Abstract
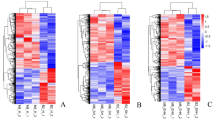
The current study attempts to investigate the differences in gene expression in longissimus thoracis muscles between sheep breeds acclimated to diverse environments. Changthangi sheep inhabits the cold arid plateau of Ladakh, at an altitude above 3000 m with prevalence of rarefied atmosphere. Muzzafarnagri sheep, on the other hand is found in the sub-tropical hot and humid plains at an altitude of about 250 m. Comparative transcriptomics was used to provide a molecular perspective of the differential adaptation of the two breeds. RNA sequencing data was generated from four biological replicates of the longissimus thoracis muscles from both breeds. The common genes expressed in both breeds were involved in muscle contraction and muscle fibre organization. The most significant pathways enriched in Changthangi muscles were glycogen metabolism, reduction of cytosolic Ca++ levels and NFE2L2 regulating anti-oxidant, while those in Muzzafarnagri were extracellular matrix organization and collagen formation. The hub genes identified in Changthangi were involved in hematopoiesis and HIF signaling pathway, suggesting the molecular acclimatization of Changthangi to the high altitude cold desert of Ladakh. The nodal genes discovered in Muzzafarnagri sheep were associated with the extracellular matrix which accentuates its significance in the development, growth and repair of muscles. The observed transcriptomic differences underscore the morphological and adaptive disparity between the two breeds. The candidate genes and pathways identified in this study will form the basis for future research on adaptation to high altitude and body size in small ruminants.





Similar content being viewed by others
Data availability
Data has been presented within the manuscript. The dataset generated in the study has been deposited in the NCBI (Short Read Archive Bioproject PRJNA995886).
References
Adeva-Andany MM, Gonzalez-Lucan M, Donapetry-Garcia C, Fernandez-Fernandez C, Ameneiros-Rodriguez E (2016) Glycogen metabolism in humans. BBA Clin 5:85–100. https://doi.org/10.1016/j.bbacli.2016.02.001
Ahlawat S, Arora R, Sharma R, Sharma U, Kaur M, Kumar A, Singh KV, Singh MK, Vijh RK (2020) Skin transcriptome profiling of Changthangi goats highlights the relevance of genes involved in pashmina production. Sci Rep 10(1):6050. https://doi.org/10.1038/s41598-020-63023-6
Al-Kershi S, Bhayadia R, Ng M, Verboon L, Emmrich S, Gack L, Schwarzer A, Strowig T, Heckl D, Klusmann JH (2019) The stem cell-specific long noncoding RNA HOXA10-AS in the pathogenesis of KMT2A-rearranged leukemia. Blood Adv 3(24):4252–4263. https://doi.org/10.1182/bloodadvances.2019032029
Arora R, Siddaraju NK, Sudarshan S, Fairoze MN, Kaur M, Sharma A, Girdhar Y, Sreesujatha RM, Devatkal SK, Ahlawat S, Vijh RK, Manjunatha SS (2019) Transcriptome profiling of longissimus thoracis muscles identifies highly connected differentially expressed genes in meat type sheep of India. PLoS ONE 14(6):e0217461. https://doi.org/10.1371/journal.pone.0217461
Arora R, Siddaraju NK, Manjunatha SS, Sudarshan S, Mohamed NF, Kumar A, Chhabra P, Kaur M, Sreesujatha RM, Ahlawat S, Vijh RK (2021) Muscle transcriptome provides the first insight into the dynamics of gene expression with progression of age in sheep. Sci Rep 11(1):22360. https://doi.org/10.1038/s41598-021-01848-5
