Abstract
Quantitative structure–activity relationship (QSAR) and read-across techniques have recently been merged into a new emerging field of read-across structure–activity relationship (RASAR) that uses the chemical similarity concepts of read-across (an unsupervised step) and finally develops a supervised learning model (like QSAR). The RASAR method has so far been used only in case of graded predictions or classification modeling. In this work, we attempt, for the first time, to apply RASAR for quantitative predictions (q-RASAR) using a case study of androgen receptor binding affinity data. We have computed a number of error-based and similarity-based measures such as weighted standard deviation of the predicted values, coefficient of variation of the computed predictions, average similarity level of close training compounds for each query molecule, standard deviation and coefficient of variation of similarity levels, maximum similarity levels to positive and negative close training compounds, a concordance measure indicating similarity to positive, negative or both classes of close training compounds, etc. We have clubbed these additional measures along with the selected chemical descriptors from the previously developed QSAR model and redeveloped new partial least squares models from the training set, and predicted the endpoint using the query data set. Interestingly, these new models outperform the internal and external validation quality of the original QSAR model. In this study, we have also introduced a new similarity-based concordance measure (Banerjee-Roy coefficient) that can significantly contribute to the model quality. A q-RASAR model also has the advantage over read-across predictions in providing easy interpretation and indicating quantitative contributions of important chemical features. The strategy described here should be applicable to other biological/toxicological/property data modeling for enhanced quality of predictions, easy interpretability, and efficient transferability.
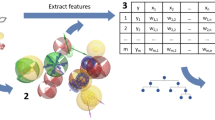
Graphical abstract










Similar content being viewed by others
Data and software availability
The data set was originally collected from the Endocrine Disruptor Knowledge Base (EDKB) database (https://www.fda.gov/science-research/bioinformatics-tools/endocrine-disruptor-knowledge-base) and is available in Supplementary Information in the Excel format. The DTC Laboratory tools used in this study are available free of charge from https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/home and http://teqip.jdvu.ac.in/QSAR_Tools/
Abbreviations
- QSAR:
-
Quantitative structure–activity relationship
- RA:
-
Read-across
- RASAR:
-
Read-across structure–activity relationship
- PLS:
-
Partial least squares
- ICP:
-
Intelligent consensus predictions
- CM:
-
Consensus model
- SD:
-
Standard deviation
- CV:
-
Coefficient of variation
References
Bowes J, Brown AJ, Hamon J, Jarolimek W, Sridhar A, Waldron G, Whitebread G (2012) Reducing safety-related drug attrition: the use of in vitro pharmacological profiling. Nat Rev Drug Discov 11: 909–922. https://www.nature.com/articles/nrd3845
Knapen D, Angrish MM, Fortin MC, Katsiadaki I, Leonard M, Margiotta-Casaluci L, Munn S, O’Brien JM, Pollesch N, Smith LC, Zhang X, Villeneuvei DL (2018) Adverse outcome pathway networks I: development and applications. Environ Toxicol Chem 37(6):1723–1733. https://doi.org/10.1002/etc.4125
Maldonado AG, Doucet JP, Petitjean M, Fan BT (2006) Molecular similarity and diversity in chemoinformatics: from theory to applications. Mol Divers 10:39–79. https://doi.org/10.1007/s11030-006-8697-1
Schultz TW, Amcoff P, Berggren E, Gautier F, Klaric M, Knight DJ, Mahony C, Schwarz M, White A, Cronin MTD (2015) A strategy for structuring and reporting a read-across prediction of toxicity. Regul Toxicol Pharmacol 72:586–601. https://doi.org/10.1016/j.yrtph.2015.05.016
Luechtefeld T, Maertens A, Russo DP, Rovida C, Zhu H, Hartung T (2016) Analysis of publically available skin sensitization data from REACH registrations 2008–2014. Altex 33(2):135–148. https://doi.org/10.14573/altex.1510055
Luechtefeld T, Marsh D, Rowlands C, Hartung T (2018) Machine learning of toxicological big data enables read-across structure activity relationships (RASAR) outperforming animal test reproducibility. Toxicol Sci 165(1):198–212. https://doi.org/10.1093/toxsci/kfy152
Hemmerich J, Ecker GF (2020) In silico toxicology: from structure–activity relationships towards deep learning and adverse outcome pathways WIREs. Comput Mol Sci 10:e1475. https://doi.org/10.1002/wcms.1475
Huang T, Sun G, Zhao L, Zhang N, Zhong R, Peng Y (2021) Quantitative structure-activity relationship (QSAR) studies on the toxic effects of nitroaromatic compounds (NACs): a systematic review. Int J Mol Sci 22:8557. https://doi.org/10.3390/ijms22168557
Diwan M, AbdulHameed M, Liu R, Schyman P, Sachs D, Xu Z, Desai V, Wallqvist A (2021) ToxProfiler: toxicity-target profiler based on chemical similarity. Comput Toxicol 18:100162. https://doi.org/10.1016/j.comtox.2021.100162
