Abstract
The Nile Tilapia (Oreochromis niloticus), a gonochoristic teleost fish with a XX/XY sex-determination system, is an ideal model for investigating gonadal sex differentiation. During gonadal differentiation, the expression of cyp19a1a in XX gonads and dmrt1 in XY gonads are required for undifferentiated tissues to develop into ovary or testis. In this study, quantitative real-time RT-PCR assessed the expression of cyp19a1a and dmrt1 genes in gonads and tail fin tissues. Differences in gene expression mean among sexually differentiated fish were analyzed using two-way analysis of variance (ANOVA) and validation of mixed model using discriminant analysis (DA) for morphometric traits and the gene expression in gonads and tail fin tissues used to validate and utilize them in discriminating sexes in sex-differentiated Nile Tilapia fish. The results revealed that, cyp19a1a gene expression in female ovaries was more significant than dmrt1 in male testis. In the other hand, the dmrt1 gene expression in the tail fin was higher in males than females. Both, cyp19a1a and dmrt1 genes, can discriminate fish sexes by 100% by using their expression in tail fin tissues. In conclusion, the cyp19a1a and dmrt1 genes could be used as a genetic marker to discriminate between the Nile Tilapia sexes, whereas used as an indicator for ovarian or testis differentiation in sexually differentiated Nile Tilapia using tail fin tissues. It is worth mentioning that this is the first investigation for using cyp19a1a and dmrt1 genes from Nile Tilapia tail fin tissues in sex determination.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A freshwater fish Nile Tilapia (O. niloticus) is the third most important farmed fish after carp and salmon, and the males grow substantially faster than females. Because of its economic value as one of the world’s most important farmed fish species, its production depends on all-male monosex culture because males grow faster, and to control reproduction until harvest, there are extensive researches into the development of sex control strategies (Palaiokostas et al. 2015). Nile Tilapia is a genetic sex determination (GSD) and temperature-dependent sex determination (TSD) fish with a XX/XY sex-determination mechanism that could be affected by the high temperature (masculinizing above 32 °C) (Wang et al. 2017).
The genes that determine sex in many fish species are quite diverse. Understanding the mechanisms of sex determination in most fish species used in aquaculture, however, is difficult due to the complicated and variable nature of this mechanism (Lin et al. 2017). The Amhy gene has been discovered as a possible sex-determining gene in Nile Tilapia (O. niloticus), and the amhy/amhrII signal has been proposed to have an important role in male sex determination (Li et al. 2015). Gain- and loss-of-function investigations indicate that gsdf is essential for male differentiation in Tilapia (Kaneko et al. 2015; Jiang et al. 2016). In the presence of sf1 (splicing factor 1), the male differentiation gene dmrt1 activates gsdf expression (Jiang et al. 2016). Dmrt1 gene directly controls sox9b (SRY-box transcription factor 9b) by binding to a cis-regulatory region in the sox9b promoter (Wei et al. 2019). Moreover, both foxl2 and cyp19a1a mutant alleles exhibit female-to-male sex reversal (Zhang et al. 2017).
Dmrt1 (Doublesex and Mab-3 (DM)-related transcription factor1) and amh (anti-müllerian) genes are both important in vertebrate’s testis development. Dmrt1 is a member of the DM domain gene family. Its duplication resulted in the medaka sex-determining DMY gene (Matsuda et al. 2002; Nanda et al. 2002). Dmrt1 is structurally conserved across phyla and has an early effect on differentiating testis in non-mammalian vertebrates (Smith and Sinclair 2004). Dmrt1 was one of the first genes elevated in Nile Tilapia XY males, becoming male-specific at later stages (Ijiri et al. 2008). During differentiation, transcriptome analysis in Nile Tilapia indicated that dmrt1 is involved in testes differentiation and development (Tao et al. 2018). It was found in the testis of normal XY and XX males, demonstrating that this is a gene unrelated to Y chromosome whose expression is linked to testicular development (Guan et al. 2000).
Cyp19a (The Cytochrome P450 family 19 subfamily A) is also known as cyp19a1, cyp19a1a, and ovarian aromatase (Guiguen et al. 2010). cyp19a1a has a high level of expression in the ovary and a lower level of expression in the testis and brain (Dalla Valle et al. 2002). cyp19a1a sex-specific expression in XX gonads during early gonadal differentiation is required for ovary differentiation in Nile Tilapia (Ijiri et al. 2008).
