Abstract
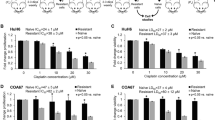
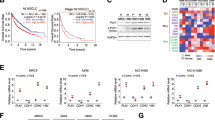
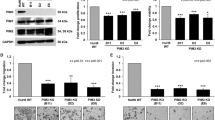
Patients presenting with metastatic hepatoblastoma have limited treatment options and survival rates as low as 25%. We previously demonstrated that Proviral Integration site in Maloney murine leukemia virus 3 (PIM3) kinase promotes tumorigenesis and cancer cell stemness in hepatoblastoma. In this study, we assessed the role of PIM3 kinase in promoting hepatoblastoma metastasis. We utilized a tail vein injection model of metastasis to evaluate the effect of CRISPR/Cas9-mediated PIM3 knockout, stable overexpression of PIM3, and pharmacologic PIM inhibition on the formation of lung metastasis. In vivo studies revealed PIM3 knockout impaired the formation of lung metastasis: 5 out of 6 mice injected with wild type hepatoblastoma cells developed lung metastasis while none of the 7 mice injected with PIM3 knockout hepatoblastoma cells developed lung metastasis. PIM3 overexpression in hepatoblastoma increased the pulmonary metastatic burden in mice and mechanistically, upregulated the phosphorylation and cell surface expression of CXCR4, a key receptor in the progression of cancer cell metastasis. CXCR4 blockade with AMD3100 decreased the metastatic phenotype of PIM3 overexpressing cells, indicating that CXCR4 contributed to PIM3’s promotion of hepatoblastoma metastasis. Clinically, PIM3 expression correlated positively with CXCR4 expression in primary hepatoblastoma tissues. In conclusion, we have shown PIM3 kinase promotes the metastatic phenotype of hepatoblastoma cells through upregulation of CXCR4 cell surface expression and these findings suggest that targeting PIM3 kinase may provide a novel therapeutic strategy for metastatic hepatoblastoma.






Similar content being viewed by others
References
Feng J, Polychronidis G, Heger U, Frongia G, Mehrabi A, Hoffmann K (2019) Incidence trends and survival prediction of hepatoblastoma in children: a population-based study. Cancer Commun (Lond) 39(1):62
Meyers RL, Tiao G, de Ville de Goyet J, Superina R, Aronson DC (2014) Hepatoblastoma state of the art: pre-treatment extent of disease, surgical resection guidelines and the role of liver transplantation. Curr Opin Pediatr 26(1):29–36
Perilongo G, Brown J, Shafford E, Brock P, De Camargo B, Keeling JW et al (2000) Hepatoblastoma presenting with lung metastases: treatment results of the first cooperative, prospective study of the International Society of Paediatric Oncology on childhood liver tumors. Cancer 89(8):1845–1853
Stafman LL, Mruthyunjayappa S, Waters AM, Garner EF, Aye JM, Stewart JE et al (2018) Targeting PIM kinase as a therapeutic strategy in human hepatoblastoma. Oncotarget 9(32):22665–22679
Maryati R, Julson JR, Bownes LV, Quinn CH, Hutchins SC, Williams AP, Markert HR, Beierle AM, Stewart JE, Hjelmeland AB, Mroczek-Musulman E, Beierle EA (2022) Metastatic human hepatoblastoma cells exhibit enhanced tumorigenicity, invasiveness and a stem-cell like phenotype. J of Pediatr Surg. ; in press
Marayati R, Stafman LL, Williams AP, Bownes LV, Quinn CH, Markert HR et al (2021) CRISPR/Cas9-mediated knockout of PIM3 suppresses tumorigenesis and cancer cell stemness in human hepatoblastoma cells.Cancer Gene Ther.
