Abstract
Cellular plasticity can occur naturally in an organism and is considered an adapting mechanism during the developmental stage. However, abnormal cellular plasticity is observed in different diseased conditions, including cancer. Cancer cell plasticity triggers the stimuli of epithelial-mesenchymal transition (EMT), abnormal epigenetic changes, expression of stem cell factors and implicated signaling pathways, etc., and helps in the maintenance of CSC phenotype. Conversely, CSC maintains the cancer cell plasticity, EMT, and epigenetic plasticity. EMT contributes to increased cell migration and greater diversity within tumors, while epigenetic changes, stem cell factors (OCT4, NANOG, and SOX2), and various signaling pathways allow cancer cells to maintain various phenotypes, giving rise to intra- and inter-tumoral heterogeneity. The intricate relationships between cancer cell plasticity and stem cell factors help the tumor cells adopt drug-tolerant states, evade senescence, and successfully acquire drug resistance with treatment dismissal. Inhibiting molecules/signaling pathways involved in promoting CSCs, cellular plasticity, EMT, and epigenetic plasticity might be helpful for successful cancer therapy management. This review discussed the role of cellular plasticity, EMT, and stem cell factors in tumor initiation, progression, reprogramming, and therapy resistance. Finally, we discussed how the intervention in this axis will help better manage cancers and improve patient survivability.
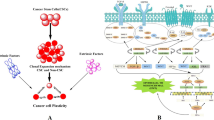
Graphical abstract




Similar content being viewed by others
Abbreviations
- AKT:
-
Protein kinase B
- BCRP:
-
Breast cancer resistance protein
- CSC:
-
Cancer stem cells
- CaP:
-
Prostate cancer
- DVL:
-
Dishevelled
- EGFR:
-
Epidermal growth factor receptor
- ESC:
-
Epithelial stem cells
- EMT:
-
Epithelial-mesenchymal transition
- FOXC1:
-
Forkhead box c1
- FZ:
-
Frizzled
- HIF:
-
Hypoxia-induced factor
- IL:
-
Interleukin
- JAK:
-
Janus kinase
- KLF4:
-
Krüppel-like factor
- KRAS:
-
Kirsten rat sarcoma
- MRP1:
-
Multidrug resistance protein 1
- OCT4:
-
Octamer-binding transcription factor 4
- P-gp:
-
P-glycoprotein
- PI3K:
-
PhosphatidylInositol 3-kinase
- ROS:
-
Reactive oxygen species
- STAT:
-
Signal transducer and activator of transcription
- SMO:
-
Smoothened
- SOX:
-
SRY-related HMG-box
- TGF:
-
Tumor growth factor
- TKI:
-
Tyrosine kinase inhibitor
- ZEB:
-
Zinc finger transcription factor
References
Sung, H., Ferlay, J., Siegel, R. L., Laversanne, M., Soerjomataram, I., Jemal, A., & Bray, F. (2021). Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a Cancer Journal for Clinicians, 71(3), 209–249. https://doi.org/10.3322/caac.21660
Vineis, P., Schatzkin, A., & Potter, J. D. (2010). Models of carcinogenesis: An overview. Carcinogenesis, 31(10), 1703–1709. https://doi.org/10.1093/carcin/bgq087
Hanahan, D., & Weinberg, R. A. (2011). Hallmarks of cancer: The next generation. Cell, 144(5), 646–674. https://doi.org/10.1016/j.cell.2011.02.013
Hanahan, D. (2022). Hallmarks of cancer: New dimensions. Cancer Discovery, 12(1), 31–46. https://doi.org/10.1158/2159-8290.CD-21-1059
Shen, S., & Clairambault, J. (2020). Cell plasticity in cancer cell populations. F1000Research, 9, F1000 Faculty Rev-635. https://doi.org/10.12688/f1000research.24803.1
Ferrao, P. T., Behren, A., Anderson, R. L., & Thompson, E. W. (2015). Editorial: Cellular and phenotypic plasticity in cancer. Frontiers in Oncology, 5, 171. https://doi.org/10.3389/fonc.2015.00171
Herreros-Villanueva, M., Bujanda, L., Billadeau, D. D., & Zhang, J. S. (2014). Embryonic stem cell factors and pancreatic cancer. World Journal of Gastroenterology, 20(9), 2247–2254. https://doi.org/10.3748/wjg.v20.i9.2247
