Abstract
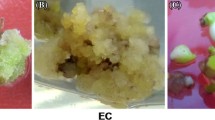
Carbon nanomaterials (CNMs) have attracted a great deal of research interest for their potential application in plants. Some types of CNMs have exhibited the ability on regulating plant growth of plants, which showed a promising future in agriculture. However, detailed mechanism of their regulation on plant growth is still not well characterized especially at the molecular level. In this study, our objective is to study the regulation mechanism of carbon nanoparticles (CNPs) on tobacco callus growth at protein level. During the incubation period, proteomic profiling of tobacco callus was investigated by using the isobaric tags for relative and absolute quantitation labelling (iTRAQ) coupled with 2D-LC MS/MS. The function of differentially expressed proteins was achieved by using a gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) analysis with InterproScan, and qRT-PCR was used to support the results of the proteomic data. The results showed that the growth of tobacco callus was enhanced by CNPs, with the optimal concentration at 50 mg/L. The expression levels of 198 cellular proteins were significantly up- or down-regulated after CNPs treatment. Some differentially expressed proteins were located in mitochondria and involved in calcium-mediated signaling pathway. The proteomic profile changes provided more deeper understanding of the highly complex regulatory mechanisms in tobacco callus exposed to CNPs.





Similar content being viewed by others
Abbreviations
- CNMs:
-
Carbon nanomaterials
- CNPs:
-
Carbon nanoparticles
- GO:
-
Gene ontology
- iTRAQ:
-
Isobaric tag for relative and absolute quantitation
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- TBY-2 cell:
-
Nicotiana tabacum L. Bright Yellow-2 cell
- TEM:
-
Transmission electron microscopy
References
Alex M, Chang HY, Louise D, Matthew F, Sarah H, Rodrigo L, Craig MA, Conor MM, Gift N, Sebastien P (2015) The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res 43:213–221
Ali MH, Sobze JM, Pham TH, Nadeem M, Liu C, Galagedara L, Cheema M, Thomas R (2020) Carbon nanoparticles functionalized with carboxylic acid improved the germination and seedling vigor in upland boreal forest species. Nanomaterials 10:176
Anderson NL, Anderson NG (1998) Proteome and proteomics: new technologies, new concepts, and new words. Electrophoresis 19:1853–1861
Cañas JE, Long M, Nations S, Vadan R, Dai L, Luo M, Ambikapathi R, Lee EH, Olszyk D (2008) Effects of functionalized and nonfunctionalized single-walled carbon nanotubes on root elongation of select crop species. Environ Toxicol Chem 27:1922–1931
Chen L, Wang H, Li X, Nie C, Liang T, **e F, Liu K, Peng X, **e J (2018) Highly hydrophilic carbon nanoparticles: uptake mechanism by mammalian and plant cells. RSC Adv 8:35246–35256
Chen L, Yang J, Li X, Liang T, Nie C, **e F, Liu K, Peng X, **e J (2020) Carbon nanoparticles enhance potassium uptake via upregulating potassium channel expression and imitating biological ion channels in BY-2 cells. Journal of Nanobiotechnology 18:21
Deng Y, Zou W, Li G, Zhao J (2014) TRANSLOCASE OF THE INNER MEMBRANE9 and 10 are essential for maintaining mitochondrial function during early embryo cell and endosperm free nucleus divisions in Arabidopsis. Plant Physiol 166:853–868
Elliott MH, Smith DS, Parker CE, Borchers C (2009) Current trends in quantitative proteomics. J Mass Spectrom 44:1637–1660
Feihong L, Chao S, Xuehan L, **nyi Y, Chao L, Yanying S, Shenchun Q (2018) The effect of graphene oxide on adventitious root formation and growth in apple. Plant Physiol Biochem 129:122–129
Hamasaki H, Yoshizumi T, Takahashi N, Higuchi M, Kuromori T, Imura Y, Shimada H, Matsui M (2012) SD3, an Arabidopsis thaliana homolog of TIM21, affects intracellular ATP levels and seedling development. Mol Plant 5:461–471
Jian Y, Taibo L, Haijiang L, Qisheng Y, Yanling Z, Han** Z, Shixiang Z (2015) Effects of nano-carbon sol on physiological characteristics of root system and potassium absorption of flue-cured tobacco. Tobacco Science & Technology 48:7–11
Khodakovskaya MV, Silva KD, Biris AS, Dervishi E, Villagarcia H (2015) Carbon nanotubes induce growth enhancement of tobacco cells. ACS Nano 6:2128–2135
Kim TY, Lee SH, Ku H, Lee SY (2019) Enhancement of drought tolerance in cucumber plants by natural carbon materials. Plants (basel, Switzerland) 8:446
Kole C, Kole P, Randunu KM, Choudhary P, Podila R, Pu CK, Rao AM, Marcus RK (2013) Nanobiotechnology can boost crop production and quality: first evidence from increased plant biomass, fruit yield and phytomedicine content in bitter melon ( Momordica charantia ). BMC Biotechnol 13:37
Kumar S, Yoshizumi T, Hongo H, Yoneda A, Hara H, Hamasaki H, Takahashi N, Nagata N, Shimada H, Matsui M (2012) Arabidopsis mitochondrial protein TIM50 affects hypocotyl cell elongation through intracellular ATP level. Plant Sci 183:212–217
Lahiani MH, Chen J, Irin F, Puretzky AA, Green MJ, Khodakovskaya MV (2015) Interaction of carbon nanohorns with plants: uptake and biological effects. Carbon 81:607–619
Lahiani MH, Dervishi E, Chen J, Nima Z, Gaume A, Biris AS, Khodakovskaya MV (2013) Impact of carbon nanotube exposure to seeds of valuable crops. ACS Appl Mater Interfaces 5:7965–7973
