Abstract
Background
Under saline conditions, Suaeda salsa, as a typical halophyte, accumulates large amounts of Na+ in its leaves during optimal growth. Key transporters involved in Na+ accumulation in plants are HKT-type protein, the plasma membrane Na+/H+ transporter SOS1, and the tonoplast Na+/H+ antiporter NHX1. In this study, the function of SsHKT1;1 and its coordinate expression with SsSOS1 and SsNHX1 to regulate Na+ homeostasis in S. salsa was investigated.
Results
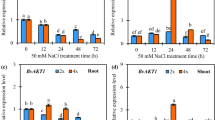
We showed, by yeast complementation assays, that SsHKT1;1 encoded a Na+-selective transporter, which located to the plasma membrane and was preferentially expressed within the stele, and was particularly abundant in xylem parenchyma and pericycle cells. When compared with a treatment of 25 mM NaCl, 150 mM NaCl greatly decreased the transcripts of SsHKT1;1, but maintained a relatively constant level of the expression of SsSOS1 in roots. Consequently, the synergistic effect of SsHKT1;1 and SsSOS1 would result in greater Na+ loading into the xylem under 150 mM NaCl than 25 mM NaCl. In leaves, 150 mM NaCl up-regulated the abundance of SsNHX1 compared with levels in 25 mM NaCl. This enabled the permanent sequestering of Na+ into leaf vacuoles.
Conclusions
Overall, SsHKT1;1 functioned in reducing Na+ retrieval from the root xylem, and played an important role in coordinating with SsSOS1 and SsNHX1 to maintain Na+ accumulation in S. salsa under saline conditions.










Similar content being viewed by others
References
Agarwal PK, Shukla PS, Gupta K, Jha B (2013) Bioengineering for salinity tolerance in plants: state of the art. Mol Biotechnol 54:102–123
Ali A, Park HC, Aman R, Ali Z, Yun DJ (2013) Role of HKT1 in Thellungiella salsuginea, a model extremophile plant. Plant Signal Behav 8:e25196. https://doi.org/10.4161/psb.25196
Ali A, Raddatz N, Aman R, Kim S, Park HC, Jan M, Baek D, Khan IU, Oh DH, Lee SY, Bressan RA, Lee KW, Maggio A, Pardo JM, Bohnert HJ, Yun DJ (2016) A single amino acid substitution in the sodium transporter HKT1 associated with plant salt tolerance. Plant Physiol 171:2112–2126
An D, Chen JG, Gao YQ, Li X, Chao ZF, Chen ZR, Li QQ, Han ML, Wang YL, Wang YF, Chao DY (2017) AtHKT1 drives adaptation of Arabidopsis thaliana to salinity by reducing floral sodium content. PLoS Genet 13:e1007086. https://doi.org/10.1371/journal.pgen.1007086
Anderson JA, Huprikar SS, Kochian LV, Lucas WJ, Gaber RF (1992) Functional expression of a probable Arabidopsis thaliana potassium channel in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 89:3736–3740
Apse MP, Blumwald E (2007) Na+ transport in plants. FEBS Lett 581:2247–2254
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285:1256–1258
Assaha DVM, Ueda A, Saneoka H, Al-Yahyai R, Yaish MW (2017) The role of Na+ and K+ transporters in salt stress adaptation in glycophytes. Front Physiol 8:509
Böhm J, Scherzer S, Shabala S, Krol E, Neher E, Mueller TD, Hedrich R (2016) Venus flytrap HKT1-type channel provides for prey sodium uptake into carnivorous plant without conflicting with electrical excitability. Mol Plant 9:428–436
Brini F, Hanin M, Mezghani I, Berkowitz GA, Masmoudi K (2007) Overexpression of wheat Na+/H+ antiporter TNHX1 and H+-pyrophosphatase TVP1 improve salt- and drought-stress tolerance in Arabidopsis thaliana plants. J Exp Bot 58:301–308
Byrt CS, Xu B, Krishnan M, Lightfoot DJ, Athman A, Jacobs AK, Watson-Haigh NS, Plett D, Munns R, Tester M, Gilliham M (2014) The Na+ transporter, TaHKT1;5-D, limits shoot Na+ accumulation in bread wheat. Plant J 80:516–526