Ayuso M, Fernandez A, Nunez Y, Benitez R, Isabel B, Barragan C, Fernandez AI, Rey AI, Medrano JF, Canovas A, Gonzalez-Bulnes A, Lopez-Bote C, Ovilo C (2015) Comparative analysis of muscle transcriptome between pig genotypes identifies genes and regulatory mechanisms associated to growth, fatness and metabolism. PLoS ONE 10(12):e0145162. https://doi.org/10.1371/journal.pone.0145162
BAHS-Basic Animal Husbandry Statistics (2022) Government of India, Ministry of Fisheries, Animal Husbandry &, Dairying, Department of Animal Husbandry &, Dairying, Krishi Bhavan, New Delhi, 1-141
Beard NA, Casarotto MG, Wei L, Varsányi M, Laver DR, Dulhunty AF (2005) Regulation of ryanodine receptors by calsequestrin: Effect of high luminal Ca2+ and phosphorylation. Biophys J 88(5):3444–3454
Bernard C, Cassar-Malek I, Le Cunff M, Dubroeucq H, Renand G, Hocquette JF (2007) New indicators of beef sensory quality revealed by expression of specific genes. J Agric Food Chem 55(13):5229–5237. https://doi.org/10.1021/jf063372l
Bhatia S, Arora R (2005) Biodiversity and Conservation of Indian Sheep Genetic Resources-An overview. Asian-australas J Anim Sci 18(10):1387–1402
Bigham AW, Lee FS (2014) Human high-altitude adaptation: forward genetics meets the HIF pathway. Genes Dev 28(20):2189–2204. https://doi.org/10.1101/gad.250167.114
Cai C, Li M, Zhang Y, Meng S, Yang Y, Gao P, Guo X, Cao G, Li B (2020) Comparative transcriptome analyses of Longissimus thoracic between Pig breeds Differing in muscle characteristics. Front Genet 11:526309. https://doi.org/10.3389/fgene.2020.526309
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin C (2014) CytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(4):1–7. https://doi.org/10.1186/1752-0509-8-S4-S11
Csapo R, Gumpenberger M, Wessner B (2020) Skeletal muscle Extracellular Matrix– what do we know about its composition, regulation, and physiological roles? A narrative review. Front Physiol 11:253. https://doi.org/10.3389/fphys.2020.00253
Dengler VL, Galbraith M, Espinosa JM (2013) Transcriptional regulation by hypoxia inducible factors. Crit Rev Biochem Mol Biol 49(1):1–15. https://doi.org/10.3109/10409238.2013.838205
Duchen MR (2000) Mitochondria and calcium: from cell signaling to cell death. J Physiol 529(1):57–68. https://doi.org/10.1111/j.1469-7793.2000.00057.x
Emara M, Turner AR, Allalunis-Turner J (2010) Hypoxic regulation of cytoglobin and neuroglobin expression in human normal and tumor tissues. Cancer Cell Int 10(1):1–16. https://doi.org/10.1186/1475-2867-10-33
Gan M, Shen L, Fan Y, Guo Z, Liu B, Chen L, Tang G, Jiang Y, Li X, Zhang S, Bai L, Zhu L (2019) High altitude adaptability and meat quality in tibetan pigs: a reference for local pork processing and genetic improvement. Anim (Basel) 3(12):1080. https://doi.org/10.3390/ani9121080
Garcia-Crespo D, Juste RA, Hurtado A (2005) Selection of ovine housekee** genes for normalization by real-time RT-PCR; analysis of PrP gene expression and genetic susceptibility to scrapie. BMC Vet Res 1(1):1–8
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID Bioinformatics resources. Nat Protoc 4(1):44–57
Huang H, Zhao Y, Shang X, Liu X, Ren H (2017) Expression of carbonic anhydrase III and skeletal muscle remodeling following selective denervation. Mol Med Rep 16(6):8289–8294
Huang X, Qu R, Ouyang J, Zhong S, Dai J (2020) An overview of the Cytoskeleton-Associated Role of PDLIM5. Front Physiol 11:975. https://doi.org/10.3389/fphys.2020.00975
Jassal B, Matthews L, Viteri G, Gong C, Lorente P, Fabregat A, Sidiropoulos K, Cook J, Gillespie M, Haw R, Loney F, May B, Milacic M, Rothfels K, Sevilla C, Shamovsky V, Shorser S, Varusai T, Weiser J, Wu G, Stein L, Hermjakob H, D’Eustachio P (2020) The reactome pathway knowledgebase. Nucleic Acids Res 48(D1):D498–D503. https://doi.org/10.1093/nar/gkz1031