Tang, F., Barbacioru, C., Bao, S., Lee, C., Nordman, E., Wang, X., Lao, K., & Surani, M. A. (2010). Tracing the derivation of embryonic stem cells from the inner cell mass by single-cell RNA-Seq analysis. Cell Stem Cell, 6(5), 468–478. https://doi.org/10.1016/j.stem.2010.03.015
Fatma, H., & Siddique, H. R. (2021). Pluripotency inducing Yamanaka factors: Role in stemness and chemoresistance of liver cancer. Expert Review of Anticancer Therapy, 21(8), 853–864. https://doi.org/10.1080/14737140.2021.1915137
Hadjimichael, C., Chanoumidou, K., Papadopoulou, N., Arampatzi, P., Papamatheakis, J., & Kretsovali, A. (2015). Common stemness regulators of embryonic and cancer stem cells. World journal of stem cells, 7(9), 1150–1184. https://doi.org/10.4252/wjsc.v7.i9.1150
Fatma, H., Siddique, H. R., & Maurya, S. K. (2021). The multiple faces of NANOG in cancer: A therapeutic target to chemosensitize therapy-resistant cancers. Epigenomics, 13(23), 1885–1900. https://doi.org/10.2217/epi-2021-0228
Boumahdi, S., & de Sauvage, F. J. (2020). The great escape: Tumor cell plasticity in resistance to targeted therapy. Nature reviews. Drug Design and Discovery, 19(1), 39–56. https://doi.org/10.1038/s41573-019-0044-1
Jacquemin, V., Antoine, M., Dom, G., Detours, V., Maenhaut, C., & Dumont, J. E. (2022). Dynamic cancer cell heterogeneity: Diagnostic and therapeutic implications. Cancers, 14(2), 280. https://doi.org/10.3390/cancers14020280
Dagogo-Jack, I., & Shaw, A. T. (2018). Tumour heterogeneity and resistance to cancer therapies. Nature reviews. Clinical Oncology, 15(2), 81–94. https://doi.org/10.1038/nrclinonc.2017.166
Cancer Genome Atlas Research Network. (2008). Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature, 455(7216), 1061–1068. https://doi.org/10.1038/nature07385
Murugaesu, N., Wilson, G. A., Birkbak, N. J., Watkins, T., McGranahan, N., Kumar, S., Abbassi-Ghadi, N., Salm, M., Mitter, R., Horswell, S., Rowan, A., Phillimore, B., Biggs, J., Begum, S., Matthews, N., Hochhauser, D., Hanna, G. B., & Swanton, C. (2015). Tracking the genomic evolution of esophageal adenocarcinoma through neoadjuvant chemotherapy. Cancer Discovery, 5(8), 821–831. https://doi.org/10.1158/2159-8290.CD-15-0412
Fanelli, G. N., Naccarato, A. G., & Scatena, C. (2020). Recent advances in cancer plasticity: Cellular mechanisms, surveillance strategies, and therapeutic optimization. Frontiers in Oncology, 10, 569. https://doi.org/10.3389/fonc.2020.00569
Cairns, J. (1975). Mutation selection and the natural history of cancer. Nature, 255(5505), 197–200. https://doi.org/10.1038/255197a0
Nowell, P. C. (1976). The clonal evolution of tumor cell populations. Science, 194(4260), 23–28. https://doi.org/10.1126/science.959840
Greaves, M., & Maley, C. C. (2012). Clonal evolution in cancer. Nature, 481(7381), 306–313. https://doi.org/10.1038/nature10762
McGranahan, N., & Swanton, C. (2017). Clonal heterogeneity and tumor evolution: Past, present, and the future. Cell, 168(4), 613–628. https://doi.org/10.1016/j.cell.2017.01.018
Ahmed, M., & Li, L. C. (2013). Adaptation and clonal selection models of castration-resistant prostate cancer: Current perspective. International Journal of Urology : Official Journal of the Japanese Urological Association, 20(4), 362–371. https://doi.org/10.1111/iju.12005
Vaquero-Siguero, N., Schleussner, N., Volk, J., Mastel, M., Meier, J., & Jackstadt, R. (2022). Modeling colorectal cancer progression reveals niche-dependent clonal selection. Cancers, 14(17), 4260. https://doi.org/10.3390/cancers14174260
Pierce, G. B., & Speers, W. C. (1988). Tumors as caricatures of the process of tissue renewal: Prospects for therapy by directing differentiation. Cancer Research, 48(8), 1996–2004.