Monosex culture includes physical separation of males and females, within-species hybridization to produce all-male fish, and artificial sex reversal using hormones. Treatment with 17-methyltestosterone included in the feed of sexually undifferentiated fry was the most common approach for all-male population production in Tilapia. Combining a sex reversal method with a breeding program to obtain broodstock that produces monosex fry following the reverse technique is one option to eliminate the hormones usage. It is important to develop YY supermales in Nile Tilapia by crossing neo-females (XY) with ordinary males (XY). The use of sex-linked DNA markers simplify the procedure by differentiating XX, XY, or YY individuals, reducing individual genotype identification through progeny testing. There is no information about gene expression of sex determination genes in tail fin tissues to determine sex in Nile Tilapia. The main objective of this work is to investigate the expression of cyp19a1a and dmrt1 genes in tail (caudal) fins of sexually differentiated fish, to assess whether they could be used in future research to differentiate and determine the sex of fish.
Materials and Methods
Sample Collection
The experimental fish used in the present study were 24 adult fish (12 males and 12 females) (aged 3–4 months); individuals were collected from Barsiq fish farm, El-Behaira, Egypt. Briefly, at sampling time, fish were measured (total length (TL), standard length (SL), and body weight (BW)) and separated based on the external morphological difference between males and females. The fish gonads and tail or caudal fins were immediately removed by dissection, frozen in liquid nitrogen, then stored at – 80 °C until analyzed. All animal maintenance and handling procedures followed the recommendations of the Institutional Animal Care Use Committee, Alexandria University, Egypt (Alex-IACUC) review report AU: 14/20/11/01/2/9.
Fulton’s Condition Factor (K)
Fulton’s condition factor (K) is a fish welfare indicator that enables the collection of meaningful data on growth, age, reproduction, nutrition, health, and welfare status. This factor is calculated through following the formula K = (100 × BW)/TL3, where BW represents body weight (g) and TL corresponds to total length (cm) (Gonzalez-Martinez et al. 2021).
RNA Extraction
The Genozol Tri RNA Kit (Geneaid) was used to extract total RNA from the ovary, testis, and apical sections of tail fin clip tissues according to the manufacturer’s procedure. The RNA yield quality and concentrations were tested using a Nanodrop spectrophotometer (BioDrop, England). For each sample, the RNA concentration was standardized to 50 ng.
RT-PCR Reaction for Genes of Interest
Topreal™ One-step RT q-PCR Kit (SYBER Green with low ROX) (enzynomics) is used to RT-PCR reactions performed according to the manufacturer procedure for expression analysis as well as β-actine gene was used as a reference gene; the reaction conditions were as follows: holding at 45 °C for 30 min (cDNA synthesis step), PCR reaction initiated at 95 °C for 10 min, followed by 95 °C for 5 s, then annealing step for 30 s for various primers cyp19a1a, dmrt1a, and β- actine genes that reaction repeated for 50 cycles (Table 1). The specificity of real-time PCR amplification was validated by analysis of melting curves, which ensured that only one PCR product was specifically amplified at the target size. The cycle threshold (Ct) was calculated for each replicate and final values were obtained from the average of two replicates per sample. The expression values of the studied genes were normalized using the expression values of a reference gene β-actine.
Studied gene expression analysis was done using 2−∆∆Ct method (Livak and Schmittgen 2001) according to the following equation:
Fold difference = 2−ΔΔCt
The ΔΔCt method is a popular methodology for comparing experimental sample data with a calibrator (e.g., untreated or wild-type sample) and a normalizer (e.g., housekee** gene expression). With this method, Ct s for the gene of interest (GOI) in both the test sample(s) and calibrator sample are now normalized in relation to a normalizer (norm) gene Ct from the same two samples. The obtaining ΔΔCt value is incorporated to calculate the fold difference in expression.
The Ct method is an effective approach for comparing the results of experimental samples that include both a calibrator (such as an untreated or wild-type sample) and a normalizer (e.g., housekee** gene expression). Using this approach, the Ct values for the gene of interest (GOI) in the test sample(s) and calibrator sample are now modified to the gene’s normalizer (norm) Ct values from the same two samples. The obtained Ct value is used to compute the fold difference in expression.
