Abstract
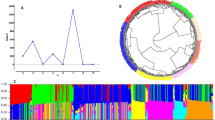
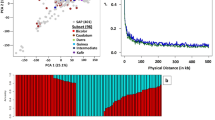
Genetic diversity in plant breeding is key to dissecting the genetic architecture of complex economical traits. Genome-wide association studies (GWAS) and genomic selection (GS) have emerged as promising tools to accelerate agronomic trait improvement efficiency, defining current breeding strategies. The key objective of this study was to investigate the genetic architecture and perform genomic prediction of yield, specific gravity (SG), and dry matter (DM) traits, using a diversity panel of 568 Solanum tuberosum Andigenum accessions. Analyses were carried out using a total of 4271 Single Nucleotide Polymorphisms (SNPs) obtained using the SolCAP SNP array 8 K version. Yield data were collected from field evaluations during three consecutive years, while tuber quality traits (SG and DM) were evaluated in 1 year. To this end, population structure was first examined and revealed two main subpopulations. GWAS analysis identified candidate loci for yield in chromosomes 1, 3, 4 and 11, with minor effects on phenotypic variation (< 1–10%). Important genomic regions were detected for SG in chromosomes 5, 6, and 8, and in DM in chromosome 7, though they explained only a small fraction of the phenotypic variation (< 1–2% for SG and 3% for DM). Functional annotation of candidate genes showed promising genomic regions related to tuber growth, quality and development that could further be verified by other tools (i.e., gene editing). Furthermore, the predicted capacity of the cross-validation process between subpopulations was low for yield and specific gravity. However, when the prediction process was conducted across the entire population, the predictive power increase, as a greater diversity in the data enables better prediction. Instead, dry matter genomic prediction accuracies were higher in accordance with its higher narrow sense heritability value. These findings enriched GWAS analyses in an Andigenum potato collection and reinforce the potential of applying GS in Colombian breeding programs using diverse populations.





Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are available from the first author on reasonable request.
References
Alvarado J, Aguilera JM (2001) Métodos para Medir Propiedades Físicas en Industrias de Alimentos. In: Acribia (ed) Métodos para Medir Propiedades Físicas en Industrias de Alimentos. Zaragosa, España, p 426
Amoros W, Salas E, Hualla V et al (2020) Heritability and genetic gains for iron and zinc concentration in diploid potato. Crop Sci 60:1884–1896. https://doi.org/10.1002/csc2.20170
Asmamaw Y, Tekalign T, Workneh TS (2010) Specific gravity, dry matter concentration, pH, and crisp-making potential of ethiopian potato (Solanum tuberosum L.) cultivars as influenced by growing environment and length of storage under ambient conditions. Potato Res 53:95–109. https://doi.org/10.1007/s11540-010-9154-1
Barone A (2004) Molecular marker-assisted selection for potato breeding. Am J Potato Res 81:111–117. https://doi.org/10.1007/BF02853608
Berdugo-Cely J, Valbuena RI, Sánchez-Betancourt E et al (2017) Genetic diversity and association map** in the Colombian central collection of Solanum tuberosum L. Andigenum group using SNPs markers. PLoS ONE 12:e0173039. https://doi.org/10.1371/journal.pone.0173039
Berdugo-Cely JA, del Céron-Lasso MS, Yockteng R (2023) Phenotypic and molecular analyses in diploid and tetraploid genotypes of Solanum tuberosum L. reveal promising genotypes and candidate genes associated with phenolic compounds, ascorbic acid contents, and antioxidant activity. Front Plant Sci 13:21. https://doi.org/10.3389/fpls.2022.1007104