Kreso, A., & Dick, J. E. (2014). Evolution of the cancer stem cell model. Cell Stem Cell, 14(3), 275–291. https://doi.org/10.1016/j.stem.2014.02.006
Kise, K., Kinugasa-Katayama, Y., & Takakura, N. (2016). Tumor microenvironment for cancer stem cells. Advanced drug delivery reviews, 99(Pt B), 197–205. https://doi.org/10.1016/j.addr.2015.08.005
van Niekerk, G., Davids, L. M., Hattingh, S. M., & Engelbrecht, A. M. (2017). Cancer stem cells: A product of clonal evolution? International Journal of Cancer, 140(5), 993–999. https://doi.org/10.1002/ijc.30448
Cabrera, M. C., Hollingsworth, R. E., & Hurt, E. M. (2015). Cancer stem cell plasticity and tumor hierarchy. World journal of stem cells, 7(1), 27–36. https://doi.org/10.4252/wjsc.v7.i1.27
Gupta, P. B., Pastushenko, I., Skibinski, A., Blanpain, C., & Kuperwasser, C. (2019). Phenotypic plasticity: Driver of cancer initiation, progression, and therapy resistance. Cell Stem Cell, 24(1), 65–78. https://doi.org/10.1016/j.stem.2018.11.011
Thankamony, A. P., Saxena, K., Murali, R., Jolly, M. K., & Nair, R. (2020). Cancer stem cell plasticity - A deadly deal. Frontiers in Molecular Biosciences, 7, 79. https://doi.org/10.3389/fmolb.2020.00079
Roberts, C. M., Cardenas, C., & Tedja, R. (2019). The Role of intra-tumoral heterogeneity and its clinical relevance in epithelial ovarian cancer recurrence and metastasis. Cancers, 11(8), 1083. https://doi.org/10.3390/cancers11081083
Stanta, G., & Bonin, S. (2018). Overview on clinical relevance of intra-tumor heterogeneity. Frontiers in Medicine, 5, 85. https://doi.org/10.3389/fmed.2018.00085
Boyer, L. A., Lee, T. I., Cole, M. F., Johnstone, S. E., Levine, S. S., Zucker, J. P., Guenther, M. G., Kumar, R. M., Murray, H. L., Jenner, R. G., Gifford, D. K., Melton, D. A., Jaenisch, R., & Young, R. A. (2005). Core transcriptional regulatory circuitry in human embryonic stem cells. Cell, 122(6), 947–956. https://doi.org/10.1016/j.cell.2005.08.020
Cole, M. F., Johnstone, S. E., Newman, J. J., Kagey, M. H., & Young, R. A. (2008). Tcf3 is an integral component of the core regulatory circuitry of embryonic stem cells. Genes & Development, 22(6), 746–755. https://doi.org/10.1101/gad.1642408
Morris, S. A. (2016). Direct lineage reprogramming via pioneer factors; A detour through developmental gene regulatory networks. Development (Cambridge, England), 143(15), 2696–2705. https://doi.org/10.1242/dev.138263
Yang, L., Shi, P., Zhao, G., Xu, J., Peng, W., Zhang, J., Zhang, G., Wang, X., Dong, Z., Chen, F., & Cui, H. (2020). Targeting cancer stem cell pathways for cancer therapy. Signal Transduction and Targeted Therapy, 5(1), 8. https://doi.org/10.1038/s41392-020-0110-5
Karamboulas, C., & Ailles, L. (2013). Developmental signaling pathways in cancer stem cells of solid tumors. Biochimica et Biophysica Acta, 1830(2), 2481–2495. https://doi.org/10.1016/j.bbagen.2012.11.008
Smith, B. A., Balanis, N. G., Nanjundiah, A., Sheu, K. M., Tsai, B. L., Zhang, Q., Park, J. W., Thompson, M., Huang, J., Witte, O. N., & Graeber, T. G. (2018). A human adult stem cell signature marks aggressive variants across epithelial cancers. Cell reports, 24(12), 3353–3366.e5. https://doi.org/10.1016/j.celrep.2018.08.062
Sneha, S., Nagare, R. P., Manasa, P., Vasudevan, S., Shabna, A., & Ganesan, T. S. (2019). Analysis of human stem cell transcription factors. Cell Reports, 21(4), 171–180. https://doi.org/10.1089/cell.2019.0005
Swain, N., Thakur, M., Pathak, J., & Swain, B. (2020). SOX2, OCT4 and NANOG: The core embryonic stem cell pluripotency regulators in oral carcinogenesis. Journal of oral and maxillofacial pathology: JOMFP, 24(2), 368–373. https://doi.org/10.4103/jomfp.JOMFP_22_20