Zheng HC, Tsuneyama K, Takahashi H, Miwa S, Sugiyama T, Popivanova BK et al (2008) Aberrant Pim-3 expression is involved in gastric adenoma-adenocarcinoma sequence and cancer progression. J Cancer Res Clin Oncol 134(4):481–488
Zhuang H, Zhao MY, Hei KW, Yang BC, Sun L, Du X et al (2015) Aberrant expression of pim-3 promotes proliferation and migration of ovarian cancer cells. Asian Pac J Cancer Prev 16(8):3325–3331
Qu Y, Zhang C, Du E, Wang A, Yang Y, Guo J et al (2016) Pim-3 is a Critical Risk Factor in Development and Prognosis of Prostate Cancer. Med Sci Monit 22:4254–4260
Stafman LL, Waldrop MG, Williams AP, Aye JM, Stewart JE, Mroczek-Musulman E et al (2019) The presence of PIM3 increases hepatoblastoma tumorigenesis and tumor initiating cell phenotype and is associated with decreased patient survival. J Pediatr Surg 54(6):1206–1213
Guo Q, Gao BL, Zhang XJ, Liu GC, Xu F, Fan QY et al (2014) CXCL12-CXCR4 Axis Promotes Proliferation, Migration, Invasion, and Metastasis of Ovarian Cancer. Oncol Res 22(5–6):247–258
Santio NM, Eerola SK, Paatero I, Yli-Kauhaluoma J, Anizon F, Moreau P et al (2015) Pim Kinases Promote Migration and Metastatic Growth of Prostate Cancer Xenografts. PLoS ONE 10(6):e0130340
Bialopiotrowicz E, Gorniak P, Noyszewska-Kania M, Pula B, Makuch-Lasica H, Nowak G et al (2018) Microenvironment-induced PIM kinases promote CXCR4-triggered mTOR pathway required for chronic lymphocytic leukaemia cell migration. J Cell Mol Med 22(7):3548–3559
Gillory LA, Stewart JE, Megison ML, Nabers HC, Mroczek-Musulman E, Beierle EA (2013) FAK Inhibition Decreases Hepatoblastoma Survival Both In Vitro and In Vivo. Transl Oncol 6(2):206–215
Chen Z, Ren X, Ren R, Wang Y, Shang J (2021) The combination of G-CSF and AMD3100 mobilizes bone marrow-derived stem cells to protect against cisplatin-induced acute kidney injury in mice.Stem Cell Research & Therapy. ; 12(1)
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S et al (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR et al (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7(3):562–578
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ et al (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28(5):511–515
Kramer A, Green J, Pollard J Jr, Tugendreich S (2014) Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30(4):523–530
Hiyama E, Ueda Y, Kurihara S, Kawashima K, Ikeda K, Morihara N, Fukazawa T, Kanawa M, Hiyama K (2019) Gene expression profiling in hepatoblastoma cases of the Japanese Study Group for Pediatric Liver Tumors-2 (JPLT-2) trial.Eur J Mol Cancer.
Carrillo-Reixach J, Torrens L, Simon-Coma M, Royo L, Domingo-Sabat M, Abril- Fornaguera J et al (2020) Epigenetic footprint enables molecular risk stratification of hepatoblastoma with clinical implications. J Hepatol 73(2):328–341
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132:365–386
Winer J, Jung CK, Shackel I, Williams PM (1999) Development and validation of real-time quantitative reverse transcriptase-polymerase chain reaction for monitoring gene expression in cardiac myocytes in vitro. Anal Biochem 270(1):41–49
Tang L, Zhang H, Zhang B (2019) A note on error bars as a graphical representation of the variability of data in biomedical research: Choosing between standard deviation and standard error of the mean. J Pancreatol 2(3):69–71
Borowicz S, Van Scoyk M, Avasarala S, Karuppusamy Rathinam MK, Tauler J, Bikkavilli RK et al The soft agar colony formation assay.J Vis Exp. 2014(92):e51998
Hatse S, Princen K, Bridger G, De Clercq E, Schols D (2002) Chemokine receptor inhibition by AMD3100 is strictly confined to CXCR4. FEBS Lett 527(1–3):255–262
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100(1):57–70
Welch DR, Hurst DR (2019) Defining the Hallmarks of Metastasis. Cancer Res 79(12):3011–3027
Chambers AF, Groom AC, MacDonald IC (2002) Dissemination and growth of cancer cells in metastatic sites. Nat Rev Cancer 2(8):563–572
