Abstract
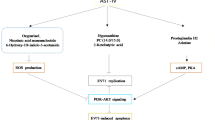
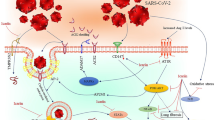
The advent of the new coronavirus, leading to the SARS-CoV-2 pandemic, has presented a substantial worldwide health hazard since its inception in the latter part of 2019. The severity of the current pandemic is exacerbated by the occurrence of re-infection or co-infection with SARS-CoV-2. Hence, comprehending the molecular process underlying the pathophysiology of sepsis and discerning possible molecular targets for therapeutic intervention holds significant importance. For the first time, 31 metabolites were tentatively identified by GC–MS analysis from Alpinia malaccensis. On the other hand, five phenolic compounds were identified and quantified from the plant in HPLC–DAD analysis, including (-) epicatechin, rutin hydrate, rosmarinic acid, quercetin, and kaempferol. Nine GC–MS and five HPLC-identified metabolites had shown interactions with 45 and 30 COVID-19-associated human proteins, respectively. Among the proteins, PARP1, FN1, PRKCA, EGFR, ALDH2, AKR1C3, AHR, and IKBKB have been found as potential therapeutic targets to mitigate SARS-CoV-2 infection. KEGG pathway analysis also showed a strong association of FN1, EGFR, and IKBKB genes with SARS-CoV-2 viral replication and cytokine overexpression due to viral infection. Protein–protein interaction (PPI) analysis also showed that TP53, MMP9, FN1, EGFR, and NOS2 proteins are highly related to the genes involved in COVID-19 comorbidity. These proteins showed interaction with the plant phytoconstituents as well. As the study offers a robust network-based procedure for identifying biomolecules relevant to COVID-19 disease, A. malaccensis could be a good source of effective therapeutic agents against COVID-19 and related viral diseases.





Similar content being viewed by others
References
Afroz N, Ahsanul Hoq M, Jahan S et al (2020) Methanol soluble fraction of fruits of Annona muricata possesses significant antidiarrheal activities. Heliyon 6:e03112. https://doi.org/10.1016/j.heliyon.2019.e03112
Alves-Silva JM, Romane A, Efferth T, Salgueiro L (2017) North African medicinal plants traditionally used in cancer therapy. Front Pharmacol. https://doi.org/10.3389/fphar.2017.00383
Amanat M, Reza MS, Shuvo MSR et al (2021) Zingiber roseum Rosc rhizome: a rich source of hepatoprotective polyphenols. Biomed Pharmacother. https://doi.org/10.1016/j.biopha.2021.111673
Amé JC, Spenlehauer C, De Murcia G (2004) The PARP superfamily. BioEssays 26:882–893
Barabási AL, Oltvai ZN (2004) Network biology: understanding the cell’s functional organization. Nat Rev Genet 5:101–113. https://doi.org/10.1038/nrg1272
Beerli C, Yakimovich A, Kilcher S et al (2019) Vaccinia virus hijacks EGFR signalling to enhance virus spread through rapid and directed infected cell motility. Nat Microbiol 4:216–225
Belpaire FM, Bogaert MG (1996) Cytochrome P450: genetic polymorphism and drug interactions. Acta Clin Belg 51:254–260. https://doi.org/10.1080/22953337.1996.11718518
Carabelli AM, Peacock TP, Thorne LG et al (2023) SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat Rev Microbiol 21:162–177. https://doi.org/10.1038/s41579-022-00841-7
Carolina D, Couto AE, Campos LC et al (2021) MMP-2 and MMP-9 levels in plasma are altered and associated with mortality in COVID-19 patients. Biomed Pharmacother 142:112067. https://doi.org/10.1016/j.biopha.2021.112067
Chang HL, Chen YH, Taiwan HC, Yang CJ (2020) EGFR tyrosine kinase inhibitor-associated interstitial lung disease during the coronavirus disease 2019 pandemic. J Thorac Oncol 15:e129–e131. https://doi.org/10.1016/j.jtho.2020.04.029
