Abstract
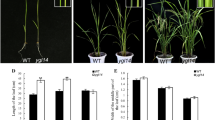
Leaf color affects the efficiency of photosynthesis, and leaf color mutants are important genetic materials for studying the mechanisms of photosynthesis, chlorophyll biosynthesis, and chloroplast development in rice. In this study, a white-striped leaf mutant, wst1, was obtained from the mutant population of the indica restorer line ‘Chuanhui 907’ (R907) when treated with 60Co-γ radiation. Compared to the wild-type, the wst1 mutant showed normal leaf color before tillering and white stripes on the leaf and leaf sheaths after tillering. The chlorophyll and carotenoid contents were significantly reduced, and the thylakoids of chloroplasts developed abnormalities in wst1 plants in the tillering stage. The results of agronomic trait analysis showed that the number of effective panicles, plant height, seed setting rate, and 1000-grain weight of the wst1 mutant were significantly lower than those of the wild-type. Genetic analysis revealed that the phenotype of the wst1 mutant is controlled by a pair of recessive nuclear genes. The candidate gene was mapped to a 72 kb region between the InDel markers M6 and M12 on the short arm of chromosome 1 using molecular marker linkage analysis. Candidate genes were sequenced on the interval, and a G base was replaced by A at the 6972nd position on the 16th exon of LOC_Os01g01920, which encoded a previously reported protein containing the HD domain, WSF3/WFSL1, leading to alternative splicing, causing a 104 bp deletion in the coding region, and resulting in mistranslation after the 490 amino acid of the encoded protein translation in wst1. RT-qPCR analysis showed that the expression levels of most genes related to chlorophyll synthesis and chloroplast development were significantly altered in wst1 plants. Our study identified a novel allele of wsf3 and wfsl1 mutant and provided a new genetic resource and theoretical basis for further understanding of the molecular mechanism of WST1 gene regulation of white-striped leaves in rice.






Similar content being viewed by others
References
Aluru MR, Rodermel SR (2004) Control of chloroplast redox by the IMMUTANS terminal oxidase. Physiol Plant 120(1):4–11. https://doi.org/10.1111/j.0031-9317.2004.0217.x
Aravind L, Koonin EV (1998) The HD domain defines a new superfamily of metal-dependent phosphohydrolases. Trends Biochem Sci 23(12):469–472. https://doi.org/10.1016/S0968-0004(98)01293-6
Dai R, Qian X, Sun J, Lu T, Jia Q, Lu T, Lu M, Rao Y (2023) Research progress on the mechanism of leaf color regulation and related genes in rice. Chin Bull Bot 58(5):799–812. https://doi.org/10.11983/CBB23055
Eberhard S, Finazzi G, Wollman F-A (2008) The dynamics of photosynthesis. Annu Rev Genet 42:463–515. https://doi.org/10.1146/annurev.genet.42.110807.091452
Fambrini M, Castagna A, Dalla Vecchia F, Degl’Innocenti E, Ranieri A, Vernieri P, Pardossi A, Guidi L, Rascio N, Pugliesi C (2004) Characterization of a pigment-deficient mutant of sunflower (Helianthus annuus L.) with abnormal chloroplast biogenesis, reduced PS II activity and low endogenous level of abscisic acid. Plant Sci 167(1):79–89. https://doi.org/10.1016/j.plantsci.2004.03.002
Ge C, Wang L, Ye W, Wu L, Cui Y, Chen P, Pan J, Zhang D, Hu J, Zeng D (2017) Single-point mutation of an histidine-aspartic domain-containing gene involving in chloroplast ribosome biogenesis leads to white fine stripe leaf in rice. Sci Rep 7(1):1–12. https://doi.org/10.1038/s41598-017-03327-2
Hiratsuka J, Shimada H, Whittier R, Ishibashi T, Sakamoto M, Mori M, Kondo C, Honji Y, Sun C-R, Meng B-Y (1989) The complete sequence of the rice (Oryza sativa) chloroplast genome: intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol Gen Genet 217(2):185–194. https://doi.org/10.1007/BF02464880
Jung K-H, Hur J, Ryu C-H, Choi Y, Chung Y-Y, Miyao A, Hirochika H, An G (2003) Characterization of a rice chlorophyll-deficient mutant using the T-DNA gene-trap system. Plant Cell Physiol 44(5):463–472. https://doi.org/10.1093/pcp/pcg064
Kusumi K, Yara A, Mitsui N, Tozawa Y, Iba K (2004) Characterization of a rice nuclear-encoded plastid RNA polymerase gene OsRpoTp. Plant Cell Physiol 45(9):1194–1201. https://doi.org/10.1093/pcp/pch133