Burlingame B, Mouillé B, Charrondière R (2009) Nutrients, bioactive non-nutrients and anti-nutrients in potatoes. J Food Compos Anal 22:494–502. https://doi.org/10.1016/j.jfca.2009.09.001
Byrne S, Meade F, Mesiti F et al (2020) Genome-wide association and genomic prediction for fry color in potato. Agronomy 10:90. https://doi.org/10.3390/agronomy10010090
Crossa J, Burgueño J, Dreisigacker S et al (2007) Association analysis of historical bread wheat germplasm using additive genetic covariance of relatives and population structure. Genetics 177:1889 LP – 1913. https://doi.org/10.1534/genetics.107.078659
Dong D, Yang Z, Ma Y et al (2022) Expression of a hydroxycinnamoyl-CoA shikimate/quinate hydroxycinnamoyl transferase 4 Gene from Zoysia japonica (ZjHCT4) causes excessive elongation and lignin composition changes in Agrostis stolonifera. Int J Mol Sci 23:9500. https://doi.org/10.3390/ijms23169500
Enciso-Rodriguez F, Douches D, Lopez-Cruz M et al (2018) Genomic Selection for late blight and common scab resistance in tetraploid potato (Solanum tuberosum). G3 Genes|Genomes|Genetics 8:2471–2481. https://doi.org/10.1534/G3.118.200273
Endelman JB, Schmitz Carley CA, Bethke PC et al (2018) Genetic variance partitioning and genome-wide prediction with allele dosage information in Autotetraploid potato. Genetics 209:77–87. https://doi.org/10.1534/genetics.118.300685/-/DC1.1
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
FAOSTAT (2021) FAOSTAT database collections. Food and Agriculture Organization of the United Nations
FEDEPAPA (2019) La papa incluída en una dieta balanceada, No.47. Revista Papa, Bogotá, Colombia
FNFP (2020) Fondo Nacional de Fomento de la Papa. Informe trimestral. IV trimestre - 2020. Sistema de información y estudios económicos. 49
Gemenet DC, Lindqvist-Kreuze H, De Boeck B et al (2020) Sequencing depth and genotype quality: accuracy and breeding operation considerations for genomic selection applications in autopolyploid crops. Theor Appl Genet 133:3345–3363. https://doi.org/10.1007/s00122-020-03673-2
Gromadka R, Cieśla J, Olszak K et al (2018) Genome-wide analysis and expression profiling of calcium-dependent protein kinases in potato (Solanum tuberosum). Plant Growth Regul 84:303–315. https://doi.org/10.1007/s10725-017-0341-9
Habier D, Fernando RL, Dekkers JCM (2007) The impact of genetic relationship information on genome-assisted breeding values. Genetics 177:2389–2397. https://doi.org/10.1534/genetics.107.081190
Habyarimana E, Parisi B, Mandolino G (2017) Genomic prediction for yields, processing and nutritional quality traits in cultivated potato (Solanum tuberosum L.). Plant Breed 136:245–252. https://doi.org/10.1111/pbr.12461
Hoffmann L, Maury S, Martz F et al (2003) Purification, cloning, and properties of an acyltransferase controlling shikimate and quinate ester intermediates in phenylpropanoid metabolism. J Biol Chem 278:95–103. https://doi.org/10.1074/jbc.M209362200
Hussen ES (2019) Review on genetic variation in potato (Solanum Tuberosum L.) for processing quality traits. J Nat Sci Res 9:31–36. https://doi.org/10.7176/jnsr/9-12-05
Jannink J-L, Lorenz AJ, Iwata H (2010) Genomic selection in plant breeding: from theory to practice. Brief Funct Genomics 9:166–177. https://doi.org/10.1093/bfgp/elq001
Jansky S (2009) Breeding, genetics, and cultivar development. In: Press A (ed) Advances in potato chemistry and technology, 1st edn. Elsevier, pp 27–62
**g S, Sun X, Yu L et al (2022) Transcription factor StABI5-like 1 binding to the FLOWERING LOCUS T homologs promotes early maturity in potato. Plant Physiol 189:1677–1693. https://doi.org/10.1093/plphys/kiac098
Krannitz PG, Aarssen LW, Lefebvre DD (1991) Short-term competition for phosphate between two genotypes of Arabidopsis thaliana (L.) Heynh. New Phytol 119:389–396. https://doi.org/10.1111/j.1469-8137.1991.tb00038.x