Cao Y, ** X, Huang H, Derebe MG, Levin EJ, Kabaleeswaran V, Pan Y, Punta M, Love J, Weng J, Quick M, Ye S, Kloss B, Bruni R, Martinez-Hackert E, Hendrickson WA, Rost B, Javitch JA, Rajashankar KR, Jiang Y, Zhou M (2011) Crystal structure of a potassium ion transporter TrkH. Nature 471:336–340
Cao Y, Liang X, Yin P, Zhang M, Jiang C (2019) A domestication-associated reduction in K+-preferring HKT transporter activity underlies maize shoot K+ accumulation and salt tolerance. New Phytol 222:301–317
Chen DC, Yang BC, Kuo TT (1992) One-step transformation of yeast in stationary phase. Curr Genet 21:83–84
Chen ZC, Yamaji N, Kashino-Fujii M, Ma JF (2016) A cation-chloride cotransporter gene is required for cell elongation and osmoregulation in rice. Plant Physiol 171:494–507
Colmenero-Flores JM, Martinez G, Gamba G, Vazquez N, Iglesias DJ, Brumos J, Talon M (2007) Identification and functional characterization of cation-chloride cotransporters in plants. Plant J 50:278–292
Davenport RJ, Muñoz-Mayor A, Jha D, Essah PA, Rus A, Tester M (2007) The Na+ transporter AtHKT1;1 controls retrieval of Na+ from the xylem in Arabidopsis. Plant Cell Environ 30:497–507
Deinlein U, Stephan AB, Horie T, Luo W, Xu GH, Schroeder JI (2014) Plant salt-tolerance mechanisms. Trends Plant Sci 19:371–379
Duan HR, Ma Q, Zhang JL, Hu J, Bao AK, Wei L, Wang Q, Luan S, Wang SM (2015) The inward-rectifying K+ channel SsAKT1 is a candidate involved in K+ uptake in the halophyte Suaeda salsa under saline condition. Plant Soil 395:173–187
Flowers TJ, Galal HK, Bromham L (2010) Evolution of halophytes: multiple origins of salt tolerance in land plants. Funct Plant Biol 37:604–612
Hamamoto S, Horie T, Hauser F, Deinlein U, Schroeder JI, Uozumi N (2015) HKT transporters mediate salt stress resistance in plants: from structure and function to the field. Curr Opin Biotechnol 32:113–120
Hauser F, Horie T (2010) A conserved primary salt tolerance mechanism mediated by HKT transporters: a mechanism for sodium exclusion and maintenance of high K+/Na+ ratio in leaves during salinity stress. Plant Cell Environ 33:552–565
Henderson SW, Wege S, Gilliham M (2018) Plant cation-chloride cotransporters (CCC): evolutionary origins and functional insights. Int J Mol Sci 19:492
Horie T, Hauser F, Schroeder JI (2009) HKT transporter-mediated salinity resistance mechanisms in Arabidopsis and monocot crop plants. Trends Plant Sci 14:660–668
Huang S, Spielmeyer W, Lagudah ES, Munns R (2008) Comparative map** of HKT genes in wheat, barley, and rice, key determinants of Na+ transport, and salt tolerance. J Exp Bot 59:927–937
Ismail AM, Horie T (2017) Genomics, physiology, and molecular breeding approaches for improving salt tolerance. Annu Rev Plant Biol 68:405–434
James RA, Blake C, Zwart AB, Hare RA, Rathjen AJ, Munns R (2012) Impact of ancestral wheat sodium exclusion genes Nax1 and Nax2 on grain yield of durum wheat on saline soils. Funct Plant Biol 39:609–618
Jha RK, Patel J, Mishra A, Jha B (2019) Introgression of halophytic salt stress-responsive genes for develo** stress tolerance in crop plants. In: Hasanuzzaman M, Shabala S, Fujita M (eds) Halophytes and climate change: adaptive mechanisms and potential uses. CABI, Boston, pp 275–286
Julkowska MM, Testerink C (2015) Tuning plant signaling and growth to survive salt. Trends Plant Sci 20:586–594