Lee S, Kim J-H, Yoo ES, Lee C-H, Hirochika H, An G (2005) Differential regulation of chlorophyll a oxygenase genes in rice. Plant Mol Biol 57(6):805–818. https://doi.org/10.1007/s11103-005-2066-9
Leister D (2003) Chloroplast research in the genomic age. Trends Genet 19(1):47–56. https://doi.org/10.1016/S0168-9525(02)00003-3
Lichtenthaler HK (1987) Chlorophylls and carotenoids: Pigments of photosynthetic biomembranes. Methods Enzymol 148C:350–382. https://doi.org/10.1016/0076-6879(87)48036-1
Liu X, Lan J, Huang Y, Cao P, Zhou C, Ren Y, He N, Liu S, Tian Y, Nguyen T (2018) WSL5, a pentatricopeptide repeat protein, is essential for chloroplast biogenesis in rice under cold stress. J Exp Bot 69(16):3949–3961. https://doi.org/10.1093/jxb/ery214
Liu L, You J, Zhu Z, Chen K, Hu M, Gu H, Liu Z, Wang Z, Wang Y, Liu S (2020) WHITE STRIPE LEAF8, encoding a deoxyribonucleoside kinase, is involved in chloroplast development in rice. Plant Cell Rep 39(1):19–33. https://doi.org/10.1007/s00299-019-02470-6
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Matin MN, Saief SA, Rahman MM, Lee DH, Kang H, Lee DS, Kang SG (2010) Comparative phenotypic and physiological characteristics of spotted leaf 6 (spl6) and brown leaf spot2 (bl2) lesion mimic mutants (LMM) in rice. Mol Cells 30(6):533–543. https://doi.org/10.1007/s10059-010-0151-7
Michelmore RW, Paran I, Kesseli R (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci 88(21):9828–9832. https://doi.org/10.1073/pnas.88.21.982
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5(2):69–76. https://doi.org/10.1007/BF00020088
Sakuraba Y, Rahman ML, Cho SH, Kim YS, Koh HJ, Yoo SC, Paek NC (2013) The rice faded green leaf locus encodes protochlorophyllide oxidoreductase B and is essential for chlorophyll synthesis under high light conditions. The Plant J 74(1):122–133. https://doi.org/10.1111/tpj.12110
Steiner S, Dietzel L, Schröter Y, Fey V, Wagner R, Pfannschmidt T (2009) The role of phosphorylation in redox regulation of photosynthesis genes psaA and psbA during photosynthetic acclimation of mustard. Mol Plant 2(3):416–429. https://doi.org/10.1093/mp/ssp007
Sugimoto H, Kusumi K, Tozawa Y, Yazaki J, Kishimoto N, Kikuchi S, Iba K (2004) The virescent-2 mutation inhibits translation of plastid transcripts for the plastid genetic system at an early stage of chloroplast differentiation. Plant Cell Physiol 45(8):985–996. https://doi.org/10.1093/pcp/pch111
Sugimoto H, Kusumi K, Noguchi K, Yano M, Yoshimura A, Iba K (2007) The rice nuclear gene, VIRESCENT 2, is essential for chloroplast development and encodes a novel type of guanylate kinase targeted to plastids and mitochondria. Plant J 52(3):512–527. https://doi.org/10.1111/j.1365-313X.2007.03251.x
Sugiyama N, Izawa T, Oikawa T, Shimamoto K (2001) Light regulation of circadian clock-controlled gene expression in rice. Plant J 26(6):607–615. https://doi.org/10.1046/j.1365-313x.2001.01063.x
Sun Y, Tian Y, Cheng S, Wang Y, Hao Y, Zhu J, Zhu X, Zhang Y, Yu M, Lei J (2019) WSL6 encoding an Era-type GTP-binding protein is essential for chloroplast development in rice. Plant Mol Biol 100(6):635–645. https://doi.org/10.1007/s11103-019-00885-z
Suzuki Y, Makino A (2012) Availability of Rubisco small subunit up-regulates the transcript levels of large subunit for stoichiometric assembly of its holoenzyme in rice. Plant Physiol 160(1):533–540. https://doi.org/10.1104/pp.112.201459
Tan J, Tan Z, Wu F, Sheng P, Heng Y, Wang X, Ren Y, Wang J, Guo X, Zhang X (2014) A novel chloroplast-localized pentatricopeptide repeat protein involved in splicing affects chloroplast development and abiotic stress response in rice. Mol Plant 7(8):1329–1349. https://doi.org/10.1093/mp/ssu054
von Caemmerer S, Quick WP, Furbank RT (2012) The development of C4 rice: current progress and future challenges. Sci 336(6089):1671–1672. https://doi.org/10.1126/science.12201
Wang P, Gao J, Wan C, Zhang F, Xu Z, Huang X, Sun X, Deng X (2010) Divinyl chlorophyll (ide) a can be converted to monovinyl chlorophyll (ide) a by a divinyl reductase in rice. Plant Physiol 153(3):994–1003. https://doi.org/10.1104/pp.110.158477
Wang L, Wang C, Wang Y, Niu M, Ren Y, Zhou K, Zhang H, Lin Q, Wu F, Cheng Z (2016) WSL3, a component of the plastid-encoded plastid RNA polymerase, is essential for early chloroplast development in rice. Plant Mol Biol 92(4):581–595. https://doi.org/10.1007/s11103-016-0533-0