Nagle JA, Ma Z, Byrne MA, White MF, Shaw LM (2004) Involvement of insulin receptor substrate 2 in mammary tumor metastasis. Mol Cell Biol 24(22):9726–9735
Martin SS, Ridgeway AG, Pinkas J, Lu Y, Reginato MJ, Koh EY et al (2004) A cytoskeleton- based functional genetic screen identifies Bcl-xL as an enhancer of metastasis, but not primary tumor growth. Oncogene 23(26):4641–4645
Luo H, Sun R, Zheng Y, Huang J, Wang F, Long D et al (2020) PIM3 Promotes the Proliferation and Migration of Acute Myeloid Leukemia Cells. Onco Targets Ther 13:6897–6905
Liu J, Qu X, Shao L, Hu Y, Yu X, Lan P et al (2018) Pim-3 enhances melanoma cell migration and invasion by promoting STAT3 phosphorylation. Cancer Biol Ther 19(3):160–168
Rashid OM, Nagahashi M, Ramachandran S, Dumur CI, Schaum JC, Yamada A et al (2013) Is tail vein injection a relevant breast cancer lung metastasis model? J Thorac Dis 5(4):385–392
Cojoc M, Peitzsch C, Trautmann F, Polishchuk L, Telegeev GD, Dubrovska A (2013) Emerging targets in cancer management: role of the CXCL12/CXCR4 axis. Onco Targets Ther 6:1347–1361
Balkwill F (2004) Cancer and the chemokine network. Nat Rev Cancer 4(7):540–550
Zou YR, Kottmann AH, Kuroda M, Taniuchi I, Littman DR (1998) Function of the chemokine receptor CXCR4 in haematopoiesis and in cerebellar development. Nature 393(6685):595–599
Burger JA, Peled A (2009) CXCR4 antagonists: targeting the microenvironment in leukemia and other cancers. Leukemia 23(1):43–52
Sun YX, Wang J, Shelburne CE, Lopatin DE, Chinnaiyan AM, Rubin MA et al (2003) Expression of CXCR4 and CXCL12 (SDF-1) in human prostate cancers (PCa) in vivo. J Cell Biochem 89(3):462–473
Dang Y, Jiang N, Wang H, Chen X, Gao Y, Zhang X et al (2020) Proto-Oncogene Serine/Threonine Kinase PIM3 Promotes Cell Migration via Modulating Rho GTPase Signaling. J Proteome Res 19(3):1298–1309
Liu B, Wang Z, Li HY, Zhang B, ** B, Li YY (2014) Pim-3 promotes human pancreatic cancer growth by regulating tumor vasculogenesis. Oncol Rep 31(6):2625–2634
Decker S, Finter J, Forde AJ, Kissel S, Schwaller J, Mack TS et al (2014) PIM kinases are essential for chronic lymphocytic leukemia cell survival (PIM2/3) and CXCR4- mediated microenvironmental interactions (PIM1). Mol Cancer Ther 13(5):1231–1245
Wang Q, Diao X, Sun J, Chen Z (2011) Regulation of VEGF, MMP-9 and metastasis by CXCR4 in a prostate cancer cell line. Cell Biol Int 35(9):897–904
Calandra G, McCarty J, McGuirk J, Tricot G, Crocker SA, Badel K et al (2008) AMD3100 plus G-CSF can successfully mobilize CD34 + cells from non-Hodgkin’s lymphoma, Hodgkin’s disease and multiple myeloma patients previously failing mobilization with chemotherapy and/or cytokine treatment: compassionate use data. Bone Marrow Transplant 41(4):331–338
Hira VVV, Van Noorden CJF, Molenaar RJ (2020) CXCR4 Antagonists as Stem Cell Mobilizers and Therapy Sensitizers for Acute Myeloid Leukemia and Glioblastoma? Biology (Basel). ; 9(2)
Abraham M, Biyder K, Begin M, Wald H, Weiss ID, Galun E et al (2007) Enhanced unique pattern of hematopoietic cell mobilization induced by the CXCR4 antagonist 4F- benzoyl-TN14003. Stem Cells 25(9):2158–2166
Abraham M, Pereg Y, Bulvik B, Klein S, Mishalian I, Wald H et al (2017) Single Dose of the CXCR4 Antagonist BL-8040 Induces Rapid Mobilization for the Collection of Human CD34(+) Cells in Healthy Volunteers. Clin Cancer Res 23(22):6790–6801
Sison EA, Magoon D, Li L, Annesley CE, Rau RE, Small D et al (2014) Plerixafor as a chemosensitizing agent in pediatric acute lymphoblastic leukemia: efficacy and potential mechanisms of resistance to CXCR4 inhibition. Oncotarget 5(19):8947–8958
Grundler R, Brault L, Gasser C, Bullock AN, Dechow T, WoetzelS et al Dissection of PIM serine/threonine kinases in FLT3-ITD–induced leukemogenesis reveals PIM1 as regulator of CXCL12-CXCR4–mediated homing and migration.J Exp Med 2009; 206:1957–70