Villeneuve DL, Angrish MM, Fortin MC, Katsiadaki I, Leonard M, Margiotta-Casaluci L, Munn S, O’Brien JM, Pollesch NL, Smith LC, Zhang X, Knapen D (2018) Adverse outcome pathway networks II: network analytics. Environ Toxicol Chem 37(6):1734–1748. https://doi.org/10.1002/etc.4124
Wu J, D’Ambrosi S, Ammann L, Stadnicka-Michalak J, Schirmer K, Baity-Jesi M (2022) Predicting chemical hazard across taxa through machine learning. Environ Int 163:107184. https://doi.org/10.1016/j.envint.2022.107184
Banerjee A, De P, Kumar V, Kar S, Roy K (2022) Quick and efficient quantitative predictions of androgen receptor binding affinity for screening endocrine disruptor chemicals using 2D-QSAR and chemical read-across. ChemRxiv. https://doi.org/10.26434/chemrxiv-2022-gcrjg
Chatterjee M, Banerjee A, De P, Gajewicz-Skretna A, Roy K (2022) A novel quantitative read-across tool designed purposefully to fill the existing gaps in nanosafety data. Environ Sci Nano 9:189–203. https://doi.org/10.1039/D1EN00725D
Wallach I, Heifets A (2018) Most ligand-based classification benchmarks reward memorization rather than generalization. J Chem Inf Model 58(5):916–932. https://doi.org/10.1021/acs.jcim.7b00403
Wold S, Sjöström M, Eriksson L (2001) PLS-regression: a basic tool of chemometrics. Chemom Intell Lab Sys 58(2):109–130. https://doi.org/10.1016/S0169-7439(01)00155-1
Roy K, Kar S, Das RN (2015) Understanding the basics of QSAR for applications in pharmaceutical sciences and risk assessment. Academic Press, NY
Roy K, Das RN, Ambure P, Aher RB (2016) Be aware of error measures further studies on validation of predictive QSAR models. Chemom Intell Lab Sys 152:18–33. https://doi.org/10.1016/j.chemolab.2016.01.008
Gramatica P, Sangion AA (2016) A historical excursus on the statistical validation parameters for QSAR Models: A clarification concerning metrics and terminology. J Chem Inf Model 56:1127–1131. https://doi.org/10.1021/acs.jcim.6b00088
Roy K, Ambure P, Kar S, Ojha PK (2018) Is it possible to improve the quality of predictions from an “intelligent” use of multiple QSAR/QSPR/QSTR models? J Chemom 32:e2992. https://doi.org/10.1002/cem.2992
Eriksson L, Byrne T, Johansson E, Trygg J, Vikström C (2013) Multi- and megavariate data analysis basic principles and applications. Umetrics Academy, Umeå
Hong H, Fang H, **e Q, Perkins R, Sheehan DM, Tong W (2003) Comparative molecular field analysis (CoMFA) model using a large diverse set of natural, synthetic and environmental chemicals for binding to the androgen receptor. SAR QSAR Environ Res 14(5–6):373–388. https://doi.org/10.1080/10629360310001623962
Piir G, Sild S, Maran U (2021) Binary and multi-class classification for androgen receptor agonists, antagonists and binders. Chemosphere 262:128313. https://doi.org/10.1016/j.chemosphere.2020.128313
Lill MA, Winiger F, Vedani A, Ernst B (2005) Impact of induced fit on ligand binding to the androgen receptor: a multidimensional QSAR study to predict endocrine-disrupting effects of environmental chemicals. J Med Chem 48:5666–5674. https://doi.org/10.1021/jm050403f
Bennett MJ, Albert RH, Jez JM, Ma H, Penning TM, Lewis M (1997) Steroid recognition and regulation of hormone action: crystal structure of testosterone and NADP+ bound to 3α-hydroxysteroid/dihydrodiol dehydrogenase. Structure 5:799–812. https://doi.org/10.1016/S0969-2126(97)00234-7
Banerjee A, Chatterjee M, De P, Roy K (2022) Quantitative predictions from chemical read-across and their confidence measures. ChemRxiv. https://doi.org/10.26434/chemrxiv-2022-4s53w
Cherkasov A, Muratov EN, Fourches D, Varnek A, Baskin II, Cronin M, Dearden J, Gramatica P, Martin YC, Todeschini R, Consonni V, Kuz’min VE, Cramer R, Benigni R, Yang C, Rathman J, Terfloth L, Gasteiger J, Richard A, Tropsha A (2014) QSAR modeling: where have you been? Where are you going to? J Med Chem 57:4977–5010. https://doi.org/10.1021/jm4004285
Fujita T, Winkler DA (2016) Understanding the roles of the “Two QSARs.” J Chem Inf Model 56(2):269–274. https://doi.org/10.1021/acs.jcim.5b00229
Funding
This research is funded by Science and Engineering Research Board (SERB), New Delhi under the MATRICS scheme (MTR/2019/000008). AB thanks Jadavpur University, Kolkata for a scholarship.
Author information
Authors and Affiliations
Contributions
The manuscript was written through contributions of both authors. Both authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Declared none.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
RASAR Descriptor Calculator v1.0 has now been made available at https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/home.
Supplementary Information
Below is the link to the electronic supplementary material.
11030_2022_10478_MOESM2_ESM.docx
Supplementary Material SI-2 contains score plots, applicability domain plots and randomization plots of models M1 to M4 (DOCX 1671 KB)
Rights and permissions
About this article
Cite this article
Banerjee, A., Roy, K. First report of q-RASAR modeling toward an approach of easy interpretability and efficient transferability. Mol Divers 26, 2847–2862 (2022). https://doi.org/10.1007/s11030-022-10478-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-022-10478-6