Liu, A., Yu, X., & Liu, S. (2013). Pluripotency transcription factors and cancer stem cells: small genes make a big difference. Chinese Journal of Cancer, 32(9), 483–487. https://doi.org/10.5732/cjc.012.10282
Islam, Z., Ali, A. M., Naik, A., Eldaw, M., Decock, J., & Kolatkar, P. R. (2021). Transcription factors: The fulcrum between cell development and carcinogenesis. Frontiers in Oncology, 11, 681377. https://doi.org/10.3389/fonc.2021.681377
He, Z., He, J., & **e, K. (2023). KLF4 transcription factor in tumorigenesis. Cell Death & Disease, 9(1), 118. https://doi.org/10.1038/s41420-023-01416-y
Qi, X. T., Li, Y. L., Zhang, Y. Q., Xu, T., Lu, B., Fang, L., Gao, J. Q., Yu, L. S., Zhu, D. F., Yang, B., He, Q. J., & Ying, M. D. (2019). KLF4 functions as an oncogene in promoting cancer stem cell-like characteristics in osteosarcoma cells. Acta Pharmacologica Sinica, 40(4), 546–555. https://doi.org/10.1038/s41401-018-0050-6
Ma, C., Wang, F., Han, B., Zhong, X., Si, F., Ye, J., Hsueh, E. C., Robbins, L., Kiefer, S. M., Zhang, Y., Hunborg, P., Varvares, M. A., Rauchman, M., & Peng, G. (2018). SALL1 functions as a tumor suppressor in breast cancer by regulating cancer cell senescence and metastasis through the NuRD complex. Molecular Cancer, 17(1), 78. https://doi.org/10.1186/s12943-018-0824-y
Yang, F., Cui, P., Lu, Y., & Zhang, X. (2019). Requirement of the transcription factor YB-1 for maintaining the stemness of cancer stem cells and reverting differentiated cancer cells into cancer stem cells. Stem Cell Research & Therapy, 10(1), 233. https://doi.org/10.1186/s13287-019-1360-4
Gong, X., Liu, W., Wu, L., Ma, Z., Wang, Y., Yu, S., Zhang, J., **e, H., Wei, G., Ma, F., Lu, L., & Chen, L. (2018). Transcriptional repressor GATA binding 1-mediated repression of SRY-box 2 expression suppresses cancer stem cell functions and tumor initiation. The Journal of Biological Chemistry, 293(48), 18646–18654. https://doi.org/10.1074/jbc.RA118.003983
Safa, A. R., Saadatzadeh, M. R., Cohen-Gadol, A. A., Pollok, K. E., & Bijangi-Vishehsaraei, K. (2015). Glioblastoma stem cells (GSCs) epigenetic plasticity and interconversion between differentiated non-GSCs and GSCs. Genes & diseases, 2(2), 152–163. https://doi.org/10.1016/j.gendis.2015.02.001
Praharaj, P. P., Panigrahi, D. P., Bhol, C. S., Patra, S., Mishra, S. R., Mahapatra, K. K., Behera, B. P., Singh, A., Patil, S., & Bhutia, S. K. (2021). Mitochondrial rewiring through mitophagy and mitochondrial biogenesis in cancer stem cells: A potential target for anti-CSC cancer therapy. Cancer Letters, 498, 217–228. https://doi.org/10.1016/j.canlet.2020.10.036
Patil, K., Khan, F. B., Akhtar, S., Ahmad, A., & Uddin, S. (2021). The plasticity of pancreatic cancer stem cells: Implications in therapeutic resistance. Cancer Metastasis Reviews, 40(3), 691–720. https://doi.org/10.1007/s10555-021-09979-x
Li, Y., Wang, Z., Ajani, J. A., & Song, S. (2021). Drug resistance and cancer stem cells. Cell Communication and Signaling: CCS, 19(1), 19. https://doi.org/10.1186/s12964-020-00627-5
Paul, R., Dorsey, J. F., & Fan, Y. (2022). Cell plasticity, senescence, and quiescence in cancer stem cells: Biological and therapeutic implications. Pharmacology & Therapeutics, 231, 107985. https://doi.org/10.1016/j.pharmthera.2021.107985
Lu, Y., Zhu, Y., Deng, S., Chen, Y., Li, W., Sun, J., & Xu, X. (2021). Targeting the sonic hedgehog pathway to suppress the expression of the cancer stem cell (CSC)-related transcription factors and CSC-driven thyroid tumor growth. Cancers, 13(3), 418. https://doi.org/10.3390/cancers13030418