Kobayashi NI, Yamaji N, Yamamoto H, Okubo K, Ueno H, Costa A, Tanoi K, Matsumura H, Fujii-Kashino M, Horiuchi T, Nayef MA, Shabala S, An G, Ma JF, Horie T (2017) OsHKT1;5 mediates Na+ exclusion in the vasculature to protect leaf blades and reproductive tissues from salt toxicity in rice. Plant J 91:657–670
Kong XQ, Gao XH, Sun W, AnJ ZYX, Zhang (2011) Cloning and functional characterization of a cation-chloride cotransporter gene OsCCC1. Plant Mol Biol 75:567–578
Kronzucker HJ, Britto DT (2011) Sodium transport in plants: a critical review. New Phytol 189:54–81
Li W, Zhang Q, Kong X, Wu C, Ma X, Zhang H, Zhao YX (2009) Salt tolerance is conferred in Arabidopsis by overexpression of the vacuolar Na+/H+ antiporter gene SsNHX2, an alternative splicing variant of SsNHX1, from Suaeda salsa. J Plant Biol 52:147–153
Liu Q, Liu R, Ma Y, Song J (2018) Physiological and molecular evidence for Na+ and cl− exclusion in the roots of two Suaeda salsa populations. Aquat Bot 146:1–7
Ma XL, Zhang Q, Shi HZ, Zhu JK, Zhao YX, Ma CL, Zhang H (2004) Molecular cloning and different expression of a vacuolar Na+/H+ antiporter gene in Suaeda salsa under salt stress. Biol Plant 48:219–225
Ma Q, Li YX, Yuan HJ, Hu J, Wei L, Bao AK, Zhang JL, Wang SM (2014) ZxSOS1 is essential for long-distance transport and spatial distribution of Na+ and K+ in the xerophyte Zygophyllum xanthoxylum. Plant Soil 374:661–676
Ma Q, Hu J, Zhou XR, Yuan HJ, Kumar T, Luan S, Wang SM (2017) ZxAKT1 is essential for K+ uptake and K+/Na+ homeostasis in the succulent xerophyte Zygophyllum xanthoxylum. Plant J 90:48–60
Mahi HE, Pérez-Hormaeche J, Luca AD et al (2019) A critical role of sodium flux via the plasma membrane Na+/H+ exchanger SOS1 in the salt tolerance of rice. Plant Physiol 180:1046–1065
Martínez-Atienza J, Jiang X, Garciadeblas B, Mendoza I, Zhu JK, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012
Mäser P, Hosoo Y, Goshima S, Horie T, Eckelman B, Yamada K, Schroeder JI (2002) Glycine residues in potassium channel-like selectivity filters determine potassium selectivity in four-loop-per-subunit HKT transporters from plants. Proc Natl Acad Sci U S A 99:6428–6433
Mbarki S, Sytar O, Cerda A, Zivcak M, Rastogi A, He X, Zoghlami A, Abdelly C, Brestic M (2018) Strategies to mitigate the salt stress effects on photosynthetic apparatus and productivity of crop plants. In: Kumar V et al (eds) Salinity responses and tolerance in plants. Springer, Switzerland, pp 85–88
Mishra A, Tanna B (2017) Halophytes: potential resources for salt stress tolerance genes and promoters. Front Plant Sci 8:829
Møller IS, Gilliham M, Jha D, Mayo GM, Roy SJ, Coates JC, Haseloff J, Tester M (2009) Shoot Na+ exclusion and increased salinity tolerance engineered by cell type-specific alteration of Na+ transport in Arabidopsis. Plant Cell 21:2163–2178
Mumberg D, Müller R, Funk M (1995) Yeast vectors for the controlled expression of heterologous proteins in different genetic backgrounds. Gene 156:119–122
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, Byrt CS, Hare RA, Tyerman SD, Tester M, Plett D, Gilliham M (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30:360–364
Oh DH, Leidi E, Zhang Q et al (2009) Loss of halophytism by interference with SOS1 expression. Plant Physiol 151:210–222
Olías R, Eljakaoui Z, Li J, De Morales PA, Marín-Manzano MC, Pardo JM, Belver A (2009) The plasma membrane Na+/H+ antiporter SOS1 is essential for salt tolerance in tomato and affects the partitioning of Na+ between plant organs. Plant Cell Environ 32:904–916
Pardo JM, Cubero B, Leidi EO, Quintero FJ (2006) Alkali cation exchangers: roles in cellular homeostasis and stress tolerance. J Exp Bot 57:1181–1199
