Abstract
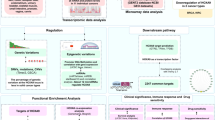
Breast cancer is the most prevalent cancer in female patients worldwide. Tissue factor pathway inhibitor 2 (TFPI-2) is identified as an important tumor suppressor in various cancers. Recent studies have shown that TFPI-2 translocates into the nucleus, where it modulates the transcription of the matrix metalloproteinase-2 (MMP-2) gene. However, its biological role and molecular mechanisms in the progression of breast cancer remain unclear. In this study, we identified 5125 differentially expressed genes (DEGs) from RNA sequencing (RNA-seq) in TFPI-2-overexpressing MDA231 cells compared with control cells. Gene ontology and Kyoto encyclopedia of genes and genomes (KEGG) analysis shown that cell cycle, cell differentiation, proteoglycans in cancer, and pathways associated with cancer were highly enriched in downregulated DEGs. Integration of the RNA-seq and ChIP-sequencing (ChIP-seq) data identified 73 genes directly controlled by TFPI-2 in MDA231 cells. Among them, melanocyte inducing transcription factor (MITF) gene expression was repressed by TFPI-2, which was further verified by a luciferase reporter assay and ChIP-quantitative PCR. Our study provides evidence of a novel role of TFPI-2 in human breast cancer involving targeting of the MITF.




Similar content being viewed by others
Data Availability
The raw sequence data of ChIP-seq and RNA-seq were obtained in SAR(BioProject ID: PRJNA871386).
References
Azamjah N, Soltan-Zadeh Y, Zayeri F (2019) Global trend of breast cancer mortality rate: a 25-year study. Asian Pac J Cancer Prev 20(7):2015–2020. https://doi.org/10.31557/APJCP.2019.20.7.2015
Bertolotto C, Lesueur F, Giuliano S, Strub T, de Lichy M, Bille K, Dessen P, d’Hayer B, Mohamdi H, Remenieras A, Maubec E, de la Fouchardière A, Molinié V, Vabres P, Dalle S, Poulalhon N, Martin-Denavit T, Thomas L, Andry-Benzaquen P, Dupin N et al (2011) A SUMOylation-defective MITF germline mutation predisposes to melanoma and renal carcinoma. Nature 480(7375):94–98. https://doi.org/10.1038/nature10539
Coazzoli M, Napoli A, Roux-Biejat P, Palma C, Moscheni C, Catalani E, Zecchini S, Conte V, Giovarelli M, Caccia S, Procacci P, Cervia D, Clementi E, Perrotta C (2020) Acid sphingomyelinase downregulation enhances mitochondrial fusion and promotes oxidative metabolism in a mouse model of melanoma. Cells 9(4):848. https://doi.org/10.3390/cells9040848
Feng C, Ho Y, Sun C, **a G, Ding Q, Gu B (2017) Epigallocatechin gallate inhibits the growth and promotes the apoptosis of bladder cancer cells. Exp Ther Med 14(4):3513–3518. https://doi.org/10.3892/etm.2017.4981
Gaston-Massuet C, Andoniadou C-L, Signore M, Jayakody S-A, Charolidi N, Kyeyune R, Vernay B, Jacques T-S, Taketo M-M, Le Tissier P, Dattani M-T, Martinez-Barbera J-P (2011) Increased wingless (Wnt) signaling in pituitary progenitor/stem cells gives rise to pituitary tumors in mice and humans. Proc Natl Acad Sci USA 108(28):11482–11487. https://doi.org/10.1073/pnas.1101553108
Hamamoto J, Soejima K, Naoki K, Yasuda H, Hayashi Y, Yoda S, Nakayama S, Satomi R, Terai H, Ikemura S, Sato T, Arai D, Ishioka K, Ohgino K, Betsuyaku T (2015) Methylation-induced downregulation of TFPI-2 causes TMPRSS4 overexpression and contributes to oncogenesis in a subset of non-small-cell lung carcinoma. Cancer Sci 106(1):34–42. https://doi.org/10.1111/cas.12569
Haraksingh R-R, Snyder M-P (2013) Impacts of variation in the human genome on gene regulation. J Mol Biol 425(21):3970–3977. https://doi.org/10.1016/j.jmb.2013.07.015
Herman M-P, Sukhova GK, Kisiel W, Foster D, Kehry M-R, Libby P, Schönbeck U (2001) Tissue factor pathway inhibitor-2 is a novel inhibitor of matrix metalloproteinases with implications for atherosclerosis. J Clin Investig 107(9):1117–1126. https://doi.org/10.1172/JCI10403
Kasetty G, Smeds E, Holmberg E, Wrange L, Adikesavan S, Papareddy P (2016) Vertebrate TFPI-2 C-terminal peptides exert therapeutic applications against Gram-negative infections. BMC Microbiol 16(1):129. https://doi.org/10.1186/s12866-016-0750-3
Kempaiah P, Chand H-S, Kisiel W (2009) Human tissue factor pathway inhibitor-2 is internalized by cells and translocated to the nucleus by the importin system. Arch Biochem Biophys 482(1–2):58–65. https://doi.org/10.1016/j.abb.2008.11.028
Kim N, Kim S, Lee M-W, Jeon H-J, Ryu H, Kim J-M, Lee H-J (2021) MITF promotes cell growth, migration and invasion in clear cell renal cell carcinoma by activating the RhoA/YAP signal pathway. Cancers 13(12):2920. https://doi.org/10.3390/cancers13122920
