Abstract
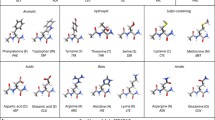
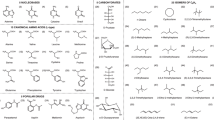
Force fields are actively used to study RNA. Development of accurate force fields relies on a knowledge of how the variation of properties of molecules depends on their structure. Detailed scrutiny of RNA’s conformational preferences is needed to guide such development. Towards this end, minimum energy structures for each of a set of 16 small RNA-derived molecules were obtained by geometry optimization at the HF/6-31G(d,p), B3LYP/apc-1, and MP2/cc-pVDZ levels of theory. The number of minima computed for a given fragment was found to be related to both its size and flexibility. Atomic electrostatic multipole moments of atoms occurring in the [HO-P(O3)-CH2-] fragment of 30 sugar-phosphate-sugar geometries were calculated at the HF/6-31G(d,p) and B3LYP/apc-1 levels of theory, and the transferability of these properties between different conformations was investigated. The atomic multipole moments were found to be highly transferable between different conformations with small standard deviations. These results indicate necessary elements of the development of accurate RNA force fields.
Graphical abstract














Similar content being viewed by others
References
Noller HF (2005) RNA structure: reading the ribosome. Science 309:1508–1514
Krutzfeldt J, Stoffel M (2006) MicroRNAs: a new class of regulatory genes affecting metabolism. Cell Metab. 4:9–12
Doudna JA, Cech TR (2002) The chemical repertoire of natural ribozymes. Nature 418:222–228
Nissen P, Hansen J, Ban N, Moore PB, Steitz TA (2000) The structural basis of ribosome activity in peptide bond synthesis. Science 289:920–930
Scott WG (2007) Ribozymes. Curr Opin Struct Biol 17:280–286
Aduri R, Psciuk BT, Saro P, Taniga H, Schlegel HB, SantaLucia J (2007) AMBER force field parameters for the naturally occurring modified nucleosides in RNA. J Chem Theory Comput 3:1464–1475
Case DA, Cheatham 3rd TE, Darden T, Gohlke H, Luo R, Merz Jr KM, Onufriev A, Simmerling C, Wang B, Woods RJ (2005) The Amber biomolecular simulation programs. J Comput Chem 26:1668–1688
Brooks BR, Brooks 3rd CL, Mackerell Jr AD, Nilsson L, Petrella RJ, Roux B, Won Y, Archontis G, Bartels C, Boresch S, Caflisch A, Caves L, Cui Q, Dinner AR, Feig M, Fischer S, Gao J, Hodoscek M, Im W, Kuczera K, Lazaridis T, Ma J, Ovchinnikov V, Paci E, Pastor RW, Post CB, Pu JZ, Schaefer M, Tidor B, Venable RM, Woodcock HL, Wu X, Yang W, York DM, Karplus M (2009) CHARMM: the biomolecular simulation program. J Comput Chem 30:1545–1614
van Gunsteren, W. F., Billeter, S. R., Eising, A. A., Hünenberger, P. H., Krüger, P. K. H. C., Mark, A. E., Scott, W. R., Tironi, I. G.(1996) Biomolecular simulation: the {GROMOS96} manual and user guide
Zichi DA (1995) Molecular dynamics of RNA with the OPLS force field. Aqueous simulation of a hairpin containing a tetranucleotide loop. J Am Chem Soc 117:2957–2969
Tubbs JD, Condon DE, Kennedy SD, Hauser M, Bevilacqua PC, Turner DH (2013) The nuclear magnetic resonance of CCCC RNA reveals a right-handed Helix, and revised parameters for AMBER force field torsions improve structural predictions from molecular dynamics. Biochemistry 52:996–1010
Schmidt JR, Yu K, McDaniel JG (2015) Transferable next-generation force fields from simple liquids to complex materials. Acc Chem Res 48:548–556
Richardson JS, Schneider B, Murray LW, Kapral GJ, Immormino RM, Headd JJ, Richardson DC, Ham D, Hershkovits E, Williams LD, Keating KS, Pyle AM, Micallef D, Westbrook J, Berman HM, Consortium RNAO (2008) RNA backbone: consensus all-angle conformers and modular string nomenclature (an RNA ontology consortium contribution). RNA 14:465–481
Perez A, Marchan I, Svozil D, Sponer J, Cheatham TE, Laughton CA, Orozco M (2007) Refinement of the AMBER force-field for nucleic acid simulations. Improving the description of α/β conformers. Biophys J 92:3817–3829
Yildirim I, Stern HA, Kennedy SD, Tubbs JD, Turner DH (2010) Reparameterization of RNA chi torsion parameters for the AMBER force field and comparison to NMR spectra for cytidine and uridine. J Chem Theory Comput 6:1520–1531