Fendler, A., Bauer, D., Busch, J., Jung, K., Wulf-Goldenberg, A., Kunz, S., Song, K., Myszczyszyn, A., Elezkurtaj, S., Erguen, B., Jung, S., Chen, W., & Birchmeier, W. (2020). Inhibiting WNT and NOTCH in renal cancer stem cells and the implications for human patients. Nature Communications, 11(1), 929. https://doi.org/10.1038/s41467-020-14700-7
Aparicio, L. A., Blanco, M., Castosa, R., Concha, Á., Valladares, M., Calvo, L., & Figueroa, A. (2015). Clinical implications of epithelial cell plasticity in cancer progression. Cancer Letters, 366(1), 1–10. https://doi.org/10.1016/j.canlet.2015.06.007
Kumar, V. E., Nambiar, R., De Souza, C., Nguyen, A., Chien, J., & Lam, K. S. (2022). Targeting epigenetic modifiers of tumor plasticity and cancer stem cell behavior. Cells, 11(9), 1403. https://doi.org/10.3390/cells1109140357
da Silva-Diz, V., Lorenzo-Sanz, L., Bernat-Peguera, A., Lopez-Cerda, M., & Muñoz, P. (2018). Cancer cell plasticity: Impact on tumor progression and therapy response. Seminars in Cancer Biology, 53, 48–58. https://doi.org/10.1016/j.semcancer.2018.08.009
Yuan, S., Norgard, R. J., & Stanger, B. Z. (2019). Cellular plasticity in cancer. Cancer Discovery, 9(7), 837–851. https://doi.org/10.1158/2159-8290.CD-19-0015
Warrier, N. M., Kelkar, N., Johnson, C. T., Govindarajan, T., Prabhu, V., & Kumar, P. (2023). Understanding cancer stem cells and plasticity: Towards better therapeutics. European Journal of Cell Biology, 102(2), 151321. https://doi.org/10.1016/j.ejcb.2023.151321
Baram, T., Rubinstein-Achiasaf, L., Ben-Yaakov, H., & Ben-Baruch, A. (2021). Inflammation-driven breast tumor cell plasticity: Stemness/EMT, therapy resistance and dormancy. Frontiers in Oncology, 10, 614468. https://doi.org/10.3389/fonc.2020.614468
Guo, H., Zeng, H., Fu, C., Huang, J., Lu, J., Hu, Y., Zhou, Y., Luo, L., Zhang, Y., Zhang, L., Chen, J., & Zeng, Q. (2021). Identification of sitogluside as a potential skin-pigmentation-reducing agent through network pharmacology. Oxidative Medicine and Cellular Longevity, 2021, 4883398. https://doi.org/10.1155/2021/4883398
Ray, T., Ryusaki, T., & Ray, P. S. (2021). Therapeutically targeting cancers that overexpress FOXC1: A transcriptional driver of cell plasticity, partial EMT, and cancer metastasis. Frontiers in Oncology, 11, 721959. https://doi.org/10.3389/fonc.2021.721959
Friedmann-Morvinski, D. (2014). Glioblastoma heterogeneity and cancer cell plasticity. Critical Reviews in Oncogenesis, 19(5), 327–336. https://doi.org/10.1615/critrevoncog.2014011777
Lee, C. C., Lin, J. C., Hwang, W. L., Kuo, Y. J., Chen, H. K., Tai, S. K., Lin, C. C., & Yang, M. H. (2018). Macrophage-secreted interleukin-35 regulates cancer cell plasticity to facilitate metastatic colonization. Nature Communications, 9(1), 3763. https://doi.org/10.1038/s41467-018-06268-0
Chaffer, C. L., San Juan, B. P., Lim, E., & Weinberg, R. A. (2016). EMT, cell plasticity and metastasis. Cancer Metastasis Reviews, 35(4), 645–654. https://doi.org/10.1007/s10555-016-9648-7
Savagner, P. (2015). Epithelial-mesenchymal transitions: from cell plasticity to concept elasticity. Current Topics in Developmental Biology, 112, 273–300. https://doi.org/10.1016/bs.ctdb.2014.11.021
Xu, Y., Lee, D. K., Feng, Z., Xu, Y., Bu, W., Li, Y., Liao, L., & Xu, J. (2017). Breast tumor cell-specific knockout of Twist1 inhibits cancer cell plasticity, dissemination, and lung metastasis in mice. Proceedings of the National Academy of Sciences of the United States of America, 114(43), 11494–11499. https://doi.org/10.1073/pnas.1618091114
