Abstract
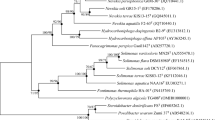
A Gram-negative, aerobic, non-motile and rod-shaped or ovoid bacterial strain, designated D1-W8T, was isolated from a tidal flat on the South Sea in South Korea. Strain D1-W8T was found to grow optimally at 25 °C, at pH 7.0–8.0 and in the presence of 2.0–3.0 % (w/v) NaCl. Neighbour-joining, maximum-likelihood and maximum-parsimony phylogenetic trees based on 16S rRNA gene sequences revealed that strain D1-W8T clustered with the type strain of Pelagicola litoralis showing 97.1 % sequence identity. 16S rRNA gene sequences of the type strains of other species exhibited lower similarity values. Strain D1-W8T was determined to contain Q-10 as the predominant ubiquinone and C18:1 ω7c as the predominant fatty acid. The major polar lipids of strain D1-W8T were identified as phosphatidylcholine, phosphatidylglycerol, phosphatidylethanolamine, one unidentified aminolipid and one unidentified lipid. The DNA G+C content of strain D1-W8T was determined to be 57.9 mol% and its DNA–DNA relatedness value with the type strain of P. litoralis was 17 %. The differential phenotypic properties, together with the phylogenetic and genetic distinctiveness, revealed that strain D1-W8T is separate from P. litoralis. On the basis of the data presented, strain D1-W8T is considered to represent a novel species of the genus Pelagicola, for which the name Pelagicola litorisediminis sp. nov. is proposed. The type strain is D1-W8T (= KCTC 32327T = CECT 8287T).


Similar content being viewed by others
References
Barrow GI, Feltham RKA (1993) Cowan and steel’s manual for the identification of medical bacteria, 3rd edn. Cambridge University Press, Cambridge
Baumann P, Baumann L (1981) The marine Gram-negative eubacteria: genera Photobacterium, Beneckea, Alteromonas, Pseudomonas, and Alcaligenes. In: Starr MP, Stolp H, Trüper HG, Balows A, Schlegel HG (eds) The prokaryotes. Springer, Berlin, pp 1302–1331
Bruns A, Rohde M, Berthe-Corti L (2001) Muricauda ruestringensis gen. nov., sp. nov., a facultatively anaerobic, appendaged bacterium from German North Sea intertidal sediment. Int J Syst Evol Microbiol 51:1997–2006
Buchan A, González JM, Moran MA (2005) Overview of the marine Roseobacter lineage. Appl Environ Microbiol 71:5665–5677
Cohen-Bazire G, Sistrom WR, Stanier RY (1957) Kinetic studies of pigment synthesis by nonsulfur purple bacteria. J Cell Comp Physiol 49:25–68
Ezaki T, Hashimoto Y, Yabuuchi E (1989) Fluorometric deoxyribonucleic acid-deoxyribonucleic acid hybridization in microdilution wells as an alternative to membrane filter hybridization in which radioisotopes are used to determine genetic relatedness among bacterial strains. Int J Syst Bacteriol 39:224–229
Garrity GM, Bell JA, Lilburn T (2005) Order III. Rhodobacterales ord. nov. In: Brenner DJ, Krieg NR, Staley JT, Garrity GM (eds) Bergey’s manual of systematic bacteriology, vol. 2, part C, 2nd edn. Springer, New York, p 161
Garrity GM, Bell JA, Lilburn T (2006) Order III. Rhodobacterales ord. nov. In List of new names and new combinations previously effectively, but not validly, published, Validation List no. 107. Int J Syst Evol Microbiol 56:1–6
Hwang CY, Cho BC (2008) Ponticoccus litoralis gen. nov., sp. nov., a marine bacterium in the family Rhodobacteraceae. Int J Syst Evol Microbiol 58:1332–1338
Hwang CY, Bae GD, Yih W, Cho BC (2009) Marivita cryptomonadis gen. nov., sp. nov. and Marivita litorea sp. nov., of the family Rhodobacteraceae, isolated from marine habitats. Int J Syst Evol Microbiol 59:1568–1575
** HM, Lee HJ, Kim JM, Park MS, Lee K, Jeon CO (2011) Litorimicrobium taeanense gen. nov., sp. nov., isolated from a sandy beach. Int J Syst Evol Microbiol 61:1392–1396
Kim YG, Hwang CY, Cho BC (2008) Pelagicola litoralis gen. nov., sp. nov., isolated from coastal water in Korea. Int J Syst Evol Microbiol 56:2102–2106