Lahiani MH, Dervishi E, Ivanov I, Chen J, Khodakovskaya M (2016) Comparative study of plant responses to carbon-based nanomaterials with different morphologies. Nanotechnology 27:265102
Lijuan Z, Li L, Aodi W, Huiling Z, Min H, Honghong W, Baoshan X, Zhenyu W, Rong J (2020) Nano-biotechnology in agriculture: use of nanomaterials to promote plant growth and stress tolerance. J Agric Food Chem 68:1935–1947
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408
Martínez-Esteso MJ, Casado-Vela J, Sellés-Marchart S, Pedreño MA, Bru-Martínez R (2014) Differential plant proteome analysis by isobaric tags for relative and absolute quantitation (iTRAQ). Methods Mol Biol 1072:155–169
Mukherjee A, Majumdar S, Servin AD, Pagano L, Dhankher OP, White JC (2016) Carbon nanomaterials in agriculture: a critical review. Front Plant Sci 7:172
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Pina C, Pinto F, Feijó JA, Becker JD (2005) Gene family analysis of the Arabidopsis pollen transcriptome reveals biological implications for cell growth, division control, and gene expression regulation. Plant Physiol 138:744–756
Rico CM, Majumdar S, Duarte-Gardea M, Peralta-Videa JR, Gardea-Torresdey JL (2011) Interaction of nanoparticles with edible plants and their possible implications in the food chain. J Agric Food Chem 59:3485–3498
Saxena M, Maity S, Sarkar S (2014) Carbon nanoparticles in ‘biochar’ boost wheat (Triticum aestivum) plant growth. RSC Adv 4:39948–39954
Schwarzländer M, Fuchs P (2017) Plant mitochondrial membranes: adding structure and new functions to respiratory physiology. Curr Opin Plant Biol 40:147–157
Team CR (2013) Team RDC.R: a language and environment for statistical computing. R Foundation for Statistical Computing: Vienna. Austria Computing 14:12–21
Tiwari DK, Dasgupta-Schubert N, Villaseñor Cendejas LM, Villegas J, Carreto Montoya L, Borjas García SE (2013) Interfacing carbon nanotubes (CNT) with plants: enhancement of growth, water and ionic nutrient uptake in maize (Zea mays) and implications for nanoagriculture. Appl Nanosci 4:577–591
Uniprot consortium (2015) UniProt: a hub for protein information. Nucleic Acids Res 43:204–212
Verma SK, Das AK, Gantait S, Kumar V, Gurel E (2019) Applications of carbon nanomaterials in the plant system: a perspective view on the pros and cons. Sci Total Environ 667:485–499
Wang X, Han H, Liu X, Gu X, Chen K, Lu D (2012) Multi-walled carbon nanotubes can enhance root elongation of wheat ( Triticum aestivum ) plants. J Nanopart Res 14:841–851
Wiśniewski JR, Zougman A, Nagaraj N, Mann M (2009) Universal sample preparation method for proteome analysis. Nat Methods 6:359–362
Yan S, Zhao L, Li H, Zhang Q, Tan J, Huang M, He S, Li L (2013) Single-walled carbon nanotubes selectively influence maize root tissue development accompanied by the change in the related gene expression. J Hazard Mater 246–247:110–118
Yang T, Poovaiah BW (2000) Molecular and biochemical evidence for the involvement of calcium/calmodulin in auxin action. J Biol Chem 275:3137–3143
Yuan H, Hu S, Huang P, Song H, Wang K, **g R, He R, Cui D (2011) Single walled carbon nanotubes exhibit dual-phase regulation to exposed Arabidopsis mesophyll cells. Nanoscale Res Lett 6:44
Yuan J, Gao H, Sui J, Duan H, Chen WN, Ching CB (2012) Cytotoxicity evaluation of oxidized single-walled carbon nanotubes and graphene oxide on human hepatoma HepG2 cells: an iTRAQ-coupled 2D LC-MS/MS proteome analysis. Toxicol Sci 126:149–161
Yuan Z, Zhang Z, Wang X, Li L, Cai K, Han H (2017) Novel impacts of functionalized multi-walled carbon nanotubes in plants: promotion of nodulation and nitrogenase activity in the rhizobium-legume system. Nanoscale 9:9921–9937
Zhang BT, Zheng X, Li HF, Lin JM (2013) Application of carbon-based nanomaterials in sample preparation: a review. Anal Chim Acta 784:1–17
Zhang H, Yue M, Zheng X, **e C, Zhou H, Li L (2017) Physiological effects of single- and multi-walled carbon nanotubes on rice seedlings. IEEE Trans Nanobioscience 16:563–570
Funding
The work described in this paper was substantially supported by Zhengzhou Tobacco Research Institute Dean Fund (112017CA0090) and Science and Technology Project of China Tobacco Corporation (110201902004).
Author information
Authors and Affiliations
Contributions
QY and TL conceived and designed the experiments. ZZ, QC, and XM performed the experiments. ZZ, LH, and HD participated in the data analysis. ZZ and TL drafted the manuscript. TL provided guidance for preparing the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
• CNPs induced growth enhancement of tobacco callus. Proteins regulated by CNPs were identified by iTRAQ-based quantitative proteomics, and the expression levels of 198 cellular proteins were significantly up- or down-regulated. The differentially expressed proteins were located in mitochondria and involved in calcium-mediated signaling pathway.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, Z., Hu, L., Chen, Q. et al. iTRAQ-Based Comparative Proteomic Analysis Provides Insights into Tobacco Callus Response to Carbon Nanoparticles. Plant Mol Biol Rep 40, 556–565 (2022). https://doi.org/10.1007/s11105-022-01337-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-022-01337-7