Wang Y, Ren Y, Zhou K, Liu L, Wang J, Xu Y, Zhang H, Zhang L, Feng Z, Wang L (2017) WHITE STRIPE LEAF4 encodes a novel P-type PPR protein required for chloroplast biogenesis during early leaf development. Front Plant Sci 8:1116. https://doi.org/10.3389/fpls.2017.01116
Wang Y, Zhong P, Zhang X, Liu J, Zhang C, Yang X, Wan C, Liu C, Zhou H, Yang B (2019) GRA78 encoding a putative S-sulfocysteine synthase is involved in chloroplast development at the early seedling stage of rice. Plant Sci 280:321–329. https://doi.org/10.1016/j.plantsci.2018.12.019
Waters MT, Langdale JA (2009) The making of a chloroplast. EMBO J 28(19):2861–2873. https://doi.org/10.1038/emboj.2009.264
Yang Y, Xu J, Huang L, Leng Y, Dai L, Rao Y, Chen L, Wang Y, Tu Z, Hu J (2016) PGL, encoding chlorophyllide a oxygenase 1, impacts leaf senescence and indirectly affects grain yield and quality in rice. J Exp Bot 67(5):1297–1310. https://doi.org/10.1093/jxb/erv529
Ye W, Hu S, Wu L, Ge C, Cui Y, Chen P, Wang X, Xu J, Ren D, Dong G (2016) White stripe leaf 12 (WSL12), encoding a nucleoside diphosphate kinase 2 (OsNDPK2), regulates chloroplast development and abiotic stress response in rice (Oryza sativa L.). Mol Breed 36(5):1–15. https://doi.org/10.1007/s11032-016-0479-6
Yoo S-C, Cho S-H, Sugimoto H, Li J, Kusumi K, Koh H-J, Iba K, Paek N-C (2009) Rice virescent3 and stripe1 encoding the large and small subunits of ribonucleotide reductase are required for chloroplast biogenesis during early leaf development. Plant Physiol 150(1):388–401. https://doi.org/10.1104/pp.109.136648
Zhang H, Liu L, Cai M, Zhu S, Zhao J, Zheng T, Xu X, Zeng Z, Niu J, Jiang L (2015) A point mutation of magnesium chelatase OsCHLI gene dampens the interaction between CHLI and CHLD subunits in rice. Plant Mol Biol Rep 33(6):1975–1987. https://doi.org/10.1007/s11105-015-0889-3
Zhang Z, Cui X, Wang Y, Wu J, Gu X, Lu T (2017) The RNA editing factor WSP1 is essential for chloroplast development in rice. Mol Plant 10(1):86–98. https://doi.org/10.1016/j.molp.2016.08.009
Zhang H, Zhou J-F, Kan Y, Shan J-X, Ye W-W, Dong N-Q, Guo T, **ang Y-H, Yang Y-B, Li Y-C (2022) A genetic module at one locus in rice protects chloroplasts to enhance thermotolerance. Sci 376(6599):1293–1300. https://doi.org/10.1126/science.abo5721
Zhao S, Long W, Wang Y, Liu L, Wang Y, Niu M, Zheng M, Wang D, Wan J (2016) A rice White-stripe leaf3 (wsl3) mutant lacking an HD domain-containing protein affects chlorophyll biosynthesis and chloroplast development. J Plant Biol 59(3):282–292. https://doi.org/10.1007/s12374-016-0459-8
Zhou S, Sawicki A, Willows RD, Luo M (2012) C-terminal residues of Oryza sativa GUN4 are required for the activation of the ChlH subunit of magnesium chelatase in chlorophyll synthesis. FEBS Lett 586(3):205–210. https://doi.org/10.1016/j.febslet.2011.12.026
Zhou K, Ren Y, Zhou F, Wang Y, Zhang L, Lyu J, Wang Y, Zhao S, Ma W, Zhang H (2017) Young Seedling Stripe1 encodes a chloroplast nucleoid-associated protein required for chloroplast development in rice seedlings. Planta 245(1):45–60. https://doi.org/10.1007/s00425-016-2590-7
Zhu X, Mou C, Zhang F, Huang Y, Yang C, Ji J, Liu X, Cao P, Nguyen T, Lan J (2020) WSL9 encodes an HNH endonuclease domain-containing protein that is essential for early chloroplast development in rice. Rice 13(1):1–13. https://doi.org/10.1186/s12284-020-00407-2
Acknowledgements
This study was supported by the National Natural Science Foundation of China-Youth Science Fund (32001531), Sichuan Science and Technology Program (2022ZDZX0012, and 2021YFYZ0016).
Funding
This study was supported by the National Natural Science Foundation of China-Youth Science Fund (32001531), Sichuan Science and Technology Program (2022ZDZX0012, and 2021YFYZ0016).
Author information
Contributions
BH, ZP, and PW planned and designed the research. BH, ZH, XX, AD, HL, MW, LW, and CZ performed the experiments. LW and PW conducted fieldwork. BH, ZH, and ZP analyzed the data. BH and XX wrote the manuscript. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, B., He, Z., ** of a white stripe leaf mutant in rice. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01897-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01897-5