Statistical Analysis
The following model was used to investigate differences in mean gene expression across sexually differentiated fish using two-way analysis of variance (ANOVA) (CoStat 2017):
where Yijl is the observation of the ijlth parameter measured; µ is the overall mean; Gi is the effect of the ith gene; Sj is the effect of jth sex; (GS)ij is the interaction genes by sex; eijl is the random error. Significant differences (P ≤ 0.05) among means were tested by the method of Duncan (1955). The T-test was used to analyze the morphometric characteristics.
Furthermore, the final data set was subjected to discriminant analysis (DA) which is a statistical procedure classify a collection of observations into two groups based on the values of independent, continuous categorical variables or predictors (the values of a categorical, dependent discriminant variable, also known as the grou** variable). In this study, sex (male or female) served as the categorical variable, while morphometric features and the levels of various genes’ expression acted as the predictors. The analysis produces a linear discriminant function that assigns values of the grou** variable based on the predictors’ values (Legendre and Legendre 2012). Based on the weighted combination of the independent variables, a discriminant score can be calculated according to the equation:
where Di is the predicted (discriminant) score, a is constant, x is the predictor, and b is the discriminant coefficient.
Results
Morphometric Characteristics
ANOVA test results reported that body weight (BW), total length (TL), and standard length (SL) differed significantly high (P ≤ 0.05) between females and males, with the mean values of these variables between 61.11 ± 14.78 and 184.736 ± 27.09, 14.2 ± 0.92 cm and 21.18 ± 1.28 cm, and 11.47 ± 0.86 cm and 17.1 ± 1.15 cm, respectively. There are no significant differences (P ≤ 0.05) between females and males in Fulton’s condition factor (K) (Table 2).
The Expression Profile of cyp19a1a and dmrt1 Genes
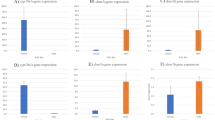
The expression of all studied genes was detected in both gonads and tail fin tissues of females and males. Comparative gene expression analysis of sex determination genes was conducted, and the results showed that in gonads, the female ovaries had higher levels of cyp19a1a gene expression than in the male testis. Also, Dmrt1 gene expression in the male’s testis was higher than in the female’s ovaries (Fig. 1A, B). In tail fin tissues, cyp19a1a gene expression was higher in the female’s tail fins than in the males. In contrast, Dmrt1 gene expression which was higher in males than females (Fig. 1C, D).
Sex-Related Differences of cyp19a1a and dmrt1 Genes in Gonads
First, we analyzed the expression of cyp19a1a and dmrt1 in gonads of 3–4 months sexually differentiated female and male Nile Tilapia fish. Statistical analysis showed that there was a highly significant difference (P ≤ 0.05) between mean gene expressions. Concerning the effect of sex on gene expression, there was a high significance between males and females. Regarding, the interaction between G × S, the results reported that cyp19a1a in females’ ovaries recorded a higher gene expression value than the other interactions, as shown in Table 3.
Sex-Related Differences of cyp19a1a and dmrt1 Genes in Tail (Caudal) Fins
The sex-related differences in gene expression of cyp19a1a and dmrt1 in tail fin tissues were analyzed in the same fish samples, where there were significant differences (P ≤ 0.05) between gene expressions between sexes, and dmrt1 was more significant than cyp19a1a. And, in terms of sex, males were higher than females. Also, the interaction between gene and sex reported that dmrt1 in male tail fins had the highest gene expression value among other interactions, as shown in Table 3.
Definition of Sex Predictor Model by Discriminant Analysis (DA)
The discriminant analysis (DA) determines which variables, including morphometric characteristics (TL, SL, K factor, and BW) and gene expression of two studied genes in gonads, can differentiate between sexes. We checked for which gene of two genes in tail fins best defined the groups (males and females) in Nile Tilapia.
For morphometric characteristics, the BW, TL, and SL factors significantly (P ≤ 0.05) contributed to differentiate sex: F = 16.043, 19.635, and 15.397; Wilks’ λ test values were 0.384, 0.337, and 0.394, and can correctly classify 83.3% of fish, respectively. However, the K factor had no significant and high value of Wilks’ λ = 0.919. as shown in Table 4.
