Abstract
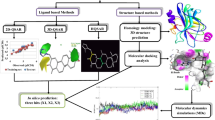
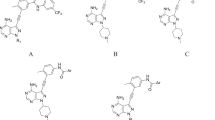
As per the World Health Organization (WHO), cancer is the second most leading cause of death after cardiovascular diseases in worldwide with around 9.88 million total new cases and 1.08 million were observed due to skin cancer in 2018. Amongst two types of skin cancer, progression of melanoma cancer is increasing day by day due to the environmental changes than non-melanoma cancer. Most of B-Raf mutation, specifically B-RafV600E, is responsible for the progression of the melanoma cancer. Here, various 3D-QSAR techniques like comparative molecular field analysis (CoMFA), comparative molecular similarity indices analysis (CoMSIA), molecular hologram QSAR (HQSAR) and topomer CoMFA were used to design novel B-Raf inhibitors by using 28 synthetic B-Raf inhibitors. Except for topomer CoMFA model, remaining models were generated by three different alignment methods in which distil-based alignment method was found best and gave prominent statistical values. After performing N-fold statistical validation, in CoMFA, q2, r2 and r2pred values were found to be 0.638, 0.969 and 0.848, respectively. Similarly, q2, r2 and r2pred values were found to be 0.796, 0.978 and 0.891 in CoMSIA (SHD) and 0.761, 0.973 and 0.852 in CoMSIA (SH) by N-fold statistical validation. In HQSAR analysis, statistical values were found for q2 as 0.984, r2 as 0.999 and r2pred as 0.634 with 97 as best hologram length (BHL). The results of topomer CoMFA showed the q2 value of 0.663 and the r2 value of 0.967. Important features of purinylpyridine were identified by contour map analysis of all 3D-QSAR techniques, which could be useful to design the novel molecules as B-Raf inhibitors for the treatment of melanoma cancer.












Similar content being viewed by others
Abbreviations
- Raf:
-
Rapidly accelerated fibrosarcoma
- RAS:
-
Retrovirus-associated DNA sequences
- MAPK:
-
Mitogen-activated protein kinase
- RTK:
-
Receptor tyrosine kinase
- ERK:
-
Extracellular regulated kinase
- 3D-QSAR:
-
Three-dimensional quantitative structural activity relationship
- CoMFA:
-
Comparative molecular field analysis
- CoMSIA:
-
Comparative molecular similarity indices analysis
- HQSAR:
-
Molecular hologram QSAR
- MD/MS:
-
Molecular docking and molecular simulation
References
http://www.who.int/en/news-room/fact-sheets/detail/cancer (Accessed on 5th October 2018)
List of rare diseases and synonyms (2018) https://www.orpha.net/orphacom/cahiers/docs/GB/List_of_rare_diseases_in_alphabetical_order.pdf. Accessed 5 Oct 2018
Liu L, Xu J, Yang J, Feng C, Miao Y (2017). Bioorg Med Chem Lett 27:4952
Neidle S (2013) Cancer drug design and discovery, 2nd edn. Acedamic Press, London
Rahman MA, Salajegheh A, Smith RA, Lam AKY (2013). Exp Mol Pathol 95:336
Verma J, Khedkar VM, Coutinho EC (2010). Curr Top Med Chem 10:95
Zhang S, Lin Z, Pu Y, Zhang Y, Zhang L, Zuo Z (2017). Comput Biol Chem 67:38
Yu S, Yuan J, Shi J, Ruan X, Zhang T, Wang Y et al (2015). Chemom Intell Lab Syst 146:34
Yang W, Chen Y, Zhou X, Gu Y, Qian W, Zhang F et al (2015). Eur J Med Chem 89:581
Sybyl X (2011) Molecular modelling software. Tripos Certara, V.1.2, St. Louis
Borisa A, Bhatt H (2015). Eur J Pharm Sci 79:1
Wang W, Tian Y, Wan Y, Gu S, Ju X, Luo X, Liu G (2018). Struct Chem 30:385–397
Elham G, Mohammad H (2018). J Chin Chem Soc 65:1
Jianbo T, Pei Z, **ang S, Wang W (2017). J Chemom 31:2934
Wold S, Ruhe A, Wold H, Dunn III WJ (1984). J Sci Stat Comput 5:735
Zambre V, Murumkar P, Giridhar R, Yadav M (2010). J. Mol. Graph. Model 29:229
Tanga H, Yanga L, Li J, Chen J (2016). J Taiwan Inst Chem Eng 68:64–73
Clark M, Cramer RD., (1993), Quant. Struct. Relationships 12:137.
Patel P, Bhatt H (2016). Bioorg Med Chem Lett 28:2328
Golbraikh A, Tropsha A (2002). J Mol Graph Model 20:269
Markus Böhm, Jörg Stürzebecher and, Gerhard Klebe (1999),. J Med Chem42:458.
Mohammed AA, Janarthanan T, Naga ST (2017). Struct Chem 28:1187–1200
Chaube U, Bhatt H (2017). Mol Divers 21:741
Chaube U, Chhatbar D, Bhatt H (2016). Bioorg Med Chem Lett 26:864
Ballu S, Itteboina R, Sivan SK, Manga V (2017). Struct Chem 29:593–605
Acknowledgements
The authors are thankful to Nirma University, Ahmedabad, India, for providing the needful facilities to carry out research work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 241 kb)
Rights and permissions
About this article
Cite this article
Chavda, J., Bhatt, H. 3D-QSAR (CoMFA, CoMSIA, HQSAR and topomer CoMFA), MD simulations and molecular docking studies on purinylpyridine derivatives as B-Raf inhibitors for the treatment of melanoma cancer. Struct Chem 30, 2093–2107 (2019). https://doi.org/10.1007/s11224-019-01334-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-019-01334-9