Abstract
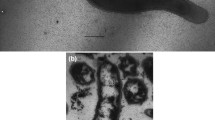
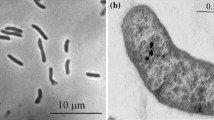
A novel alkaliphilic sulfate-reducing bacterium, strain M08butT, was isolated from a salsa lake of terrestrial mud volcano (Taman Peninsula, Russia). Cells were rod-shaped, motile and Gram-stain-negative. The temperature range for growth was 15–42 °C (optimum at 30 °C). The pH range for growth was 7.0–11.0, with an optimum at pH 8.5–9.0 Strain M08butT used sulfate, thiosulfate, sulfite, dimethyl sulfoxide and arsenate as electron acceptors. Acetate, formate, butyrate, fumarate, succinate, glycerol and pyruvate were utilized as electron donors with sulfate. Fermentative growth was observed with fumarate, pyruvate, crotonate. Strain M08butT grew chemolithoautotrophically with H2 and CO2. The G + C content of the genomic DNA was 60.1%. The fatty acid profile of strain M08butT was characterized by the presence of anteiso-C15:0 as the major component (68.8%). The closest phylogenetic relative of strain M08butT was Desulfatitalea tepidiphila (the order Desulfobacterales) with 96.3% 16S rRNA gene sequence similarity. Based on the phenotypic, genotypic and phylogenetic characteristics of the isolate, strain M08butT is considered to represent a novel species of the genus Desulfatitalea, with proposed name Desulfatitalea alkaliphila sp. nov. The type strain of Desulfatitalea alkaliphila is M08butT (= KCTC 25382T = VKM B-3560T = DSM 113909T = JCM 39202T = UQM 41473T).


Similar content being viewed by others
References
Alain K, Holler T, Musat F et al (2006) Microbiological investigation of methane- and hydrocarbon-discharging mud volcanoes in the Carpathian Mountains, Romania. Environ Microbiol 8:574–590. https://doi.org/10.1111/j.1462-2920.2005.00922.x
Altschul S, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Benson D, Boguski M, Lipman D et al (1999) GenBank. Nucleic Acids Res 27:38–43. https://doi.org/10.1093/nar/27.1.12
Beverige TJ, Lawrence JR, Murray RGE (2007) Sampling and staining for light microscopy. In: Reddy CA, Beverige TJ, Breznak JA, Marzluf GA, Scmidt TM et al (eds) Methods for general and molecular microbiology, 3rd edn. ASM Press, Washington, pp 19–33
Chen IA, Chu K, Palaniappan K et al (2021) The IMG/M data management and analysis system vol 6.0: new tools and advanced capabilities. Nucleic Acids Res 49(D1):D751–D763. https://doi.org/10.1093/nar/gkaa939
Chun J, Oren A, Ventosa A et al (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466. https://doi.org/10.1099/ijsem.0.002516
Frolova AA, Merkel AY, Novikov AA et al (2021a) Anaerotalea alkaliphila gen. nov., sp. nov., an alkaliphilic, anaerobic, fermentative bacterium isolated from a terrestrial mud volcano. Extremophiles 25:301–309. https://doi.org/10.1007/s00792-021-01229-w
Frolova AA, Merkel AY, Kuchierskaya AA et al (2021b) Pseudodesulfovibrio alkaliphilus, sp. nov., an alkaliphilic sulfate-reducing bacterium isolated from a terrestrial mud volcano. Antonie Van Leeuwenhoek 114:1387–1397. https://doi.org/10.1007/s10482-021-01608-5
Hartig C (2008) Rapid identification of fatty acid methyl esters using a multidimensional gas chromatography-mass spectrometry database. J Chromatogr 1177:159–169. https://doi.org/10.1016/j.chroma.2007.10.089
Higashioka Y, Kojima H, Watanabe M, Fukui M (2013) Desulfatitalea tepidiphila gen. nov., sp. nov., a sulfate-reducing bacterium isolated from tidal flat sediment. Int J Syst Evol Microbiol 63:761–765. https://doi.org/10.1099/ijs.0.043356-0
Hilton JC, Rajagopalan KV (1996) Molecular cloning of dimethyl sulfoxide reductase from Rhodobacter sphaeroides. Biochem Biophys Acta 1294:111–114. https://doi.org/10.1016/0167-4838(96)00015-5