Krishnappa G, Savadi S, Tyagi BS et al (2021) Integrated genomic selection for rapid improvement of crops. Genomics 113:1070–1086. https://doi.org/10.1016/J.YGENO.2021.02.007
Li X, Xu J, Duan S et al (2018) Map** and QTL Analysis of early-maturity traits in tetraploid potato (Solanum tuberosum L.). Int J Mol Sci 19:3065. https://doi.org/10.3390/ijms19103065
Li J, Wang Y, Wen G et al (2019) Map** QTL underlying tuber starch content and plant maturity in tetraploid potato. Crop J 7:261–272. https://doi.org/10.1016/j.cj.2018.12.003
Manrique-Carpintero NC, Coombs JJ, Cui Y et al (2015) Genetic map and QTL analysis of agronomic traits in a diploid potato population using single nucleotide polymorphism markers. Crop Sci 55:2566–2579. https://doi.org/10.2135/cropsci2014.10.0745
Manrique-Carpintero NC, Berdugo-Cely JA, Cerón-Souza I et al (2023) Defining a diverse core collection of the Colombian central collection of potatoes: a tool to advance research and breeding. Front Plant Sci 14:1046400. https://doi.org/10.3389/FPLS.2023.1046400
Massa AN, Manrique-Carpintero NC, Coombs JJ et al (2015) Genetic linkage map** of economically important traits in cultivated tetraploid potato (Solanum tuberosum L.). G3 Genes, Genomes, Genet 5:2357–2364. https://doi.org/10.1534/g3.115.019646
Massa AN, Manrique-Carpintero NC, Coombs J et al (2018) Linkage analysis and QTL map** in a tetraploid russet map** population of potato. BMC Genet 19:87. https://doi.org/10.1186/s12863-018-0672-1
McCord PH, Sosinski BR, Haynes KG et al (2011) Linkage map** and QTL analysis of agronomic traits in tetraploid potato (Solanum tuberosum subsp. tuberosum). Crop Sci 51:771–785. https://doi.org/10.2135/cropsci2010.02.0108
Meuwissen THE, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829. https://doi.org/10.1093/genetics/157.4.1819
Moreno JD, Valbuena RI (2006) Colección central colombiana de papa: riqueza de variabilidad genética para el mejoramiento del cultivo. Rev Innovación y Cambio Tecnológico, pp 16–24
Moskvina V, Schmidt KM (2008) On multiple-testing correction in genome-wide association studies. Genet Epidemiol 32:567–573. https://doi.org/10.1002/GEPI.20331
National Oceanic and Atmospheric Administration NOAA (2023) Cold & Warm Episodes by Season. In: Natl. Centers Environ. Predict. https://origin.cpc.ncep.noaa.gov/products/analysis_monitoring/ensostuff/ONI_v5.php. Accessed 20 Oct 2023
Oliver SN, Lunn JE, Urbanczyk-Wochniak E et al (2008) Decreased expression of cytosolic pyruvate kinase in potato tubers leads to a decline in pyruvate resulting in an in vivo repression of the alternative oxidase. Plant Physiol 148:1640–1654. https://doi.org/10.1104/pp.108.126516
Ottoman RJ, Hane DC, Brown CR et al (2009) Validation and implementation of marker-assisted selection (MAS) for PVY resistance (Ryadggene) in a tetraploid potato breeding program. Am J Potato Res 86:304–314. https://doi.org/10.1007/s12230-009-9084-0
Pandey J, Scheuring DC, Koym JW, Vales MI (2022) Genomic regions associated with tuber traits in tetraploid potatoes and identification of superior clones for breeding purposes. Front Plant Sci 13:1–16. https://doi.org/10.3389/fpls.2022.952263
Park J, Massa AN, Douches D et al (2021) Linkage and QTL map** for tuber shape and specific gravity in a tetraploid map** population of potato representing the russet market class. BMC Plant Biol 21:1–18. https://doi.org/10.1186/s12870-021-03265-2