Also, in Table 4, all expression levels of the two genes as determined by real-time RT-PCR on samplings from gonads were used to perform the discriminant analysis (DA). The results revealed that cyp19a1a and dmrt1 expression levels were significantly high (P ≤ 0.05) contributing to differentiate sex, where F = 46.690 and 79.547; Wilks’ λ test values were 0.176 and 0.112 and able to correctly classify 100% of fish, respectively. This concluded that cyp19a1a and dmrt1 had a significant effect on discriminating sex in sexually differentiated Nile Tilapia.
The result of gene expression levels of the two genes from tail fin tissues revealed that cyp19a1a and dmrt1 expression levels are significantly high (P ≤ 0.05) contributing to separate sex, where Wilks’ λ test values were 0.185 and 0.135 and were capable to correctly classify 100% of the female fish, respectively (Table 4). Wilks’ λ values range from 0 to 1.0, the smaller the λ value, the more the variable discriminates well among groups. This statistically concluded that cyp19a1a and dmrt1 were best for discriminating sex in sexually differentiated Nile Tilapia using tail fin tissues.
Discussion
Owing to sex growth differences, it is generally advantageous in aquaculture to produce fish monosex populations, for example, male Nile Tilapia grow faster than females (Chakraborty and Banerjee 2010). Nile Tilapia is a genetic sex determination (GSD) plus temperature-dependent sex determination (TSD) fish, with a XX/XY sex-determination system (Wang et al. 2017). Even so, understanding the mechanisms behind sex determination in most fish species used in aquaculture is difficult due to the complicated and varied nature of these processes; we investigated the differences in morphometric parameters and gene expression of cyp19a1a and dmrt1 in gonads and tail fin tissues between Nile Tilapia females and males.
While the gene expression of sex determination genes becomes less accurate in tissues not directly implicated in reproduction, we detected cyp19a1a and dmrt1 expression in female and male tail fins besides gonads; these findings along with Hofsten and Olsson (2005) who reported that Sox9a (a gene that plays an important role in sex determination) are present in zebrafish (Danio rerio) pectoral fin. Our results reported a high expression of the cyp19a1a gene in female gonads which is consistent with previous research on the Southern flounder (Luckenbach et al. 2005), Atlantic halibut (Matsuoka et al. 2006), and rainbow trout (Vizziano et al. 2007) which suggested that the expression of the cyp19a1a gene acts as an early marker of sex differentiation in these species. Furthermore, this research, combined with studies on Nile Tilapia conducted by (Ijiri et al. 2008) demonstrated that cyp19a1a in XX gonads during early gonadal differentiation is crucial for ovary differentiation. In males’ tail fins unlike gonads, we found a higher expression of the dmrt1 gene than cyp19a1a. These findings were consistent with the research of (Guan et al. 2000) who discovered that the dmrt1 gene is not linked to the Y chromosome, whose expression is correlated with testicular development because it was expressed in the testis of normal XY males and XX males. It is likely to Panagiotopoulou et al. (2023) who found genetic sex markers in Siberian (Acipenser baerii) and Atlantic (A. oxyrinchus) sturgeons; however, they utilized a different technique, extracting genetic material from female and male tail fins.
Discriminant analysis is used in a variety of research fields, including numerical ecology applied to fisheries management to classify the manipulation of a particular ecosystem (Tudela et al. 2005); in microbiology, to differentiate between human and animal sources of contaminates in surface waters (Kaneene et al. 2007); in forensics, to estimate the sex of unknown skeletal remains (Kemkes-Grottenthaler 2005); also in medical research, for instance to discriminate between different types of anemia (Ahluwalia et al. 1995); and for the first time, it used in study sex differentiation of seabass (Blázquez et al. 2009).
In this study, discriminant analysis was used first to define which variables among morphometric parameters and genes (cyp19a1a and dmrt1) could discriminate sex in adult Nile Tilapia fish gonads and then in tail fins. Wilks’ λ values range from 0 to 1.0; the smaller the λ value, the more the variable discriminates well among groups. For morphometric characteristics, the results reported that BW, TL, and SL contributed to differentiate sex: F = 16.043, 19.635, and 15.397; Wilks’ λ test values were 0.384, 0.337, and 0.394, respectively, and could discriminate between females and males by 83.3%. This is agreed with the fact that males grow faster than females (Chakraborty and Banerjee 2010) and with Blázquez et al. (2009) who indicate that SL can classify 75.7% of European seabass fish. For gene expression, when using the expression levels of the two genes from gonads and tail fin tissues, the result revealed that cyp19a1a and dmrt1 expression levels are significantly high (P ≤ 0.05), and these two genes contribute to a 100% accurate discrimination and classification of sexes. The research on seabass fish conducted by (Blázquez et al. 2009) revealed that the cyp19a1a gene was able to correctly classify 100% of the fish.