Hoang DT, Chernomor O, von Haeseler A et al (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Huang X, Miller W (1991) A time efficient, linear space local similarity algorithm. Adv Appl Math 12:337–357. https://doi.org/10.1016/0196-8858(91)90017-D
Janto B, Ahmed A, Ito M et al (2011) Genome of alkaliphilic Bacillus pseudofirmus OF4 reveals adaptations that support the ability to grow in an external pH range from 7.5 to 11.4. Environ Microbiol 13:3289–3309. https://doi.org/10.1111/j.1462-2920.2011.02591.x
Khomyakova MA, Merkel AY, Petrova DA et al (2020) Alkalibaculum sporogenes sp. nov., isolated from a terrestrial mud volcano and emended description of the genus Alkalibaculum. Int J Syst Evol Microbiol 70:4914–4919. https://doi.org/10.1099/ijsem.0.004361
Khomyakova MA, Merkel AY, Kopitsyn DS, Slobodkin AI (2022) Pelovirga terrestris gen. nov., sp. nov, anaerobic, alkaliphilic, fumarate-, arsenate-, Fe (III)- and sulfur-reducing bacterium isolated from a terrestrial mud volcano. Syst Appl Microbiol 45(2):126304. https://doi.org/10.1016/j.syapm.2022.126304
Knittel K, Boetius A (2009) Anaerobic oxidation of methane: progress with an unknown process. Annu Rev Microbiol 63:311–334. https://doi.org/10.1146/annurev.micro.61.080706.093130
Krulwich TA, Sachs G, Padan E (2011) Molecular aspects of bacterial pH sensing and homeostasis. Nat Rev Microbiol 9:330–343. https://doi.org/10.1038/nrmicro2549
Kuever J, Rainey FA, Widdel F (2005) Order III. Desulfobacterales ord. nov. In: Brenner DJ, Krieg NR, Staley JT, Garrity GM (eds) Bergey’s manual of systematic bacteriology (the Proteobacteria), part C (The alpha-, beta-, delta-, and Epsilonproteobacteria), 2nd edn. Springer, New York, p 959
Lavrushin VY, Kopf A, Dehle A, Stepanets MI (2003) Formation of mudvolcanic fluids in Taman (Russia) and Kakhetia (Georgia): evidence from boron isotopes. Lithol Miner Resour 38:120–153. https://doi.org/10.1023/A:1023452025440
Malasarn D, Keeffe JR, Newman DK (2008) Characterization of the arsenate respiratory reductase from Shewanella sp. strain ANA-3. J Bacteriol 190:135–142. https://doi.org/10.1128/JB.01110-07
Mazzini A, Etiope G (2017) Mud volcanism: an updated review. Earth-Sci Rev 168:81–112
Merkel AY, Chernyh NA, Pimenov NV et al (2021) Diversity and metabolic potential of the terrestrial mud volcano microbial community with a high abundance of archaea mediating the anaerobic oxidation of methane. Life 11:953. https://doi.org/10.3390/life11090953
Mulkidjanian AY, Galperin MY, Makarova KS et al (2008) Evolutionary primacy of sodium bioenergetics. Biol Direct 3:13. https://doi.org/10.1186/1745-6150-3-13
Nakamura T, Yamada KD, Tomii K, Katoh K (2018) Parallelization of MAFFT for large-scale multiple sequence alignments. Bioinformatics 34:2490–2492. https://doi.org/10.1093/bioinformatics/bty121
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Overbeek R, Olson R, Pusch GD et al (2014) The seed and the rapid annotation of microbial genomes using subsystems technology (RAST). Nucleic Acids Res 42:D206–D214. https://doi.org/10.1093/nar/gkt1226
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP et al (2020) List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607–5612. https://doi.org/10.1099/ijsem.0.004332
Pereira IAC, Ramos AR, Grein F et al (2011) A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea. Front Microbiol 2:69. https://doi.org/10.3389/fmicb.2011.00069
Ren G, Ma A, Zhang Y, Deng Y et al (2018) Electron acceptors for anaerobic oxidation of methane drive microbial community structure and diversity in mud volcanoes. Environ Microbiol 20:2370–2385. https://doi.org/10.1111/1462-2920.14128