Pérez-Rodríguez P, Crossa J, Rutkoski J et al (2017) Single-step genomic and pedigree genotype × environment interaction models for predicting wheat lines in international environments. Plant Genome. https://doi.org/10.3835/PLANTGENOME2016.09.0089
Pritchard JK (2010) Documentation for structure software: version 2.3
R Core Team (2021) R: a language and environment for statistical computing
Rosyara UR, De JWS, Douches DS, Endelman JB (2016) Software for genome-wide association studies in autopolyploids and its application to potato. Plant Genome 9:10. https://doi.org/10.3835/PLANTGENOME2015.08.0073
Rousseeuw PJ (1987) Silhouettes: a graphical aid to the interpretation and validation of cluster analysis. J Comput Appl Math 20:53–65. https://doi.org/10.1016/0377-0427(87)90125-7
Ruiz de Arcaute R, Carrasco A, Ortega F et al (2022) Evaluation of genetic resources in a potato breeding program for chip quality. Agronomy 12:1–21. https://doi.org/10.3390/agronomy12051142
Santa JD, Berdugo-Cely J, Cely-Pardo L et al (2018) QTL analysis reveals quantitative resistant loci for Phytophthora infestans and Tecia solanivora in tetraploid potato (Solanum tuberosum L.). PLoS ONE 13:e0199716. https://doi.org/10.1371/journal.pone.0199716
Schmitz Carley CA, Coombs JJ, Douches DS et al (2017) Automated tetraploid genotype calling by hierarchical clustering. Theor Appl Genet 130:717–726. https://doi.org/10.1007/S00122-016-2845-5
Schönhals EM, Ding J, Ritter E et al (2017) Physical map** of QTL for tuber yield, starch content and starch yield in tetraploid potato (Solanum tuberosum L.) by means of genome wide genoty** by sequencing and the 8.3 K SolCAP SNP array. BMC Genomics 18:1–20. https://doi.org/10.1186/S12864-017-3979-9
Seid E, Tessema L, Abebe T et al (2023) Genetic variability for micronutrient content and tuber yield traits among biofortified potato (Solanum tuberosum L.) clones in Ethiopia. Plants. https://doi.org/10.3390/plants12142625
Selga C, Reslow F, Pérez-Rodríguez P, Ortiz R (2022) The power of genomic estimated breeding values for selection when using a finite population size in genetic improvement of tetraploid potato. G3 Genes, Genomes, Genet 12:11. https://doi.org/10.1093/G3JOURNAL/JKAB362
Singh P, Arif Y, Siddiqui H et al (2023) Critical factors responsible for potato tuberization. Bot Rev. https://doi.org/10.1007/s12229-023-09289-7
Slater AT, Cogan NOI, Hayes BJ et al (2014) Improving breeding efficiency in potato using molecular and quantitative genetics. Theor Appl Genet 127:2279–2292. https://doi.org/10.1007/s00122-014-2386-8
Sørensen KK, Kirk HG, Olsson K et al (2008) A major QTL and an SSR marker associated with glycoalkaloid content in potato tubers from Solanum tuberosum x S. sparsipilum located on chromosome I. Theor Appl Genet 117:1–9. https://doi.org/10.1007/s00122-008-0745-z
Sowokinos JR, Preiss J (1976) Pyrophosphorylases in Solanum tuberosum. Plant Physiol 57:1459–1466. https://doi.org/10.1104/pp.69.6.1459
Sverrisdóttir E, Sundmark EHR, Johnsen HØ et al (2018) The value of expanding the training population to improve genomic selection models in tetraploid potato. Front Plant Sci 9:1–14. https://doi.org/10.3389/fpls.2018.01118
Tagliotti ME, Deperi SI, Bedogni MC et al (2018) Use of easy measurable phenotypic traits as a complementary approach to evaluate the population structure and diversity in a high heterozygous panel of tetraploid clones and cultivars. BMC Genet 19:1–12. https://doi.org/10.1186/S12863-017-0556-9
Tibbs Cortes L, Zhang Z, Yu J (2021) Status and prospects of genome-wide association studies in plants. Plant Genome 14:e20077. https://doi.org/10.1002/TPG2.20077
Tiwari JK, Siddappa S, Singh BP et al (2013) Molecular markers for late blight resistance breeding of potato: an update. Plant Breed 132:237–245. https://doi.org/10.1111/pbr.12053