Kitami K, Yoshihara M, Koya Y, Sugiyama M, Iyoshi S, Uno K, Mogi K, Tano S, Fujimoto H, Nawa A, Kikkawa F, Kajiyama H (2020) Microphthalmia-associated transcription factor-dependent melanoma cell adhesion molecule activation promotes peritoneal metastasis of ovarian cancer. Int J Mol Sci 21(24):9776. https://doi.org/10.3390/ijms21249776
Liao Y, Smyth G-K, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics (oxford, England) 30(7):923–930. https://doi.org/10.1093/bioinformatics/btt656
Love M-I, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Ng L-F, Kaur P, Bunnag N, Suresh J, Sung I-C-H, Tan Q-H, Gruber J, Tolwinski N-S (2019) WNT signaling in disease. Cells 8(8):826. https://doi.org/10.3390/cells8080826
Nicolini A, Ferrari P, Duffy M-J (2018) Prognostic and predictive biomarkers in breast cancer: past, present and future. Semin Cancer Biol 52(Pt 1):56–73. https://doi.org/10.1016/j.semcancer.2017.08.010
Nooron N, Ohba K, Takeda K, Shibahara S, Chiabchalard A (2017) Dysregulated expression of MITF in subsets of hepatocellular carcinoma and cholangiocarcinoma. Tohoku J Exp Med 242(4):291–302. https://doi.org/10.1620/tjem.242.291
Ota Y, Koizume S, Nakamura Y, Yoshihara M, Takahashi T, Sato S, Myoba S, Ohtake N, Kato H, Yokose T, Miyagi E, Miyagi Y (2021) Tissue factor pathway inhibitor-2 is specifically expressed in ovarian clear cell carcinoma tissues in the nucleus, cytoplasm and extracellular matrix. Oncol Rep 45(3):1023–1032. https://doi.org/10.3892/or.2021.7944
Ploper D, Taelman V-F, Robert L, Perez B-S, Titz B, Chen H-W, Graeber T-G, von Euw E, Ribas A, De Robertis E-M (2015) MITF drives endolysosomal biogenesis and potentiates Wnt signaling in melanoma cells. Proc Natl Acad Sci USA 112(5):E420–E429. https://doi.org/10.1073/pnas.1424576112
Rao C-N, Cook B, Liu Y, Chilukuri K, Stack M-S, Foster D-C, Kisiel W, Woodley D-T (1998) HT-1080 fibrosarcoma cell matrix degradation and invasion are inhibited by the matrix-associated serine protease inhibitor TFPI-2/33 kDa MSPI. Int J Cancer 76(5):749–756. https://doi.org/10.1002/(SICI)1097-0215(19980529)76:5%3c749::AID-IJC21%3e3.0.CO;2-Y
Rollin J, Régina S, Vourc’h P, Iochmann S, Bléchet C, Reverdiau P, Gruel Y (2007) Influence of MMP-2 and MMP-9 promoter polymorphisms on gene expression and clinical outcome of non-small cell lung cancer. Lung Cancer (amsterdam, Netherlands) 56(2):273–280. https://doi.org/10.1016/j.lungcan.2006.11.021
So K-K, Peng X-L, Sun H, Wang H (2017) Whole genome chromatin IP-sequencing (ChIP-Seq) in skeletal muscle cells. Methods Mol Biol (clifton, N.J.) 1668:15–25. https://doi.org/10.1007/978-1-4939-7283-8_2
Sung H, Ferlay J, Siegel R-L, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Wang G, Zeng Y, Chen S, Li D, Li W, Zhou Y, Singer R-H, Gu W (2017) Localization of TFPI-2 in the nucleus modulates MMP-2 gene expression in breast cancer cells. Sci Rep 7(1):13575. https://doi.org/10.1038/s41598-017-14148-8
Wang G, **e B, Su Y, Gu Q, Hao D, Liu H, Wang C, Hu Y, Zhang M (2021) Expression analysis of tissue factor pathway inhibitors TFPI-1 and TFPI-2 in Paralichthys olivaceus and antibacterial and anticancer activity of derived peptides. Vet Res 52(1):32. https://doi.org/10.1186/s13567-021-00908-y
Yamashita R, Wakaguri H, Sugano S, Suzuki Y, Nakai K (2010) DBTSS provides a tissue specific dynamic view of transcription start sites. Nucleic Acids Res 38:D98–D104. https://doi.org/10.1093/nar/gkp1017
Acknowledgements
We are indebted to Dr. Wei Gu for providing us with MDA231 cells and technical advice.
Funding
This research was supported by funding from the Guangxi Natural Science Foundation (Grant: 2018JJB140005) and Guangxi Bagui Scholars Special Project.
Author information
Authors and Affiliations
Contributions
GW designed the study and final manuscript revisions, and obtained funding. NZ helped GW in designing for the study. GZ performed experiments, and wrote the manuscript. GZ, WY, and BT performed some of the study. CZ, MK, and ZN supervised and directed the study. RC completed the data curation.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare no competing interests or any competing financial or non-financial interests.
Ethical Approval
The Ethics Committee at the Affiliated Hospital of Guilin Medical School reviewed and approved this research (YJSLL2021104).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, G., Zhang, G., Zhu, N. et al. Integrative analyses of RNA-seq and ChIP-seq Reveal MITF as a Target Gene of TFPI-2 in MDA231 Cells. Biochem Genet 61, 1745–1757 (2023). https://doi.org/10.1007/s10528-023-10340-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-023-10340-w