Ronquist F, Teslenko M, van der Mark P et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Schauder R, Widdel F, Fuchs G (1987) Carbon assimilation pathways in sulfate-reducing bacteria. II. Enzymes of a reductive citric acid cycle in the autotrophic Desulfobacter hydrogenophilus. Arch Microbiol 148:218–225. https://doi.org/10.1007/BF00414815
Slobodkin AI, Tourova TP, Kuznetsov BB et al (1999) Thermoanaerobacter siderophilus sp. nov., a novel dissimilatory Fe(III)-reducing anaerobic thermophilic bacterium. Int J Syst Bacteriol 49:1471–1478. https://doi.org/10.1099/00207713-49-4-1471
Slobodkin AI, Reysenbach AL, Slobodkina GB et al (2012) Thermosulfurimonas dismutans gen. nov., sp. nov., an extremely thermophilic sulfur-disproportionating bacterium from a deep-sea hydrothermal vent. Int J Syst Evol Microbiol 62:2565–2571. https://doi.org/10.1099/ijs.0.034397-0
Slobodkina GB, Baslerov RV, Novikov AA et al (2016) Inmirania thermothiophila gen. nov., sp. nov., a thermophilic, facultatively autotrophic, sulfur–oxidizing gammaproteobacterium isolated from a shallow–sea hydrothermal vent. Int J Syst Evol Microbiol 66:701–706. https://doi.org/10.1099/ijsem.0.000773
Søndergaard D, Pedersen CN, Greening C (2016) HydDB: a web tool for hydrogenase classification and analysis. Sci Rep 27:34212. https://doi.org/10.1038/srep34212
Takami H, Nakasone K, Takaki Y et al (2000) Complete genome sequence of the alkaliphilic bacterium Bacillus halodurans and genomic sequence comparison with Bacillus subtilis. Nucleic Acids Res 28:4317–4331. https://doi.org/10.1093/nar/28.21.4317
Tatusova T, DiCuccio M, Badretdin A et al (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Trüper HG, Schlegel HG (1964) Sulphur metabolism in Thiorhodaceae. I. Quantitative measurements on growing cells of Chromatium okenii. Antonie Van Leeuwenhoek 30:225–238. https://doi.org/10.1007/BF02046728
Tu TH, Wu LW, Lin YS et al (2017) Microbial community composition and functional capacity in a terrestrial ferruginous, sulfate-depleted mud volcano. Front Microbiol 8:2137. https://doi.org/10.3389/fmicb.2017.02137
Waite DW, Chuvochina M, Pelikan C et al (2020) Proposal to reclassify the proteobacterial classes Deltaproteobacteria and Oligoflexia, and the phylum Thermodesulfobacteria into four phyla reflecting major functional capabilities. Int J Syst Evol Microbiol 70:5972–6016. https://doi.org/10.1099/ijsem.0.004213
Wallace KK, Bao ZY, Dai H et al (1995) Purification of crotonyl-CoA reductase from Streptomyces collinus and cloning, sequencing and expression of the corresponding gene in Escherichia coli. Eur J Biochem 233:954–962. https://doi.org/10.1111/j.1432-1033.1995.954_3.x
Wolin EA, Wolin MJ, Wolfe RS (1963) Formation of methane by bacterial extracts. J Biol Chem 238:2882–2886
Yoon SH, Ha SM, Kwon SJ et al (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Funding
This work was funded by the Russian Science Foundation, grant number 22–14–00011 (MK, AM, AS, isolation and characterization) and by the Ministry of Science and Higher Education of the Russian Federation (genome sequencing and annotation).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Human and animal rights
The article does not contain any studies related to human participants or animals.
Additional information
Communicated by Oren.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The GenBank accession number for the 16S rRNA gene sequence of strain M08butT is OP380633. This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession JALJRB000000000.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khomyakova, M.A., Merkel, A.Y., Segliuk, V.S. et al. Desulfatitalea alkaliphila sp. nov., an alkalipilic sulfate- and arsenate- reducing bacterium isolated from a terrestrial mud volcano. Extremophiles 27, 12 (2023). https://doi.org/10.1007/s00792-023-01297-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00792-023-01297-0