In conclusion, cyp19a1a and dmrt1 genes could be used as a genetic marker to discriminate between the fish sexes and as an indicator for ovarian or testis differentiation in sexually differentiated Nile Tilapia. Additionally, the tissues from tail fins had significant levels of cyp19a1a and dmrt1 gene expression. Therefore, they could be utilized in future studies to distinguish between various fish sexes and to identify their sex using molecular markers with the advantage that it is an un-scarified method. The use of sex-linked DNA markers could shorten the process by distinguishing XX, XY, or YY individuals in breeding programs of fish stocks, thus avoiding the identification of individual genotypes by progeny testing. Moreover, this work is the first investigation for using cyp19a1a and dmrt1 genes from Nile Tilapia tail fin tissues in sex determination.
Data Availability
No datasets were generated or analyzed during the current study.
References
Ahluwalia N, Lammi-Keefe CJ, Bendel RB et al (1995) Iron deficiency and anemia of chronic disease in elderly women: a discriminant-analysis approach for differentiation. Am J Clin Nutr 61:590–596
Blázquez M, Navarro-Martín L, Piferrer F (2009) Expression profiles of sex differentiation-related genes during ontogenesis in the european sea bass acclimated to two different temperatures. J Exp Zool Part B Mol Dev Evol 312:686–700
Chakraborty SB, Banerjee S (2010) Comparative growth performance of mixed-sex and monosex Nile Tilapia population in freshwater cage culture system under Indian perspective. Int J Biol. https://doi.org/10.5539/ijb.v2n1p44
CoStat 6.451 (2017) Copyright(c) 1998–2017, CoHort Software. http://www.cohort.com
Dalla Valle L, Lunardi L, Colombo L, Belvedere P (2002) European sea bass (Dicentrarchus labrax L.) cytochrome P450arom: cDNA cloning, expression and genomic organization. J Steroid Biochem Mol Biol 80:25–34
Duncan DB (1955) Multiple range and multiple F tests. Biometrics 11:1
Gonzalez-Martinez A, De-Pablos-Heredero C, González M et al (2021) Usefulness of discriminant analysis in the morphometric differentiation of six native freshwater species from Ecuador. Animals. https://doi.org/10.3390/ani11010111
Guan G, Kobayashi T, Nagahama Y (2000) Sexually dimorphic expression of two types of DM (Doublesex/Mab-3)-domain genes in a teleost fish, the Tilapia (Oreochromis niloticus). Biochem Biophys Res Commun 272:662–666
Guiguen Y, Fostier A, Piferrer F, Chang CF (2010) Ovarian aromatase and estrogens: a pivotal role for gonadal sex differentiation and sex change in fish. Gen Comp Endocrinol 165:352–366
Hofsten J, Olsson PE (2005) Zebrafish sex determination and differentiation: involvement of FTZ-F1 genes. Reprod Biol Endocrinol 3:1–11
Ijiri S, Kaneko H, Kobayashi T et al (2008) Sexual dimorphic expression of genes in gonads during early differentiation of a teleost fish, the Nile Tilapia Oreochromis niloticus. Biol Reprod 78:333–341
Jiang DN, Yang HH, Li MH et al (2016) gsdf is a downstream gene of dmrt1 that functions in the male sex determination pathway of the Nile Tilapia. Mol Reprod Dev 83:497–508
Kaneene JB, Miller RA, Sayan R et al (2007) Considerations when using discriminant function analysis of antimicrobial resistance profiles to identify sources of fecal contamination of surface water in Michigan. Appl Environ Microbiol 73:2878–2890
Kaneko H, Ijiri S, Kobayashi T et al (2015) Gonadal soma-derived factor (gsdf), a TGF-beta superfamily gene, induces testis differentiation in the teleost fish Oreochromis niloticus. Mol Cell Endocrinol 415:87–99
Kemkes-Grottenthaler A (2005) Sex determination by discriminant analysis: an evaluation of the reliability of patella measurements. Forensic Sci Int 147:129–133
Legendre P, Legendre L (2012) Numerical ecology - P. Elsevier, Legendre, Louis Legendre, Third Engl