Banas P, Hollas D, Zgarbova M, Jurecka P, Orozco M, Cheatham TE, Sponer J, Otyepka M (2010) Performance of molecular mechanics force fields for RNA simulations: stability of UUCG and GNRA hairpins. J Chem Theory Comput 6:3836–3849
Denning EJ, Priyakumar UD, Nilsson L, Mackerell Jr AD (2011) Impact of 2′-hydroxyl sampling on the conformational properties of RNA: update of the CHARMM all-atom additive force field for RNA. J Comput Chem 32:1929–1943
Zgarbova M, Otyepka M, Sponer J, Mladek A, Banas P, Cheatham TE, Jurecka P (2011) Refinement of the Cornell et al. Nucleic acids force field based on reference quantum chemical calculations of glycosidic torsion profiles. J Chem Theory Comput 7:2886–2902
Wales DJ, Yildirim I (2017) Improving computational predictions of single-stranded RNA tetramers with revised α/γ torsional parameters for the Amber force field. J Phys Chem B 121:2989–2999
Robertson MJ, Tirado-Rives J, Jorgensen WL (2017) Improved treatment of nucleosides and nucleotides in the OPLS-AA force field. Chem Phys Lett 683:276–280
Yildirim I, Stern HA, Tubbs JD, Kennedy SD, Turner DH (2011) Benchmarking AMBER force fields for RNA: comparisons to NMR spectra for single-stranded r (GACC) are improved by revised χ torsions. J Phys Chem B 115:9261–9270
Yildirim I, Kennedy SD, Stern HA, Hart JM, Kierzek R, Turner DH (2012) Revision of AMBER torsional parameters for RNA improves free energy predictions for tetramer duplexes with GC and iGiC base pairs. J Chem Theory Comput 8:172–181
Kuhrova P, Mlynsky V, Zgarbova M, Krepl M, Bussi G, Best RB, Otyepka M, Sponer J, Banas P (2019) Improving the performance of the Amber RNA force field by tuning the hydrogen-bonding interactions. J Chem Theory Comput 15:3288–3305
Kuhrova P, Mlynsky V, Zgarbova M, Krepl M, Bussi G, Best RB, Otyepka M, Sponer J, Banas P (2020) Correction to “improving the performance of the Amber RNA force field by tuning the hydrogen-bonding interactions”. J Chem Theory Comput 16:818–819 erratum
Sponer J, Bussi G, Krepl M, Banas P, Bottaro S, Cunha RA, Gil-Ley A, Pinamonti G, Poblete S, Jurecka P, Walter NG, Otyepka M (2018) RNA structural dynamics as captured by molecular simulations: a comprehensive overview. Chem Rev 118:4177–4338
Svozil D, Sponer JE, Marchan I, Perez A, Cheatham 3rd TE, Forti F, Luque FJ, Orozco M, Sponer J (2008) Geometrical and electronic structure variability of the sugar-phosphate backbone in nucleic acids. J Phys Chem B 112:8188–8197
Sponer J, Mladek A, Sponer JE, Svozil D, Zgarbova M, Banas P, Jurecka P, Otyepka M (2012) The DNA and RNA sugar-phosphate backbone emerges as the key player. An overview of quantum-chemical, structural biology and simulation studies. Phys Chem Chem Phys 14:15257–15277
Schneider B, Neidle S, Berman HM (1997) Conformations of the sugar-phosphate backbone in helical DNA crystal structures. Biopolymers 42:113–124
Antson AA, Dodson EJ, Dodson G, Greaves RB, Chen X, Gollnick P (1999) Structure of the Trp RNA-binding attenuation protein, TRAP, bound to RNA. Nature 401:235–242
Mladek A, Banas P, Jureccka P, Otyepka M, Zgarbova M, Sponer J (2014) Energies and 2′-hydroxyl group orientations of RNA backbone conformations. Benchmark CCSD (T)/CBS database, electronic analysis, and assessment of DFT methods and MD simulations. J Chem Theory Comput 10:463–480
Gorb L, Podolyan Y, Dziekonski P, Sokalski WA, Leszczynski J (2004) Double-proton transfer in adenine-thymine and guanine-cytosine base pairs. A post-hartree-fock ab initio study. J Am Chem Soc 126:10119–10129
Kelly REA, Kantorovich LN (2006) Planar nucleic acid base super-structures. J Mater Chem 16:1894–1905
Sharma P, Lait LA, Wetmore SD (2013) yDNA versus yyDNA pyrimidines: computational analysis of the effects of unidirectional ring expansion on the preferred sugar-base orientation, hydrogen-bonding interactions and stacking abilities. Phys Chem Chem Phys 15:2435–2448
MacKerell Jr AD (2009) Contribution of the intrinsic mechanical energy of the phosphodiester linkage to the relative stability of the a, BI, and BII forms of duplex DNA. J Phys Chem B 113:3235–3244
Ode H, Matsuo Y, Neya S, Hoshino T (2008) Force field parameters for rotation around chi torsion axis in nucleic acids. J Comput Chem. 29:2531–2542
