Abstract
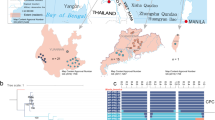
The green peafowl (Pavo muticus, Linnaeus 1766) is an endangered species native to Southeast Asia. Despite considerable morphological diversity, the intraspecific genetic structure of green peafowl has not been comprehensively addressed. We used public whole-genome sequencing data of one blue and 52 green peafowls to characterise their genetic diversity, differentiation, identify Ancestry Informative Markers (AIMs) and compare their demographic histories. We found evidence of substantial population structure, with at least three distinct clusters and diverse demographic histories that may result from different responses to biogeoclimatic events. The genetic structure of native populations follows the pattern of the geographic distribution of the green peafowl with the highest autosomal pairwise FST between Yunnan and Vietnam (~ 0.1) and intermediate estimates for Thailand comparisons (~ 0.077). We identify AIMs to distinguish between these three native populations. The captive green peafowls from **nxing clustered with Vietnam, and those from Qinhuangdao (QHD) formed a separate cluster. The two QHD individuals appear to have varying levels of blue peafowl ancestry based on PCA and admixture analysis and are mirrored in their demographic histories. Our study establishes the occurrence of genetically distinct natural populations of green peafowl that can be considered separate management units (MU) when planning conservation actions.




Similar content being viewed by others
Data availability
All datasets used in this study are compiled from public repositories. The scripts and associated data from the analysis are available here: https://github.com/A**kya-IISERB/Pavo/tree/main/Conservation and https://doi.org/10.17632/ddwbwfjtrj.1.
References
Allendorf FW, Leary RF, Spruell P, Wenburg JK (2001) The problems with hybrids: setting conservation guidelines. Trends Ecol Evol 16:613–622. https://doi.org/10.1016/S0169-5347(01)02290-X
Brickle NW (2002) Habitat use, predicted distribution and conservation of green peafowl (Pavo muticus) in Dak Lak Province, Vietnam. Biol Conserv 105:189–197. https://doi.org/10.1016/S0006-3207(01)00182-3
Browning BL, Tian X, Zhou Y, Browning SR (2021) Fast two-stage phasing of large-scale sequence data. Am J Hum Genet 108:1880–1890. https://doi.org/10.1016/J.AJHG.2021.08.005
Cahill JA, Soares AER, Green RE, Shapiro B (2016) Inferring species divergence times using pairwise sequential markovian coalescent modelling and low-coverage genomic data. Philos Trans R Soc B Biol Sci 371:20150138. https://doi.org/10.1098/rstb.2015.0138
Chen D, Hosner PA, Dittmann DL et al (2021) Divergence time estimation of Galliformes based on the best gene shop** scheme of ultraconserved elements. BMC Ecol Evol 21. https://doi.org/10.1186/S12862-021-01935-1
Cheng SC, Liu CB, Yao XQ et al (2023) Hologenomic insights into mammalian adaptations to myrmecophagy. Natl Sci Rev 10. https://doi.org/10.1093/NSR/NWAC174
Coates DJ, Byrne M, Moritz C (2018) Genetic diversity and conservation units: dealing with the species-population continuum in the age of genomics. Front Ecol Evol 6:165. https://doi.org/10.3389/FEVO.2018.00165/BIBTEX
Conner K, Hartl DL (2004) A primer of Ecological Genetics: a textbook. ISBN: 9780878932023
Conway W (2003) The role of zoos in the 21st century1. Int Zoo Yearb 38:7–13. https://doi.org/10.1111/J.1748-1090.2003.TB02059.X
Conway WG (2011) Buying time for wild animals with zoos. Zoo Biol 30:1–8. https://doi.org/10.1002/ZOO.20352
Darriba Di, Posada D, Kozlov AM et al (2020) ModelTest-NG: a New and Scalable Tool for the selection of DNA and protein evolutionary models. Mol Biol Evol 37:291–294. https://doi.org/10.1093/MOLBEV/MSZ189
Delacour J, Harrison JC, John C, World Pheasant Association (1977). The pheasants of the world. 395. ISBN-10 : 0904558371
Dong F, Kuo H-CC, Chen G-LL et al (2021) Population genomic, climatic and anthropogenic evidence suggest the role of human forces in endangerment of green peafowl (Pavo muticus). Proc Biol Sci. 288(1948):20210073. https://doi.org/10.1098/rspb.2021.0073