Tiwari JK, Buckseth T, Singh RK et al (2020) Prospects of improving nitrogen use efficiency in potato: Lessons from transgenics to genome editing strategies in plants. Front Plant Sci 11:1–6. https://doi.org/10.3389/fpls.2020.597481
Urbany C, Stich B, Schmidt L et al (2011) Association genetics in Solanum tuberosum provides new insights into potato tuber bruising and enzymatic tissue discoloration. BMC Genomics 12:14. https://doi.org/10.1186/1471-2164-12-7
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423. https://doi.org/10.3168/JDS.2007-0980
Wellenreuther M, Hansson B (2016) Detecting polygenic evolution: problems, pitfalls, and promises. Trends Genet 32:155–164. https://doi.org/10.1016/j.tig.2015.12.004
Wilson S, Zheng C, Maliepaard C et al (2021) Understanding the effectiveness of genomic prediction in tetraploid potato. Front Plant Sci 12:1–13. https://doi.org/10.3389/fpls.2021.672417
Acknowledgements
The authors gratefully acknowledge Raul Iván Valbuena-Benavides as the CCC collection leader and curator; he designed and developed the field trials. Furthermore, the authors want to acknowledge Arturo Díaz and Hugo Gómez for their collaboration in the field trials and Shane Teachworth for copy editing.
Funding
This research was supported by the Ministerio de Agricultura y Desarrollo Rural of Colombia under the project “BGA Conservación en Campo BG Vegetal” with 1002443 code, managed by Agrosavia.
Author information
Authors and Affiliations
Contributions
FEE-R, LFF, JB-C and GAG-M conceived the study. ZLL-P and BCO collected, generated, and organized yield and quality data. GAG-M and FEE-R organized and analysed the phenotypic data. JB-C organized and generated the SNP data. CFA performed the GWAS and Genomic selection analysis. The first draft of the manuscript was written by GAG-M, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests to declare that are relevant to the content of this article.
Ethics approval and consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10681_2024_3337_MOESM2_ESM.tiff
(A) Delta K value estimated using the Evanno method (Evanno et al. 2005) and (B) Optimal number of clusters determined by the Silhouette method. (TIFF 217 KB)
10681_2024_3337_MOESM3_ESM.tiff
Boxplots showing the effect of the significant markers associated with (A) Yield, (B) Specific gravity and (C) Dry matter. The letters on the X axis represent allele variants and p represents the variation across variants, 0: AAAA, 1: AAAB, 2: AABB, 3: ABBB or 4: BBBB. (TIFF 111220 KB)
10681_2024_3337_MOESM4_ESM.tiff
Partitioning of the genetic variance for three traits; yield, specific gravity and dry matter. Vg represents additive variance; Vge represents the interaction between year and genotype, and Ve represents residual genetic variance. (TIFF 4903 KB)
10681_2024_3337_MOESM5_ESM.xlsx
List of accessions of the diversity panel from the Colombian Central Collection of S. tuberosum group Andigenum used in this study. (XLSX 40 KB)
10681_2024_3337_MOESM6_ESM.xlsx
List of all QTLs identified by year for the diversity panel from the Colombian Central Collection of S. tuberosum group Andigenum. AD: additive, DOM: 1-dominant, DD: 2-dominant, DGEN: diplo-General, GEN: General. (XLSX 16 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Garzón-Martínez, G.A., Azevedo, C.F., Berdugo-Cely, J.A. et al. Genetic dissection of yield and quality-related traits in a Colombian Andigenum potato collection, revealed by genome-wide association and genomic prediction analyses. Euphytica 220, 79 (2024). https://doi.org/10.1007/s10681-024-03337-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-024-03337-y