Chaudhuri AG, Tiwary BK, Chakrabarti N (2020) Matrix metallopeptidase 9 as a host protein target of chloroquine and melatonin for immunoregulation in COVID-19: a network-based meta-analysis. Life Sci 257:118096. https://doi.org/10.1016/j.lfs.2020.118096
Chen Y, Tian Y, Gao Y et al (2020) In silico design of novel HIV-1 NNRTIs based on combined modeling studies of Dihydrofuro[3,4-d] pyrimidines. Front Chem 8:164. https://doi.org/10.3389/fchem.2020.00164
Coperchini F, Chiovato L, Croce L et al (2020) The cytokine storm in COVID-19: an overview of the involvement of the chemokine/chemokine-receptor system. Cytokine Growth Factor Rev 53:25–32. https://doi.org/10.1016/j.cytogfr.2020.05.003
Daina A, Michielin O, Zoete V (2017) SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
D’Amours D, Desnoyers S, D’Silva I et al (1999) Poly (ADP-ribosyl) ation reactions in the regulation of nuclear functions. Biochem J 342:249–268
Dash S, Dash C, Pandhare J (2021) Therapeutic significance of microRNA-mediated regulation of PARP-1 in SARS-CoV-2 infection. Non-Coding RNA 7:60
Davis AP, Grondin CJ, Johnson RJ et al (2019) The comparative toxicogenomics database: update 2019. Nucleic Acids Res 47:D948–D954. https://doi.org/10.1093/nar/gky868
Di Sotto A, Gullì M, Acquaviva A et al (2022) Phytochemical and pharmacological profiles of the essential oil from the inflorescences of the Cannabis sativa L. Ind Crops Prod 183:114980
Dolan ME, Hill DP, Mukherjee G et al (2020) Investigation of COVID-19 comorbidities reveals genes and pathways coincident with the SARS-CoV-2 viral disease. Sci Rep 10:20848. https://doi.org/10.1038/s41598-020-77632-8
Fishilevich S, Zimmerman S, Kohn A et al (2016) Genic insights from integrated human proteomics in genecards. Database 2016:1–17. https://doi.org/10.1093/database/baw030
Fu Y, Fang Y, Gong S et al (2023) Deep learning-based network pharmacology for exploring the mechanism of licorice for the treatment of COVID-19. Sci Rep 13:5844. https://doi.org/10.1038/s41598-023-31380-7
Gavriatopoulou M, Korompoki E, Fotiou D et al (2020) Organ-specific manifestations of COVID-19 infection. Clin Exp Med 20:493–506. https://doi.org/10.1007/s10238-020-00648-x
Giovannoni F, Li Z, Garcia CC, Quintana FJ (2020) A potential role for AHR in SARS-CoV-2 pathology. Res Sq. https://doi.org/10.21203/rs.3.rs-25639/v1
Gomathi D, Kalaiselvi M, Ravikumar G et al (2015) GC-MS analysis of bioactive compounds from the whole plant ethanolic extract of Evolvulus alsinoides (L.) L. J Food Sci Technol 52:1212–1217. https://doi.org/10.1007/s13197-013-1105-9
Hacker K, Maas R, Kornhuber J, Fromm MF, Zolk O (2015) Substrate-dependent inhibition of the human organic cation transporter OCT2: a comparison of metformin with experimental substrates. PloS one 10(9):e0136451
Harford JB, Kim SS, Pirollo KF et al (2022) TP53 gene therapy as a potential treatment for patients with COVID-19. Viruses 14(4):739. https://doi.org/10.3390/v14040739
Hazra A, Collison M, Pisano J, Kumar M, Oehler C, Ridgway JP (2020) Coinfections with SARS-CoV-2 and other respiratory pathogens. Infect Control Hosp Epidemiol 41(10):1228–1229. https://doi.org/10.1017/ice.2020.322
Huang YF, Bai C, He F et al (2020) Review on the potential action mechanisms of Chinese medicines in treating coronavirus disease 2019 (COVID-19). Pharmacol Res 158:104939. https://doi.org/10.1016/j.phrs.2020.104939
Ison MG, Wolfe C, Boucher H (2020) Emergency use authorization for intravenous peramivir: the need for a transparent distribution process. JAMA 323:2365–2366. https://doi.org/10.1093/cid/cis351
Kaul TN, Middleton E, Ogra PL (1985) Antiviral effect of flavonoids on human viruses. J Med Virol 15:71–79. https://doi.org/10.1002/jmv.1890150110