Du HY, Zhang XY, Dinh TD et al (2020) Identification of hybrid green peafowl using mitochondrial and nuclear markers. Conserv Genet Resour 12:669–683. https://doi.org/10.1007/S12686-020-01159-3/TABLES/5
Ernst M, Jønsson KA, Ericson PGP et al (2022) Utilising museomics to trace the complex history and species boundaries in an avian-study system of conservation concern. Hered 2022 1283 128:159–168. https://doi.org/10.1038/s41437-022-00499-0
Espindola-Hernandez P, Mueller JC, Kempenaers B (2022) Genomic signatures of the evolution of a diurnal lifestyle in Strigiformes. G3 Genes|Genomes|Genetics 12(8):jkac135. https://doi.org/10.1093/G3JOURNAL/JKAC135
European Conservation Breeding Group (2023) PMI TN Stud book program E. In: Details regarding Studb. PMI-TN. http://pfauenfarm.de/Home-English/Our-Peafowl/Pavo-Mutisus-Imperator-E/PMI-TN-Stud-book-program-E/pmi-tn-stud-book-program-e.html. Accessed 4 May 2023
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/J.1365-294X.2005.02553.X
Fraser J, Wharton D (2007) The future of Zoos: a New Model for Cultural Institutions. Curator Museum J 50:41–54. https://doi.org/10.1111/J.2151-6952.2007.TB00248.X
Funk WC, McKay JK, Hohenlohe PA, Allendorf FW (2012) Harnessing genomics for delineating conservation units. Trends Ecol Evol 27:489–496. https://doi.org/10.1016/J.TREE.2012.05.012
Gautier M, Klassmann A, Vitalis R (2017) Rehh 2.0: a reimplementation of the R package rehh to detect positive selection from haplotype structure. Mol Ecol Resour 17:78–90. https://doi.org/10.1111/1755-0998.12634
Gu B, Wang F (2021) A review on the ecology and conservation biology of green peafowl (Pavo muticus). Biodivers Sci 29:1554. https://doi.org/10.17520/BIODS.2021144
Hernowo JB, Mardiastuti ANI, Alikodra HS, Kusmana CECEP (2011) Behavior Ecology of the Javan Green Peafowl (Pavo muticus muticus Linnaeus 1758) in baluran and alas Purwo National Park, East Java. HAYATI J Biosci 18:164–176. https://doi.org/10.4308/hjb.18.4.164
Höglund J, Laurila A, Rödin-Mörch P (2019) Population Genomics and Wildlife Adaptation in the Face of Climate Change. 333–355. https://doi.org/10.1007/13836_2019_69
Jackson CE (2006) Peacock. Reaktion Books, London, U.K. ISBN-10: 1861892934
Jaiswal SK, Gupta A, Saxena R et al (2018) Genome sequence of peacock reveals the Peculiar Case of a glittering bird. Front Genet 9:392. https://doi.org/10.3389/fgene.2018.00392
Johnson CN, Balmford A, Brook BW et al (2017) Biodiversity losses and conservation responses in the Anthropocene. Sci (80-) 356:270–275. https://doi.org/10.1126/SCIENCE.AAM9317/SUPPL_FILE/AAM9317_JOHNSON_SM.PDF
Kardos M, Shafer ABA (2018) The peril of gene-targeted conservation. Trends Ecol Evol 33:827–839. https://doi.org/10.1016/J.TREE.2018.08.011
Kishida T (2017) Population history of Antarctic and common minke whales inferred from individual whole-genome sequences. Mar Mammal Sci 33:645–652. https://doi.org/10.1111/mms.12369
Ko S, Chu BB, Peterson D et al (2023) Unsupervised discovery of ancestry-informative markers and genetic admixture proportions in biobank-scale datasets. Am J Hum Genet 110:314–325. https://doi.org/10.1016/J.AJHG.2022.12.008
Kong D, Wu F, Shan P et al (2018) Status and distribution changes of the endangered green peafowl (Pavo muticus) in China over the past three decades (1990s-2017). Avian Res 9:1–9. https://doi.org/10.1186/S40657-018-0110-0/TABLES/3
Kopelman NM, Mayzel J, Jakobsson M et al (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191. https://doi.org/10.1111/1755-0998.12387
Korneliussen TS, Albrechtsen A, Nielsen R (2014) ANGSD: analysis of next generation sequencing data. BMC Bioinformatics 15:356. https://doi.org/10.1186/s12859-014-0356-4
Korunes KL, Samuk K (2021) Pixy: unbiased estimation of nucleotide diversity and divergence in the presence of missing data. Mol Ecol Resour 21:1359–1368. https://doi.org/10.1111/1755-0998.13326
Kozlov AM, Darriba D, Flouri T et al (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455. https://doi.org/10.1093/BIOINFORMATICS/BTZ305