Komagata K, Suzuki KI (1987) Lipid and cell wall analysis in bacterial systematics. Methods Microbiol 19:161–207
Lányí B (1987) Classical and rapid identification methods for medically important bacteria. Methods Microbiol 19:1–67
Minnikin DE, Patel PV, Alshamaony L, Goodfellow M (1977) Polar lipid composition in the classification of Nocardia and related bacteria. Int J Syst Bacteriol 27:104–117
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Park S, Yoon JH (2012) Ruegeria arenilitoris sp. nov., isolated from the seashore sand around a seaweed farm. Antonie Van Leeuwenhoek 102:581–589
Park MS, Chung BS, Lee HJ, ** HM, Lee SS, Oh YK, Jeon CO (2011) Citreicella aestuarii sp. nov., isolated from a tidal flat. Int J Syst Evol Microbiol 61:2595–2599
Park S, Lee MH, Lee JS, Oh TK, Yoon JH (2012) Thalassobius maritimus sp. nov., isolated from seawater. Int J Syst Evol Microbiol 62:8–12
Park S, Jung YT, Kim SI, Yoon JH (2013) Thalassococcus lentus sp. nov., an alphaproteobacterium isolated from seawater of a seaweed farm. Antonie Van Leeuwenhoek 103:465–473
Rappé MS, Vergin K, Giovannoni SJ (2000) Phylogenetic comparisons of a coastal bacterioplankton community with its counterparts in open ocean and freshwater systems. FEMS Microbiol Ecol 33:219–232
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI technical note 101. Microbial ID, Inc, Newark
Staley JT (1968) Prosthecomicrobium and Ancalomicrobium: new prosthecate freshwater bacteria. J Bacteriol 95:1921–1942
Tamaoka J, Komagata K (1984) Determination of DNA base composition by reverse-phase high-performance liquid chromatography. FEMS Microbiol Lett 25:125–128
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Wayne LG, Brenner DJ, Colwell RR et al (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Yoon JH, Kim H, Kim SB, Kim HJ, Kim WY, Lee ST, Goodfellow M, Park YH (1996) Identification of Saccharomonospora strains by the use of genomic DNA fragments and rRNA gene probes. Int J Syst Bacteriol 46:502–505
Yoon JH, Lee ST, Park YH (1998) Inter- and intraspecific phylogenetic analysis of the genus Nocardioides and related taxa based on 16S rDNA sequences. Int J Syst Bacteriol 48:187–194
Yoon JH, Kang KH, Park YH (2003) Psychrobacter jeotgali sp. nov., isolated from jeotgal, a traditional Korean fermented seafood. Int J Syst Evol Microbiol 53:449–454
Yoon JH, Kang SJ, Oh TK (2007) Donghicola eburneus gen. nov., sp. nov., isolated from seawater of the East Sea in Korea. Int J Syst Evol Microbiol 57:73–76
Yoon JH, Kang SJ, Lee SY, Oh KH, Oh TK (2009) Seohaeicola saemankumensis gen. nov., sp. nov., isolated from a tidal flat. Int J Syst Evol Microbiol 59:2675–2679
Yoon JH, Kang SJ, Lee SY (2012) Salinimonas lutimais sp. nov., a polysaccharide-degrading bacterium isolated from a tidal flat. Antonie Van Leeuwenhoek 102:803–810
Acknowledgments
This study was supported by a grant from the National Institute of Biological Resources (NIBR), funded by the Ministry of Environment (MOE) and the Program for Collection, Management and Utilization of Biological Resources and BK21 program from the Ministry of Science, ICT & Future Planning (MSIP) of the Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain D1-W8T is KC708867.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Park, S., Jung, YT. & Yoon, JH. Pelagicola litorisediminis sp. nov., a novel alphaproteobacterium isolated from tidal flat sediment. Antonie van Leeuwenhoek 104, 103–110 (2013). https://doi.org/10.1007/s10482-013-9930-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-013-9930-4