Li M, Sun Y, Zhao J et al (2015) A tandem duplicate of anti-Müllerian hormone with a missense SNP on the Y chromosome is essential for male sex determination in Nile Tilapia. Oreochromis Niloticus Plos Genet 11:e1005678
Lin A, **ao S, Xu S et al (2017) Identification of a male-specific DNA marker in the large yellow croaker (Larimichthys crocea). Aquaculture 480:116–122
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Luckenbach JA, Early LW, Rowe AH et al (2005) Aromatase cytochrome P450: cloning, intron variation, and ontogeny of gene expression in southern flounder (Paralichthys lethostigma). J Exp Zool Part A Comp Exp Biol 303A:643–656
Matsuda M, Nagahama Y, Shinomiya A et al (2002) DMY is a Y-specific DM-domain gene required for male development in the medaka fish. Nature 417:559–563
Matsuoka MP, van Nes S, Andersen Ø et al (2006) Real-time PCR analysis of ovary- and brain-type aromatase gene expression during Atlantic halibut (Hippoglossus hippoglossus) development. Comp Biochem Physiol Part B Biochem Mol Biol 144:128–135
Nanda I, Kondo M, Hornung U et al (2002) A duplicated copy of DMRT1 in the sex-determining region of the Y chromosome of the medaka, Oryzias latipes. Proc Natl Acad Sci U S A 99:11778–11783
Palaiokostas C, Bekaert M, Khan MGQ et al (2015) A novel sex-determining QTL in Nile Tilapia (Oreochromis niloticus). BMC Genomics. https://doi.org/10.1186/s12864-015-1383-x
Panagiotopoulou H, Marzecki K, Gawor J et al (2023) Extensive search of genetic sex markers in Siberian (Acipenser baerii) and Atlantic (A. oxyrinchus) sturgeons. Aquaculture 573:739517
Poonlaphdecha S, Pepey E, Canonne M et al (2013) Temperature induced-masculinisation in the Nile Tilapia causes rapid up-regulation of both dmrt1 and amh expressions. Gen Comp Endocrinol 193:234–242
Rengmark AH, Slettan A, Lee WJ et al (2007) Identification and map** of genes associated with salt tolerance in Tilapia. J Fish Biol 71:409–422
Smith CA, Sinclair AH (2004) Sex determination: insights from the chicken. BioEssays 26:120–132
Tao W, Chen J, Tan D et al (2018) Transcriptome display during Tilapia sex determination and differentiation as revealed by RNA-Seq analysis. BMC Genomics 19:1–12
Tudela S, Coll M, Palomera I (2005) Develo** an operational reference framework for fisheries management on the basis of a two-dimensional index of ecosystem impact. ICES Journal of Marine Science. Oxford Academic, pp 585–591
Vizziano D, Randuineau G, Baron D et al (2007) Characterization of early molecular sex differentiation in rainbow trout, Oncorhynchus mykiss. Dev Dyn 236:2198–2206
Wang YY, Sun LX, Zhu JJ et al (2017) Epigenetic control of cyp19a1a expression is critical for high temperature induced Nile Tilapia masculinization. J Therm Biol 69:76–84
Wei L, Li X, Li M et al (2019) Dmrt1 directly regulates the transcription of the testis-biased Sox9b gene in Nile Tilapia (Oreochromis niloticus). Gene 687:109–115
Zhang X, Li M, Ma H et al (2017) Mutation of foxl2 or cyp19a1a results in female to male sex reversal in XX Nile Tilapia. Endocrinology 158:2634–2647
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). Open access funding was provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study. Samy Y. El-Zaeem put the experimental idea and design and revised the manuscript. Amr El-Hanafy and Alaa A. El-Dahhar revised the manuscript. Ayaat M. Elmaghraby helped in the practical part, writing, and revising the manuscript. Amany M. Hendy performed most of the experiments, analyzed the data, and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
El-Zaeem, S.Y., El-Hanafy, A., El-Dahhar, A.A. et al. A New Investigation to Discriminate Sexes in Alive Nile Tilapia (Oreochromis niloticus) Using Cyp19a1a and Dmrt1 Gene Expression in Tail Fin Tissues. Mar Biotechnol (2024). https://doi.org/10.1007/s10126-024-10340-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10126-024-10340-w