Platten JD, Cotsaftis O, Berthomieu P, Bohnert H, Davenport RJ, Fairbairn DJ, Horie T, Leigh RA, Lin HX, Luan S, Mäser P, Pantoja O, Rodríguez-Navarro A, Schachtman DP, Schroeder JI, Sentenac H, Uozumi N, Véry AA, Zhu JK, Dennis ES, Tester M (2006) Nomenclature for HKT transporters, key determinants of plant salinity tolerance. Trends Plant Sci 11:372–374
Pyo YJ, Gierth M, Schroeder JI, Cho MH (2010) High-affinity K+ transport in Arabidopsis: AtHAK5 and AKT1 are vital for seedling establishment and postgermination growth under low-potassium conditions. Plant Physiol 153:863–875
Qi Z, Spalding EP (2004) Protection of plasma membrane K+ transport by the salt overlay sensitive Na+/H+ antiporter during salinity stress. Plant Physiol 136:2548–2555
Quintero FJ, Garciadeblas B, Rodríguez-Navarro A (1996) The SAL1 gene of Arabidopsis, encoding an enzyme with 3′(2′), 5′-bisphosphate nucleotidase and inositol polyphosphate 1-phosphatase activities, increases salt tolerance in yeast. Plant Cell 8:529–537
Reguera M, Bassil E, Blumwald E (2014) Intracellular NHX-type cation/H+ antiporters in plants. Mol Plant 7:261–263
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146
Shabala S (2013) Learning from halophytes: physiological basis and strategies to improve abiotic stress tolerance in crops. Ann Bot 112:1209–1221
Shi H, Ishitani M, Kim C, Zhu JK (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci U S A 97:6896–6901
Shi H, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477
Song J, Wang B (2015) Using euhalophytes to understand salt tolerance and to develop saline agriculture: Suaeda salsa as a promising model. Ann Bot 115:541–553
Sunarpi HT, Motoda J et al (2005) Enhanced salt tolerance mediated by AtHKT1 transporter-induced Na+ unloading from xylem vessels to xylem parenchyma cells. Plant J 44:928–938
Takahashi R, Liu SK, Takano T (2009) Isolation and characterization of plasma membrane Na+/H+ antiporter genes from salt-sensitive and salt-tolerant reed plants. J Plant Physiol 166:301–309
Tang X, Mu X, Shao H, Wang H, Brestic M (2015) Global plant-responding mechanisms to salt stress: physiological and molecular levels and implications in biotechnology. Crit Rev Biotechnol 35:425–437
Uozumi N, Kim EJ, Rubio F, Yamaguchi T, Muto S, Tsuboi A, Bakker EP, Nakamura T, Schroeder JI (2000) The Arabidopsis HKT1 gene homolog mediates inward Na+ currents in Xenopus laevis oocytes and Na+ uptake in Saccharomyces cerevisiae. Plant Physiol 122:1249–1260
Vieira-Pires RS, Szollosi A, Morais-Cabral JH (2013) The structure of the KtrAB potassium transporter. Nature 496:323–328
Wang BS, Lüttge U, Ratajczak R (2001) Effects of salt treatment and osmotic stress on V-ATPase and V-PPase in leaves of the halophyte Suaeda salsa. J Exp Bot 52:2355–2365
Wang SM, Zhao GQ, Gao YS, Tang ZC, Zhang CL (2004) Puccinellia tenuiflora exhibits stronger selectivity for K+ over Na+ than wheat. J Plant Nutr 27:1841–1857
Wang B, Davenport RJ, Volkov V, Amtmann A (2006) Low unidirectional sodium influx into root cells restricts net sodium accumulation in Thellungiella halophila, a salt-tolerant relative of Arabidopsis thaliana. J Exp Bot 57:1161–1170
Wang SM, Zhang JL, Flowers TJ (2007) Low-affinity Na+ uptake in the halophyte Suaeda maritima. Plant Physiol 145:559–571
Wang CM, Zhang JL, Liu XS, Li Z, Wu GQ, Cai JY, Flowers TJ, Wang SM (2009) Puccinellia tenuiflora maintains a low Na+ level under salinity by limiting unidirectional Na+ influx resulting in a high selectivity for K+ over Na+. Plant Cell Environ 32:486–496