Kozma R, Melsted P, Magnússon KP, Höglund J (2016) Looking into the past – the reaction of three grouse species to climate change over the last million years using whole genome sequences. Mol Ecol 25:570–580. https://doi.org/10.1111/MEC.13496
Lee TH, Guo H, Wang X et al (2014) SNPhylo: a pipeline to construct a phylogenetic tree from huge SNP data. BMC Genomics 15:162. https://doi.org/10.1186/1471-2164-15-162
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arxiv. https://doi.org/10.1093/BIOINFORMATICS/BTR509
Li H, Durbin R (2011) Inference of human population history from individual whole-genome sequences. Nature 475:493–496. https://doi.org/10.1038/nature10231
Li H, Handsaker B, Wysoker A et al (2009) The sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Lin F (2010) A Monograph of Peafowl of the Genus Pavo. Can be accessed at https://github.com/A**kya-IISERB/Pavo/blob/main/Conservation/A-monograph-of-peafowl-of-the-genus-pavo-by-frank-lin-photo.pdf
Mason N, Ward M, Watson JEM et al (2020) Global opportunities and challenges for transboundary conservation. Nat Ecol Evol 2020 45 4:694–701. https://doi.org/10.1038/s41559-020-1160-3
McGowan PJK, Duckworth JW, **anji W et al (1998) A review of the status of the Green Peafowl Pavo muticus and recommendations for future action. Bird Conserv Int 8:331–348. https://doi.org/10.1017/S0959270900002100
Meisner J, Albrechtsen A (2018) Inferring Population structure and admixture proportions in low-depth NGS data. Genetics 210:719–731. https://doi.org/10.1534/GENETICS.118.301336
Mikkelson GM, Gonzalez A, Peterson GD (2007) Economic Inequality predicts Biodiversity loss. PLoS ONE 2:e444. https://doi.org/10.1371/JOURNAL.PONE.0000444
Nuttall M, Nut M, Ung V, O’Kelly H (2017) Abundance estimates for the endangered green peafowl Pavo muticus in Cambodia: identification of a globally important site for conservation. Bird Conserv Int 27:127–139. https://doi.org/10.1017/S0959270916000083
Palsbøll PJ, Bérubé M, Allendorf FW (2007) Identification of management units using population genetic data. Trends Ecol Evol 22:11–16. https://doi.org/10.1016/J.TREE.2006.09.003
Raj A, Stephens M, Pritchard JK (2014) FastSTRUCTURE: variational inference of population structure in large SNP data sets. Genetics 197:573–589. https://doi.org/10.1534/GENETICS.114.164350/-/DC1
Rellstab C, Dauphin B, Exposito-Alonso M (2021) Prospects and limitations of genomic offset in conservation management. Evol Appl 14:1202–1212. https://doi.org/10.1111/EVA.13205
Salles T, Mallard C, Husson L et al (2021) Quaternary landscape dynamics boosted species dispersal across Southeast Asia. Commun Earth Environ 2021 21 2:1–12. https://doi.org/10.1038/s43247-021-00311-7
Saridnirun G, Sukumal N, Grainger MJ, Savini T (2021) Living with human encroachment: Status and distribution of Green Peafowl in northern stronghold of Thailand. Glob Ecol Conserv 28:e01674. https://doi.org/10.1016/J.GECCO.2021.E01674
Segelbacher G, Bosse M, Burger P et al (2021) New developments in the field of genomic technologies and their relevance to conservation management. Conserv Genet 2021 232 23:217–242. https://doi.org/10.1007/S10592-021-01415-5
Sih A, Jonsson BG, Luikart G (2000) Habitat loss: ecological, evolutionary and genetic consequences. Trends Ecol Evol 15:132–134. https://doi.org/10.1016/S0169-5347(99)01799-1
Skotte L, Korneliussen TS, Albrechtsen A (2013) Estimating individual admixture proportions from next generation sequencing data. Genetics 195:693–702. https://doi.org/10.1534/GENETICS.113.154138
Sodhi NS, Koh LP, Brook BW, Ng PKL (2004) Southeast asian biodiversity: an impending disaster. Trends Ecol Evol 19:654–660. https://doi.org/10.1016/J.TREE.2004.09.006
Song S, Sliwerska E, Emery S, Kidd JM (2017) Modeling human population separation history using physically phased genomes. Genetics 205:385–395. https://doi.org/10.1534/genetics.116.192963
Song K, Gao B, Halvarsson P et al (2020) Genomic analysis of demographic history and ecological niche modeling in the endangered chinese Grouse Tetrastes sewerzowi. BMC Genomics 21:1–9. https://doi.org/10.1186/S12864-020-06957-5/FIGURES/4