Cortes J, Tamura K, DeAngelo DJ, de Bono J, Lorente D, Minden M et al (2018) Phase I studies of AZD1208, a proviral integration Moloney virus kinase inhibitor in solid and haematological cancers. Br J Cancer 118(11):1425–1433
Gomez-Cuadrado L, Tracey N, Ma R, Qian B, Brunton VG (2017) Mouse models of metastasis: progress and prospects. Dis Model Mech 10(9):1061–1074
Khanna C, Hunter K, Vlodavsky I (2005) Tail vein assay of cancer metastasis. Curr Protoc Cell Biol. 2001;Chap. 19:19 2 1–2 7
Harms JF, Welch DR (2003) MDA-MB-435 human breast carcinoma metastasis to bone. Clin Exp Metastasis 20(4):327–334
Pettaway CA, Pathak S, Greene G, Ramirez E, Wilson MR, Killion JJ et al (1996) Selection of highly metastatic variants of different human prostatic carcinomas using orthotopic implantation in nude mice. Clin Cancer Res 2(9):1627–1636
Meyers RL, Katzenstein HM, Krailo M, McGahren ED 3, Malogolowkin MH (2007) Surgical resection of pulmonary metastatic lesions in children with hepatoblastoma. J Pediatr Surg 42(12):2050–2056
Kuznetsov HS, Marsh T, Markens BA, Castano Z, Greene-Colozzi A, Hay SA et al (2012) Identification of luminal breast cancers that establish a tumor-supportive macroenvironment defined by proangiogenic platelets and bone marrow-derived cells. Cancer Discov 2(12):1150–1165
Kang SY, Halvorsen OJ, Gravdal K, Bhattacharya N, Lee JM, Liu NW et al (2009) Prosaposin inhibits tumor metastasis via paracrine and endocrine stimulation of stromal p53 and Tsp-1. Proc Natl Acad Sci U S A 106(29):12115–12120
Hart IR, Fidler IJ (1980) Cancer invasion and metastasis. Q Rev Biol 55(2):121–142
Acknowledgements
The authors wish to thank Vidya Sagar Hanumanthu from the UAB Comprehensive Flow Cytometry Core and Dr. Michael Crowley from the UAB Genomics Core.
Funding
This work was supported in part by funding from National Institutes of Health (NIH) T32 CA091078 Surgical Oncology Research Training Program (LLS); NIH T32 CA229102 Surgical Oncology Research Training Program (JRJ, LVB); NIH 5T32GM008361 MSTP Training Program (CHQ); Vince Lombardi Cancer Research Fund Bart Starr Award, Hyundai Hope on Wheels, Kaul Pediatric Research Foundation, Sid Strong Foundation, Elaine Roberts Foundation, and Open Hands Overflowing Hearts (EAB). This work utilized the University of Alabama at Birmingham (UAB) Comprehensive Flow Cytometry Core (supported by NIH P30 AR048311 and NIH P30 AI27667), and the Comprehensive Genomics and Preclinical Imaging Cores, shared facilities which are supported by the O’Neal Comprehensive Cancer Center (P30 CA013148).
Author information
Authors and Affiliations
Contributions
R Marayati was involved in study concept and design, development of methodology, data collection, data analysis, and manuscript preparation. J Julson contributed to data analysis and manuscript preparation. LV Bownes, CH Quinn, AM Beierle, HR Markert, SC Hutchins, and JE Stewart contributed with data collection and analysis. LL Stafman developed of the CRISPR/Cas9 PIM3 knockout. DK Crossman performed bioinformatics analyses of the RNA sequencing data. E Mroczek-Musulman was involved in the histology evaluation of lung sections. AB Hjelmeland provided the luciferase labeling of the hepatoblastoma cells. EA Beierle provided senior guidance with study concept and design, data analysis, and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests:
The authors declare no financial conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Raoud Marayati and Janet Julson contributed equally to the work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Marayati, R., Julson, J., Bownes, L.V. et al. PIM3 kinase promotes tumor metastasis in hepatoblastoma by upregulating cell surface expression of chemokine receptor cxcr4. Clin Exp Metastasis 39, 899–912 (2022). https://doi.org/10.1007/s10585-022-10186-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-022-10186-3