Krebs, A. M., Mitschke, J., Lasierra Losada, M., Schmalhofer, O., Boerries, M., Busch, H., Boettcher, M., Mougiakakos, D., Reichardt, W., Bronsert, P., Brunton, V. G., Pilarsky, C., Winkler, T. H., Brabletz, S., Stemmler, M. P., & Brabletz, T. (2017). The EMT-activator Zeb1 is a key factor for cell plasticity and promotes metastasis in pancreatic cancer. Nature Cell Biology, 19(5), 518–529. https://doi.org/10.1038/ncb3513
Zhou, W., Ye, X. L., Xu, J., Cao, M. G., Fang, Z. Y., Li, L. Y., Guan, G. H., Liu, Q., Qian, Y. H., & **e, D. (2017). The lncRNA H19 mediates breast cancer cell plasticity during EMT and MET plasticity by differentially sponging miR-200b/c and let-7b. Science Signaling, 10(483), eaak9557. https://doi.org/10.1126/scisignal.aak9557
Fan, C., Wang, Q., Kuipers, T. B., Cats, D., Iyengar, P. V., Hagenaars, S. C., Mesker, W. E., Devilee, P., Tollenaar, R. A. E. M., Mei, H., & Ten Dijke, P. (2023). LncRNA LITATS1 suppresses TGF-β-induced EMT and cancer cell plasticity by potentiating TβRI degradation. The EMBO journal, 42(10), e112806. https://doi.org/10.15252/embj.2022112806
Lüönd, F., Tiede, S., & Christofori, G. (2021). Breast cancer as an example of tumour heterogeneity and tumour cell plasticity during malignant progression. British Journal of Cancer, 125(2), 164–175. https://doi.org/10.1038/s41416-021-01328-7
Singh, S. K., Chen, N. M., Hessmann, E., Siveke, J., Lahmann, M., Singh, G., Voelker, N., Vogt, S., Esposito, I., Schmidt, A., Brendel, C., Stiewe, T., Gaedcke, J., Mernberger, M., Crawford, H. C., Bamlet, W. R., Zhang, J. S., Li, X. K., Smyrk, T. C., et al. (2015). Antithetical NFATc1-Sox2 and p53-miR200 signaling networks govern pancreatic cancer cell plasticity. The EMBO journal, 34(4), 517–530. https://doi.org/10.15252/embj.201489574
Flavahan, W. A., Gaskell, E., & Bernstein, B. E. (2017). Epigenetic plasticity and the hallmarks of cancer. Science, 357(6348), eaal2380. https://doi.org/10.1126/science.aal2380
Burdziak, C., Alonso-Curbelo, D., Walle, T., Reyes, J., Barriga, F. M., Haviv, D., **e, Y., Zhao, Z., Zhao, C. J., Chen, H. A., Chaudhary, O., Masilionis, I., Choo, Z. N., Gao, V., Luan, W., Wuest, A., Ho, Y. J., Wei, Y., Quail, D. F., et al. (2023). Epigenetic plasticity cooperates with cell-cell interactions to direct pancreatic tumorigenesis. Science, 380(6645), eadd5327. https://doi.org/10.1126/science.add5327
Bedi, U., Mishra, V. K., Wasilewski, D., Scheel, C., & Johnsen, S. A. (2014). Epigenetic plasticity: a central regulator of epithelial-to-mesenchymal transition in cancer. Oncotarget, 5(8), 2016–2029. https://doi.org/10.18632/oncotarget.1875
Anatskaya, O. V., & Vinogradov, A. E. (2022). Polyploidy and Myc proto-oncogenes promote stress adaptation via epigenetic plasticity and gene regulatory network rewiring. International Journal of Molecular Sciences, 23(17), 9691. https://doi.org/10.3390/ijms23179691
De Smedt, E., Lui, H., Maes, K., De Veirman, K., Menu, E., Vanderkerken, K., & De Bruyne, E. (2018). The epigenome in multiple myeloma: Impact on tumor cell plasticity and drug response. Frontiers in Oncology, 8, 566. https://doi.org/10.3389/fonc.2018.00566
Tam, W. L., & Weinberg, R. A. (2013). The epigenetics of epithelial-mesenchymal plasticity in cancer. Nature Medicine, 19(11), 1438–1449. https://doi.org/10.1038/nm.3336
He, R., Xhabija, B., Gopi, L. K., Kurup, J. T., Xu, Z., Liu, Z., & Kidder, B. L. (2022). H3K4 demethylase KDM5B regulates cancer cell identity and epigenetic plasticity. Oncogene, 41(21), 2958–2972. https://doi.org/10.1038/s41388-022-02311-z
Yao, J., Chen, J., Li, L. Y., & Wu, M. (2020). Epigenetic plasticity of enhancers in cancer. Transcription, 11(1), 26–36. https://doi.org/10.1080/21541264.2020.1713682
Fagnocchi, L., Poli, V., & Zippo, A. (2018). Enhancer reprogramming in tumor progression: A new route towards cancer cell plasticity. Cellular and Molecular Life Sciences: CMLS, 75(14), 2537–2555. https://doi.org/10.1007/s00018-018-2820-1