Zgarbova M, Luque FJ, Sponer J, Cheatham TE, Otyepka M, Jurecka P (2013) Toward improved description of DNA backbone: revisiting epsilon and zeta torsion force field parameters. J Chem Theory Comput 9:2339–2354
Savelyev A, MacKerell Jr AD (2014) All-atom polarizable force field for DNA based on the classical Drude oscillator model. J Comput Chem 35:1219–1239
Ponder JW, Wu C, Ren P, Pande VS, Chodera JD, Schnieders MJ, Haque I, Mobley DL, Lambrecht DS, DiStasio RA, Head-Gordon M, Clark GNI, Johnson ME, Head-Gordon T (2010) Current status of the AMOEBA polarizable force field. J Phys Chem B 114:2549–2564
Li X, Ponomarev SY, Sa Q, Sigalovsky DL, Kaminski GA (2013) Polarizable simulations with second order interaction model (POSSIM) force field: develo** parameters for protein sidechain analogues. J Comput Chem 34:1241–1250
Gresh N, Naseem-Khan S, Lagardere L, Piquemal J-P, Sponer JE, Sponer J (2017) Channeling through two stacked guanine quartets of one and two alkali cations in the Li+, Na+, K+, and Rb+ series. Assessment of the accuracy of the SIBFA anisotropic polarizable molecular mechanics potential. J Phys Chem B 121:3997–4014
Ren P, Ponder JW (2003) Polarizable atomic multipole water model for molecular mechanics simulation. J. Phys. Chem. B 107:5933–5947
Ponder JW, Wu C, Ren P, Pande VS, Chodera JD, Schnieders MJ, Haque I, Mobley DL, Lambrecht DS, DiStasio Jr RA, Head-Gordon M, Clark GNI, Johnson ME, Head-Gordon T (2010) Current status of the AMOEBA polarizable force field. J. Phys. Chem. B 114:2549–2564
Ren P, Wu C, Ponder JW (2011) Polarizable atomic multipole-based molecular mechanics for organic molecules. J. Chem. Theory Comput. 7:3143–3161
Shi Y, **a Z, Zhang J, Best R, Wu C, Ponder JW, Ren P (2013) Polarizable atomic multipole-based AMOEBA force field for proteins. J. Chem. Theory Comput. 9:4046–4063
Zhang C, Lu C, **g Z, Wu C, Piquemal JP, Ponder JW, Ren P (2018) AMOEBA polarizable atomic multipole force field for nucleic acids. J. Chem. Theory Comput. 14:2084–2108
Savelyev A, MacKerell Jr AD (2014) All-atom polarizable force field for DNA based on the classical Drude oscillator model. J. Comput. Chem. 35:1219–1239
Lemkul, J. A., MacKerell, A. D. Jr.(2017) Polarizable force field for DNA based on the classical Drude oscillator: I. Refinement using quantum mechanical base stacking and conformational energetics. J. Chem. Theory Comput., 13:2053–2071
Lemkul JA, MacKerell Jr AD (2017) Polarizable force field for DNA based on the classical Drude oscillator: II. Microsecond molecular dynamics simulations of duplex DNA. J. Chem. Theory Comput 13:2072–2085
Lemkul JA, MacKerell Jr AD (2018) Polarizable force field for RNA based on the classical Drude oscillator. J. Comput. Chem. 39:2624–2646
Poirier R, Kari R, Csizmadia IG (1985) Handbook of Gaussian basis sets : a compendium for ab-initio molecular orbital calculations. Elsevier, Amsterdam
Roothaan CCJ (1951) New developments in molecular orbital theory. Rev. Mod. Phys. 23:69–89
Ditchfield R, Hehre WJ, Pople JA (1971) Self-consistent molecular-orbital methods. IX. An extended Gaussian-type basis for molecular-orbital studies of organic molecules. J. Chem. Phys 54:724–728
Hehre WJ, Ditchfield R, Pople JA (1972) Self—consistent molecular orbital methods. XII. Further extensions of Gaussian—type basis sets for use in molecular orbital studies of organic molecules. J. Chem. Phys 56:2257–2261
Hariharan PC, Pople JA (1973) The influence of polarization functions on molecular orbital hydrogenation energies. Theor. Chim. Acta 28:213–222
Francl MM, Pietro WJ, Hehre WJ, Binkley JS, Gordon MS, DeFrees DJ, Pople JA (1982) Self-consistent molecular orbital methods. XXIII. A polarization-type basis set for second-row elements. J. Chem. Phys 77:3654–3665
Gordon MS, Binkley JS, Pople JA, Pietro WJ, Hehre WJ (1982) Self-consistent molecular-orbital methods. 22. Small Split-valence basis sets for second-row elements. J. Am. Chem. Soc. 104:2797–2803
Jensen F (2002) Polarization consistent basis sets. III. The importance of diffuse functions. J. Chem. Phys 117:9234–9240