Kaushik D, Kumar A, Kaushik P, Rana AC (2012) Analgesic and anti-inflammatory activity of pinus roxburghii sarg. Adv Pharmacol Sci. https://doi.org/10.1155/2012/245431
Klann K, Bojkova D, Tascher G et al (2020) Growth factor receptor signaling inhibition prevents SARS-CoV-2 replication. Mol Cell 80(164–174):e164
Kuleshov MV, Jones MR, Rouillard AD et al (2016) Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 44:W90–W97. https://doi.org/10.1093/nar/gkw377
Kushner P, McCarberg BH, Grange L et al (2022) The use of non-steroidal anti-inflammatory drugs (NSAIDs) in COVID-19. NPJ Prim Care Respir Med. https://doi.org/10.1038/s41533-022-00300-z
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26. https://doi.org/10.1016/j.addr.2012.09.019
Liu Y, Lv J, Liu J et al (2020) Mucus production stimulated by IFN-AhR signaling triggers hypoxia of COVID-19. Cell Res 30:1078–1087
Low Z, Lani R, Tiong V et al (2023) COVID-19 therapeutic potential of natural products. Int J Mol Sci 24:9589. https://doi.org/10.3390/ijms24119589
Mazumder T, Hasan T, Ahmed KS et al (2022) Phenolic compounds and extracts from Crotalaria calycina Schrank potentially alleviate pain and inflammation through inhibition of cyclooxygenase-2: an in vivo and molecular dynamics studies. Heliyon 8:e12368. https://doi.org/10.1016/j.heliyon.2022.e12368
Miller R, Wentzel A, Richards G (2020) COVID-19: NAD+ deficiency may predispose the aged, obese and type2 diabetics to mortality through its effect on SIRT1 activity. Med Hypotheses 144:110044
Mueller AL, McNamara MS, Sinclair DA (2020) Why does COVID-19 disproportionately affect older people? Aging (albany NY) 12:9959
Pathan M, Keerthikumar S, Ang CS et al (2015) FunRich: An open access standalone functional enrichment and interaction network analysis tool. Proteomics 15:2597–2601. https://doi.org/10.1002/pmic.201400515
Sakle NS, More SA, Mokale SN (2020) A network pharmacology-based approach to explore potential targets of Caesalpinia pulcherima: an updated prototype in drug discovery. Sci Rep 10:17217. https://doi.org/10.1038/s41598-020-74251-1
Schreiber V, Amé J-C, Dollé P et al (2002) Poly (ADP-ribose) polymerase-2 (PARP-2) is required for efficient base excision DNA repair in association with PARP-1 and XRCC1. J Biol Chem 277:23028–23036
Schreiber V, Amé J-C et al (2002) Poly (ADP-ribose) polymerase-2 (PARP-2) is required for efficient base excision DNA repair in association with PARP-1 and XRCC1. J Biol Chem. https://doi.org/10.1074/jbc.M202390200
Seah I, Agrawal R (2020) Can the coronavirus disease 2019 (COVID-19) affect the eyes? a review of coronaviruses and ocular implications in humans and animals. Ocul Immunol Inflamm 28:391–395. https://doi.org/10.1080/09273948.2020.1738501
Sethi S, Prakash O, Pant AK, Kumar M (2017) phytochemical analysis and pharmacological activities of methanolic extract and essential oil from rhizomes of Alpinia malaccensis (Burm. f.) Roscoe. J Essent Oil-Bearing Plants 20:1018–1029. https://doi.org/10.1080/0972060X.2017.1374216
Sevrioukova IF, Poulos TL (2015) Current approaches for investigating and predicting cytochrome P450 3A4-ligand interactions. Adv Exp Med Biol 851:83–105. https://doi.org/10.1007/978-3-319-16009-2_3
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models. Genome Res 23:2499–2504. https://doi.org/10.1101/gr.1239303.metabolite
Shi XQ, Yue SJ, Tang YP et al (2019) A network pharmacology approach to investigate the blood enriching mechanism of Danggui buxue Decoction. J Ethnopharmacol 235:227–242. https://doi.org/10.1016/j.jep.2019.01.027