Wang SY, Ma Q, Wang SM (2012) Cloning and sequence analysis of a plasma membrane Na+/H+ antiporter fragment from halophyte Suaeda salsa. Pratacultural Science 29:918–923
Wang TT, Ren ZJ, Liu ZQ, Feng X, Guo RQ, Li BG, Li LG, **g HC (2014) SbHKT1;4, a member of the high-affinity potassium transporter gene family from Sorghum bicolor, functions to maintain optimal Na+/K+ balance under Na+ stress. J Integr Plant Biol 56:315–332
Waters S, Gilliham M, Hrmova M (2013) Plant high-affinity potassium (HKT) transporters involved in salinity tolerance: structural insights to probe differences in ion selectivity. Int J Mol Sci 14:7660–7680
Xu H, Jiang X, Zhan K, Cheng X, Chen X, Pardo JM, Cui D (2008) Functional characterization of a wheat plasma membrane Na+/H+ antiporter in yeast. Arch Biochem Biophys 473:8–15
Yamaguchi T, Hamamoto S, Uozumi N (2013) Sodium transport system in plant cells. Front Plant Sci 4:410
Yang CW, Zheng SS, Huang HL, Liu ZX, Zheng W, Liu B, Shi DC (2012) Comparison of osmotic adjustment and ion balance strategies in nineteen alkali-tolerant halophyte species during adaptation to salt-alkalinized habitats in Northeast China. Aust J Crop Sci 6:141
Yeo AR, Flowers TJ (1986) Ion transport in Suaeda maritima: its relation to growth and implications for the pathway of radial transport of ions across the root. J Exp Bot 37:143–159
Yuan HJ, Ma Q, Wu GQ, Wang P, Hu J, Wang SM (2015) ZxNHX controls Na+ and K+ homeostasis at the whole-plant level in Zygophyllum xanthoxylum through feedback regulation of the expression of genes involved in their transport. Ann Bot 115:495–507
Yue LJ, Li XS, Ma Q, Zhou XR, Wu GQ, Bao AK, Wang SM (2012) NaCl stimulates growth and alleviates water stress in the xerophyte Zygophyllum xanthoxylum. J Arid Environ 87:153–160
Zhang WD, Wang P, Bao Z, Ma Q, Duan LJ, Bao AK, Zhang JL, Wang SM (2017) SOS1, HKT1;5, and NHX1 synergistically modulate Na+ homeostasis in the halophytic grass Puccinellia tenuiflora. Front Plant Sci 8:576
Zhang M, Cao Y, Wang Z, Wang ZQ, Shi J, Liang X, Song W, Chen Q, Lai J, Jiang C (2018) A retrotransposon in an HKT1 family sodium transporter causes variation of leaf Na+ exclusion and salt tolerance in maize. New Phytol 217:1161–1176
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324
Zhu M, Shabala L, Cuin TA, Huang X, Zhou M, Munns R, Shabala S (2015) Nax loci affect SOS1-like Na+/H+ exchanger expression and activity in wheat. J Exp Bot 67:835–844
Zhu M, Zhou M, Shabala L, Shabala S (2017) Physiological and molecular mechanisms mediating xylem Na+ loading in barley in the context of salinity stress tolerance. Plant Cell Environ 40:1009–1020
Acknowledgements
We thank Prof. Caifu Jiang (China Agricultural University) for technical guidance on in situ PCR. This work was supported by the National Natural Science Foundation of China (31730093) and the Fundamental Research Funds for the Central Universities (lzujbky-2018-k01).
Funding
National Natural Science Foundation of China, Grant/Award Number: 31730093; the Fundamental Research Funds for the Central Universities, Grant/Award Number: lzujbky-2018-k01.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOC 42.0 kb)
Rights and permissions
About this article
Cite this article
Wang, WY., Liu, YQ., Duan, HR. et al. SsHKT1;1 is coordinated with SsSOS1 and SsNHX1 to regulate Na+ homeostasis in Suaeda salsa under saline conditions. Plant Soil 449, 117–131 (2020). https://doi.org/10.1007/s11104-020-04463-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-020-04463-x