Huang, T., Song, X., Xu, D., Tiek, D., Goenka, A., Wu, B., Sastry, N., Hu, B., & Cheng, S. Y. (2020). Stem cell programs in cancer initiation, progression, and therapy resistance. Theranostics, 10(19), 8721–8743. https://doi.org/10.7150/thno.41648
Huang, J., Liu, K., Song, D., Ding, M., Wang, J., **, Q., & Ni, J. (2016). Krüppel-like factor 4 promotes high-mobility group box 1-induced chemotherapy resistance in osteosarcoma cells. Cancer Science, 107(3), 242–249. https://doi.org/10.1111/cas.12864
Huang, C. P., Tsai, M. F., Chang, T. H., et al. (2013). ALDH-positive lung cancer stem cells confer resistance to epidermal growth factor receptor tyrosine kinase inhibitors. Cancer Letters, 328(1), 144–151. https://doi.org/10.1016/j.canlet.2012.08.021
Shlush, L. I., Mitchell, A., Heisler, L., Abelson, S., Ng, S. W. K., Trotman-Grant, A., Medeiros, J. J. F., Rao-Bhatia, A., Jaciw-Zurakowsky, I., Marke, R., McLeod, J. L., Doedens, M., Bader, G., Voisin, V., Xu, C., McPherson, J. D., Hudson, T. J., Wang, J. C. Y., Minden, M. D., et al. (2017). Tracing the origins of relapse in acute myeloid leukaemia to stem cells. Nature, 547(7661), 104–108. https://doi.org/10.1038/nature22993
Steinbichler, T. B., Dudás, J., Skvortsov, S., Ganswindt, U., Riechelmann, H., & Skvortsova, I. I. (2018). Therapy resistance mediated by cancer stem cells. Seminars in Cancer Biology, 53, 156–167. https://doi.org/10.1016/j.semcancer.2018.11.006
Cojoc, M., Mäbert, K., Muders, M. H., & Dubrovska, A. (2015). A role for cancer stem cells in therapy resistance: Cellular and molecular mechanisms. Seminars in Cancer Biology, 31, 16–27. https://doi.org/10.1016/j.semcancer.2014.06.004
Zhou, H. M., Zhang, J. G., Zhang, X., & Li, Q. (2021). Targeting cancer stem cells for reversing therapy resistance: Mechanism, signaling, and prospective agents. Signal Transduction and Targeted Therapy, 6(1), 62. https://doi.org/10.1038/s41392-020-00430-1
Mohiuddin, I. S., Wei, S. J., & Kang, M. H. (2020). Role of OCT4 in cancer stem-like cells and chemotherapy resistance. Biochimica et biophysica acta. Molecular basis of disease, 1866(4), 165432. https://doi.org/10.1016/j.bbadis.2019.03.005
Liau, B. B., Sievers, C., Donohue, L. K., Gillespie, S. M., Flavahan, W. A., Miller, T. E., Venteicher, A. S., Hebert, C. H., Carey, C. D., Rodig, S. J., Shareef, S. J., Najm, F. J., van Galen, P., Wakimoto, H., Cahill, D. P., Rich, J. N., Aster, J. C., Suvà, M. L., Patel, A. P., et al. (2017). Adaptive chromatin remodeling drives glioblastoma stem cell plasticity and drug tolerance. Cell stem cell, 20(2), 233–246.e7. https://doi.org/10.1016/j.stem.2016.11.003
Rabé, M., Dumont, S., Álvarez-Arenas, A., Janati, H., Belmonte-Beitia, J., Calvo, G. F., Thibault-Carpentier, C., Séry, Q., Chauvin, C., Joalland, N., Briand, F., Blandin, S., Scotet, E., Pecqueur, C., Clairambault, J., Oliver, L., Perez-Garcia, V., Nadaradjane, A., Cartron, P. F., et al. (2020). Identification of a transient state during the acquisition of temozolomide resistance in glioblastoma. Cell Death & Disease, 11(1), 19. https://doi.org/10.1038/s41419-019-2200-2
Sharma, S. V., Lee, D. Y., Li, B., Quinlan, M. P., Takahashi, F., Maheswaran, S., McDermott, U., Azizian, N., Zou, L., Fischbach, M. A., Wong, K. K., Brandstetter, K., Wittner, B., Ramaswamy, S., Classon, M., & Settleman, J. (2010). A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell, 141(1), 69–80. https://doi.org/10.1016/j.cell.2010.02.027