Sukumal N, McGowan PJK, Savini T (2015) Change in status of green peafowl Pavo muticus (Family Phasianidae) in Southcentral Vietnam: a comparison over 15 years. Glob Ecol Conserv 3:11–19. https://doi.org/10.1016/J.GECCO.2014.10.007
Sukumal N, Dowell SD, Savini T (2017) Micro-habitat selection and population recovery of the Endangered Green Peafowl Pavo muticus in western Thailand: implications for conservation guidance. Bird Conserv Int 27:414–430. https://doi.org/10.1017/S095927091600037X
Sukumal N, Dowell SD, Savini T (2020) Modelling occurrence probability of the endangered green peafowl Pavo muticus in mainland Southeast Asia: applications for landscape conservation and management. Oryx 54:30–39. https://doi.org/10.1017/S003060531900005X
Talla V, Mrazek V, Höglund J, Backström N (2023) Whole genome re-sequencing uncovers significant population structure and low genetic diversity in the endangered clouded apollo (Parnasssius mnemosyne) in Sweden. Conserv Genet 1:1–10. https://doi.org/10.1007/S10592-023-01502-9/FIGURES/4
Teerlink CC, Jurynec MJ, Hernandez R et al (2021) A role for the MEGF6 gene in predisposition to osteoporosis. Ann Hum Genet 85:58–72. https://doi.org/10.1111/AHG.12408
van Balen S, Prawiradilaga DM, Indrawan M (1995) The distribution and status of green peafowl Pavo muticus in Java. Biol Conserv 71:289–297. https://doi.org/10.1016/0006-3207(94)00048-U
Voris HK (2000) Maps of Pleistocene sea levels in Southeast Asia: shorelines, river systems and time durations. J Biogeogr 27:1153–1167. https://doi.org/10.1046/J.1365-2699.2000.00489.X
Wharton D (2008) The future of zoo biology. Zoo Biol 27:498–504. https://doi.org/10.1002/ZOO.20204
Wolf JBW, Ellegren H (2017) Making sense of genomic islands of differentiation in light of speciation. Nat Rev Genet 18:87–100. https://doi.org/10.1038/nrg.2016.133
Woodruff DS (2010) Biogeography and conservation in Southeast Asia: how 2.7 million years of repeated environmental fluctuations affect today’s patterns and the future of the remaining refugial-phase biodiversity. Biodivers Conserv 19:919–941. https://doi.org/10.1007/S10531-010-9783-3/FIGURES/3
Wright AE, Harrison PW, Zimmer F et al (2015) Variation in promiscuity and sexual selection drives avian rate of Faster-Z evolution. Mol Ecol 24:1218–1235. https://doi.org/10.1111/MEC.13113
Wright BR, Farquharson KA, McLennan EA et al (2020) A demonstration of conservation genomics for threatened species management. Mol Ecol Resour 20:1526–1541. https://doi.org/10.1111/1755-0998.13211
Zhang X, Lin C, Li H et al (2022) Chromosome-Level Genome Assembly of the Green Peafowl (Pavo muticus). Genome Biol Evol 14. https://doi.org/10.1093/GBE/EVAC015
Zhou H, Sinsheimer JS, Bates DM et al (2020) OpenMendel: a Cooperative Programming Project for Statistical Genetics. Hum Genet 139:61. https://doi.org/10.1007/S00439-019-02001-Z
Acknowledgements
We want to thank Kermit Blackwood for the extensive discussion regarding the morphological diversity associated with different landscapes within green peafowl. We want to thank the two anonymous reviewers from the first round of peer review for their insightful comments regarding the writing and additional analysis that immensely helped improve the manuscript. The two new anonymous reviewers in the second round of peer review provided critical new perspectives and further enhanced the quality of the manuscript. We thank the Ministry of Human Resource Development for awarding a fellowship to ABP. The Department of Biotechnology, Ministry of Science and Technology, India (Grant no. BT/11/IYBA/2018/03) and Science and Engineering Research Board (Grant no. ECR/2017/001430) provided computational resources (i.e., Har Gobind Khorana Computational Biology cluster).
Author information
Authors and Affiliations
Contributions
ABP analyzed the genomic data and generated all the results. ABP wrote the manuscript with inputs from NV.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Patil, A.B., Vijay, N. Conservation implications of diverse demographic histories: the case study of green peafowl (Pavo muticus, Linnaeus 1766). Conserv Genet 25, 455–468 (2024). https://doi.org/10.1007/s10592-023-01580-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-023-01580-9