Singh M, de Wit EJCRM (2022) Antiviral agents for the treatment of COVID-19: Progress and challenges. Cell Rep Med 3(3):100549
Somarathna T, Fernando WMADB, Ranaweera KKDS et al (2018) Antimicrobial activity and phytochemical screening of Alpinia malaccensis (Ran-kiriya) against food-borne bacteria. J Appl Microbiol 125:1276–1285. https://doi.org/10.1111/jam.14039
Ueki IF, Min-Oo G, Kalinowski A et al (2013) Respiratory virus–induced EGFR activation suppresses IRF1-dependent interferon λ and antiviral defense in airway epithelium. J Exp Med 210:1929–1936
Van HT, Thang TD, Luu TN, Doan VD (2021) An overview of the chemical composition and biological activities of essential oils from: Alpinia genus (Zingiberaceae). RSC Adv 11:37767–37783. https://doi.org/10.1039/d1ra07370b
Von Mering C, Huynen M, Jaeggi D et al (2003) STRING: a database of predicted functional associations between proteins. Nucleic Acids Res 31:258–261. https://doi.org/10.1093/nar/gkg034
Wang H, Paulson KR, Pease SA et al (2022) Estimating excess mortality due to the COVID-19 pandemic: a systematic analysis of COVID-19-related mortality, 2020–21. Lancet 399:1513–1536. https://doi.org/10.1016/S0140-6736(21)02796-3
Warde-Farley D, Donaldson SL, Comes O et al (2010) The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res 38:W214–W220. https://doi.org/10.1093/nar/gkq537
Wicik Z, Eyileten C, Jakubik D et al (2020) ACE2 interaction networks in COVID-19: a physiological framework for prediction of outcome in patients with cardiovascular risk factors. J Clin Med 9:3743. https://doi.org/10.3390/jcm9113743
Wruck W, Adjaye J (2020) SARS-CoV-2 receptor ACE2 is co-expressed with genes related to transmembrane serine proteases, viral entry, immunity and cellular stress. Sci Rep 10:21415. https://doi.org/10.1038/s41598-020-78402-2
Xu J, Xu X, Jiang L et al (2020) SARS-CoV-2 induces transcriptional signatures in human lung epithelial cells that promote lung fibrosis. Respir Res 21:1–12
Yang N, Shen HM (2020) Targeting the endocytic pathway and autophagy process as a novel therapeutic strategy in COVID-19. Int J Biol Sci 16:1724–1731. https://doi.org/10.7150/ijbs.45498
Yarden Y (2001) The EGFR family and its ligands in human cancer: signalling mechanisms and therapeutic opportunities. Eur J Cancer 37:3–8
Zhang W, Zhao Y, Zhang F et al (2020) The use of anti-inflammatory drugs in the treatment of people with severe coronavirus disease 2019 (COVID-19): the experience of clinical immunologists from China. Clin Immunol 214:108393. https://doi.org/10.1016/j.clim.2020.108393
Zhou G, Soufan O, Ewald J et al (2019) NetworkAnalyst 3.0: a visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res 47:W234–W241. https://doi.org/10.1093/nar/gkz240
Funding
This study did not receive any specific funding from government, commercial, or non-profit funding bodies.
Author information
Authors and Affiliations
Contributions
E.J, S.J.S. and T.M: Methodology, Software, Data curation, Writing- Original draft preparation. K.S.A and H.H: HPLC-DAD analysis, Software, Validation. M.A, T.H. and I.J.L: Molecular docking, Software, Data curation, Visualization. S.R. and S.D: Conceptualization, Supervision, Writing- Reviewing and Editing. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflicts of interest
There is no conflict of interest declared by the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jahan, E., Mazumder, T., Hasan, T. et al. Metabolomic Approach to Identify the Potential Metabolites from Alpinia malaccensis for Treating SARS-CoV-2 Infection. Biochem Genet (2024). https://doi.org/10.1007/s10528-024-10869-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10528-024-10869-4