Vosko SH, Wilk L, Nusair M (1980) Accurate spin-dependent electron liquid correlation energies for local spin density calculations: a critical analysis. Can. J. Phys. 58:1200–1211
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 37:785–789
Becke, A. D.(1993) Density-functional thermochemistry. III. The role of exact exchange. J. Chem. Phys., 98:5648–5652
Jensen F (2001) Polarization consistent basis sets: principles. J. Chem. Phys. 115:9113–9125
Jensen, F.(2002) Polarization consistent basis sets. II. Estimating the Kohn–Sham basis set limit. J. Chem. Phys., 116:7372–7379
Jensen, F., Helgaker, T.(2004) Polarization consistent basis sets. V. The elements Si-Cl. J. Chem. Phys., 121:3463–3470
Dunning TH (1989) Gaussian basis sets for use in correlated molecular calculations. I. The atoms boron through neon and hydrogen. J. Chem. Phys 90:1007–1023
Møller C, Plesset MS (1934) Note on an approximation treatment for many-electron systems. Phys. Rev. 46:618–622
Head-Gordon M, Pople J, Frisch M (1988) MP2 energy evaluation by direct methods. Chem. Phys. Lett. 153:503–506
Stephens PJ, Devlin FJ, Chabalowski CF, Frisch MJ (1994) Ab initio calculation of vibrational circular dichroism spectra using large basis set MP2 force fields. Chem. Phys. Lett. 225:247–257
Woon DE, Dunning TH (1993) Gaussian basis sets for use in correlated molecular calculations. III. The atoms aluminum through argon. J. Chem. Phys 98:1358–1371
Yuan Y, Mills MJ, Popelier PLA, Jensen F (2014) Comprehensive analysis of energy minima of the 20 natural amino acids. J. Phys. Chem. A 118:7876–7891
Yuan Y, Zhang Z, Mills MJL, Hu R, Zhang R (2018) Assessing force field potential energy function accuracy via a multipolar description of atomic electrostatic interactions in RNA. J. Chem. Inf. Model. 58:2239–2254
Roost C, Lynch SR, Batista PJ, Qu K, Chang HY, Kool ET (2015) Structure and thermodynamics of N6-methyladenosine in RNA: a spring-loaded base modification. J. Am. Chem. Soc. 137:2107–2115
Popelier PLA, Logothetis G (1998) Characterization of an agostic bond on the basis of the electron density. J. Organomet. Chem. 555:101–111
Frisch MJ, Truchs GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Schmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery JJA, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam NJ, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2015) Gaussian 09, Revision E.01. Gaussian, Inc., Wallingford
Jakli I, Perczel A, Farkas O, Hollosi M, Csizmadia IG (1998) Peptide models XXII. A conformational model for aromatic amino acid residues in proteins. A comprehensive analysis of all the RHF/6–31+G* conformers of for-L-Phe–NH2. J. Mol. Struct.: THEOCHEM 455:303–314
Baldoni HA, Rodriguez AM, Zamora MA, Zamarbide GN, Enriz RD, Farkas Ö, Csàszàr P, Torday LL, Sosa CP, Jàkli I (1999) Peptide models XXIV: an Ab initio study on N-formyl-L-prolinamide with trans peptide bond. The existence or non-existence of αL and ϵL conformations. J. Mol. Struct.: THEOCHEM 465:79–91
Bader RFW (1990) Atoms in molecules. A quantum theory. Oxford, Oxford University Press
Popelier PLA (2000) Atoms in molecules. An introduction. London, Pearson Education
Keith, T. A., AIMAll, 1997-2019. Available at: http://aim.tkgristmill.com. Accessed 1 Mar 2020
Availability of data and material
N/A
Code availability
N/A
Funding
This work was supported by the National Natural Science Foundation of China (No. 21503101), the Fundamental Research Funds for the Central Universities (Grant No. lzujbky-2019-cd05), the National Natural Science Foundation of China (No.81973786), and the Science Foundation of Guangxi (AA17204096, AD16380076).
Author information
Authors and Affiliations
Contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supporting Information Available: Supporting Information contains Cartesian coordinates of all the 22 B3LYP/apc-1 minima of SPS.
ESM 1
(DOCX 118 kb)
Rights and permissions
About this article
Cite this article
Yuan, Y., Mills, M.J.L., Zhang, Z. et al. A general RNA force field: comprehensive analysis of energy minima of molecular fragments of RNA. J Mol Model 27, 137 (2021). https://doi.org/10.1007/s00894-021-04746-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-021-04746-9