of phenazine-1-carboxylic acid. Arch Biochem Biophys 106:194–199. https://doi.org/10.1016/0003-9861(64)90175-4
Liaudanskaya AI, Vychik PV, Maximova NP, Verameyenka KG (2022) Genome analysis of Pseudomonas chlororaphis subsp. aurantiaca mutant strains with increased production of phenazines. Arch Microbiol 204(247): 1–10. https://doi.org/10.1007/s00203-021-02648-1
Muhl D, Filloux A (2014) Site-directed mutagenesis and gene deletion using reverse genetics. In: Filloux A, Ramos JL (eds) Pseudomonas methods and protocols. Methods in molecular biology, 1149. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-0473-0_40
Osamura T, Kawakami T, Kido R, Ishii M, Arai H (2017) Specific expression and function of the A-type cytochrome c oxidase under starvation conditions in Pseudomonas aeruginosa. PLoS One 12(5): 1–16. https://doi.org/10.1371/journal.pone.0177957
Pereira MM, Santana M, Teixeira M (2001) A novel scenario for the evolution of haem-copper oxygen reductases. Biochim Biophys Acta 1505(2–3):185–208. https://doi.org/10.1016/S0005-2728(01)00169-4
Pierson LS 3rd, Pierson EA (2010) Metabolism and function of phenazines in bacteria: impacts on the behavior of bacteria in the environment and biotechnological processes. Appl Microbiol Biotechnol 86(6): 1659–1670. https://doi.org/10.1007/s00253-010-2509-3
Sakhtah H, Koyama L, Zhang Y, Morales DK, Fields BL, Price-Whelan A, Hogan DA, Shepard K, Dietrich LEP (2016) The Pseudomonas aeruginosa efflux pump MexGHI-OpmD transports a natural phenazine that controls gene expression and biofilm development. Proc Nat Acad Sci 113(25): 3538–3547. https://doi.org/10.1073/pnas.1600424113
Schiessl KT, Hu F, Jo J, Nazia SZ, Wang B, Price-Whelan A, Wei Min W, Dietrich LEP (2019) Phenazine production promotes antibiotic tolerance and metabolic heterogeneity in Pseudomonas aeruginosa biofilms. Nat Commun 10:762–772. https://doi.org/10.1038/s41467-019-08733-w
Shanmugaiah V, Mathivanan N, Varghese B (2010) Purification, crystal structure and antimicrobial activity of phenazine-1-carboxamide produced by a growth-promoting biocontrol bacterium, Pseudomonas aeruginosa MML2212. J Appl Microbiol 108:703–711. https://doi.org/10.1111/j.1365-2672.2009.04466.x
Shapira MA, Verameyenka KG, Liavonchyk KV, Dobysh AA, Yantsevich AV, Maksimova NP (2021) Novel approach of phenazine derivatives isolation from Pseudomonas culture medium. Process Biochem 111:325–331. https://doi.org/10.1016/j.procbio.2021.11.004
Taniguchi M, Lindsey JS (2018) Database of absorption and fluorescence spectra of & 300 common compounds for use in PhotochemCAD. Photochem Photobiol 94:290–327. https://doi.org/10.1111/php.12860
Veremeenko EG, Maksimova NP (2010) Activation of the antioxidant complex in Pseudomonas aurantiaca—producer of phenazine antibiotics. Microbiology 79(4):439–444. https://doi.org/10.1134/S0026261710040041
Wikström M, Krab K, Sharma V (2018) Oxygen activation and energy conservation by cytochrome c oxidase. Chem Rev 118(5):2469–2490. https://doi.org/10.1021/acs.chemrev.7b00664
Yan J, Liu W, Cai J, Wang Y, Li D, Hua H, Cao H (2021) Advances in phenazines over the past decade: review of their pharmacological activities, mechanisms of action, biosynthetic pathways and synthetic strategies. Mar Drugs 19(11):610–638. https://doi.org/10.3390/md19110610
Yu JM, Wang D, Pierson LS 3rd, Pierson EA (2018) Effect of producing different phenazines on bacterial fitness and biological control in Pseudomonas chlororaphis 30–84. Plant Pathol J 34(1): 44–58. https://doi.org/10.5423/PPJ.FT.12.2017.0277
Zhao Y, Liu J, Jiang T (2021) Resistance-nodulation-division efflux pump, lexABC, contributes to self-resistance of the phenazine di-n-oxide natural product myxin in Lysobacter antibioticus. Front Microbiol 12: 1–12. https://doi.org/10.3389/fmicb.2021.618513
Funding
This work was supported by the Belarusian State Program of Scientific Research (project number 20211293).
Author information
Authors and Affiliations
Contributions
Katsiaryna G. Verameyenka was involved in the conception and design of the study. Katsiaryna G. Verameyenka and Volha A. Naumouskaya prepared the material, collected and analysed the data. The first draft of the manuscript was written by Katsiaryna G. Verameyenka and all authors commented on earlier versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
There are no ethical issues relevant to this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.





Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Verameyenka, K.G., Naumouskaya, V.A. & Maximova, N.P. Cytochrome c oxidase is one of the key enzymes providing the ability to produce phenazines in Pseudomonas chlororaphis subsp. aurantiaca. World J Microbiol Biotechnol 39, 279 (2023). https://doi.org/10.1007/s11274-023-03685-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03685-2