Shen, S., Faouzi, S., Bastide, A., Martineau, S., Malka-Mahieu, H., Fu, Y., Sun, X., Mateus, C., Routier, E., Roy, S., Desaubry, L., André, F., Eggermont, A., David, A., Scoazec, J. Y., Vagner, S., & Robert, C. (2019). An epitranscriptomic mechanism underlies selective mRNA translation remodelling in melanoma persister cells. Nature Communications, 10(1), 5713. https://doi.org/10.1038/s41467-019-13360-6
Sánchez-Danés, A., Larsimont, J. C., Liagre, M., Muñoz-Couselo, E., Lapouge, G., Brisebarre, A., Dubois, C., Suppa, M., Sukumaran, V., Del Marmol, V., Tabernero, J., & Blanpain, C. (2018). A slow-cycling LGR5 tumour population mediates basal cell carcinoma relapse after therapy. Nature, 562(7727), 434–438. https://doi.org/10.1038/s41586-018-0603-3
De Angelis, M. L., Francescangeli, F., La Torre, F., & Zeuner, A. (2019). Stem cell plasticity and dormancy in the development of cancer therapy resistance. Frontiers in Oncology, 9, 626. https://doi.org/10.3389/fonc.2019.00626
Sharma, N., Bhushan, A., He, J., Kaushal, G., & Bhardwaj, V. (2020). Metabolic plasticity imparts erlotinib-resistance in pancreatic cancer by upregulating glucose-6-phosphate dehydrogenase. Cancer & metabolism, 8, 19. https://doi.org/10.1186/s40170-020-00226-5
Lorenzo-Sanz, L., & Muñoz, P. (2019). Tumor-infiltrating immunosuppressive cells in cancer-cell plasticity, tumor progression and therapy response. Cancer microenvironment : official journal of the International Cancer Microenvironment Society, 12(2-3), 119–132. https://doi.org/10.1007/s12307-019-00232-2
Wang, J. Y., Ma, J., Lin, Y. N., Wang, J., Shen, H., Gui, F. M., Han, C., Li, Q. H., Song, Z., & Wang, X. J. (2017). Zhonghua xue ye xue za zhi. Zhonghua Xueyexue Zazhi, 38(3), 192–197. https://doi.org/10.3760/cma.j.issn.0253-2727.2017.03.004
Tanabe, S., Quader, S., Cabral, H., & Ono, R. (2020). Interplay of EMT and CSC in cancer and the potential therapeutic strategies. Frontiers in Pharmacology, 11, 904. https://doi.org/10.3389/fphar.2020.00904
Jeng, K. S., Chang, C. F., & Lin, S. S. (2020). Sonic hedgehog signaling in organogenesis, tumors, and tumor microenvironments. International Journal of Molecular Sciences, 21(3), 758. https://doi.org/10.3390/ijms21030758
Gong, L., Yan, Q., Zhang, Y., Fang, X., Liu, B., & Guan, X. (2019). Cancer cell reprogramming: a promising therapy converting malignancy to benignity. Cancer communications, 39(1), 48. https://doi.org/10.1186/s40880-019-0393-5
Srivastava, D., & DeWitt, N. (2016). In vivo cellular reprogramming: The next generation. Cell, 166(6), 1386–1396. https://doi.org/10.1016/j.cell.2016.08.055
Lin, S. L., Chang, D. C., Chang-Lin, S., Lin, C. H., Wu, D. T., Chen, D. T., & Ying, S. Y. (2008). Mir-302 reprograms human skin cancer cells into a pluripotent ES-cell-like state. RNA, 14(10), 2115–2124. https://doi.org/10.1261/rna.1162708
Acknowledgements
The authors are thankful to Aligarh Muslim University, India, for providing the necessary facilities. HF expresses her sincere gratitude for the fellowship to UGC (F.82-27/2019(SA-III)), India. HRS is grateful to The UGC (Grant no.F.30-377/2017(BSR)) and DST-SERB (Grant no. EMR/2017/001758), New Delhi, India, for providing financial help.
Author information
Authors and Affiliations
Contributions
Collection of literature and writing the manuscript: H. Fatma and H.R. Siddique.
Concept and Supervision: H.R. Siddique.
Corresponding author
Ethics declarations
Conflict of interest
The author declares no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fatma, H., Siddique, H.R. Cancer cell plasticity, stem cell factors, and therapy resistance: how are they linked?. Cancer Metastasis Rev 43, 423–440 (2024). https://doi.org/10.1007/s10555-023-10144-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-023-10144-9