Jiang J, Cao Y, Shan H, Wu J, Song X, Jiang Y (2021) The GWAS analysis of body size and Population Verification of related SNPs in Hu Sheep. Front Genet 12:642552. https://doi.org/10.3389/fgene.2021.642552
Jiao D, Ji K, Liu H, Wang W, Wu X, Zhou J, Zhang Y, Zhou H, Hickford JGH, Degen AA, Yang G (2021) Transcriptome analysis reveals genes involved in thermogenesis in two cold-exposed Sheep breeds. Genes (Basel) 12(3):375. https://doi.org/10.3390/genes12030375
Kamburov A, Pentchev K, Galicka H, Wierling C, Lehrach H, Herwig R (2011) ConsensusPathDB: toward a more complete picture of cell biology. Nucleic Acids Res 39(1):D712–717
Kanatous SB, Mammen PP, Rosenberg PB, Martin CM, White MD, Dimaio JM, Huang G, Muallem S, Garry DJ (2009) Hypoxia reprograms calcium signaling and regulates myoglobin expression. Am J Physiol Cell Physiol 296(3):C393–402. https://doi.org/10.1152/ajpcell.00428.2008
Kaur M, Kumar A, Naveen Kumar S, Mohamed Nadeem F, Ahlawat S, Vijh RK, Yadav A, Arora R (2020a) Exploring the skeletal muscle miRNAome of Bandur sheep using RNA sequencing. Indian J Anim Sci 90(8):118–122
Kaur M, Kumar A, Siddaraju NK, Fairoze MN, Chhabra P, Ahlawat S, Vijh RK, Yadav A, Arora R (2020b) Differential expression of miRNAs in skeletal muscles of Indian sheep with diverse carcass and muscle traits. Sci Rep 10(1):16332. https://doi.org/10.1038/s41598-020-73071-7
Khan NN, Ganai NA, Ahmad T, Majid R, Shanaz S, Shiekh FD (2022) Changthangi Sheep: the pride of Ladakh. Trends Agric Sci 2(3):49–51. https://doi.org/10.5281/zenodo.6535189
Kim H, Na YR, Kim SY, Yang EG (2016) Protein kinase C isoforms differentially regulate hypoxia-inducible Factor-1α Accumulation in Cancer cells. J Cell Biochem 117(3):647–658. https://doi.org/10.1002/jcb.25314
Kong BW, Hudson N, Seo D, Lee S, Khatri B, Lassiter K, Cook D, Piekarski A, Dridi S, Anthony N, Bottje W (2017) RNA sequencing for global gene expression associated with muscle growth in a single male modern broiler line compared to a foundational barred Plymouth Rock chicken line. BMC Genom 18(1):1–19. https://doi.org/10.1186/s12864-016-3471-y
Kowalczyk AP, Green KJ (2013) Structure, function, and regulation of desmosomes. Prog Mol Biol Transl Sci 116:95–118. https://doi.org/10.1016/B978-0-12-394311-8.00005-4
Kumar A, Kaur M, Ahlawat S, Sharma U, Singh MK, Singh KV, Chhabra P, Vijh RK, Yadav A, Arora R (2021) Transcriptomic diversity in longissimus thoracis muscles of Barbari and Changthangi goat breeds of India. Genom 113(4):1639–1646. https://doi.org/10.1016/j.ygeno.2021.04.019
Lin Y, Zhu J, Wang Y, Li Q, Lin S (2017) Identification of differentially expressed genes through RNA sequencing in goats (Capra hircus) at different postnatal stages. PLoS ONE 12(8):e0182602. https://doi.org/10.1371/journal.pone.0182602
Liu N, Yu Z, **ang S, Zhao S, Tjärnlund-Wolf A, **ng C, Zhang J, Wang X (2012) Transcriptional regulation mechanisms of hypoxia-induced neuroglobin gene expression. Biochem J 443(1):153–164. https://doi.org/10.1042/BJ20111856
Liu T, Tang J, Li X, Lin Y, Yang Y, Ma K, Hui Z, Ma H, Qin Y, Lei H, Yang Y (2022) The Key Network of mRNAs and miRNAs regulated by HIF1A in hypoxic Hepatocellular Carcinoma cells. Front Genet 13:857507. https://doi.org/10.3389/fgene.2022.857507
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25(4):402–408
Magnusson M, Brun AC, Miyake N, Larsson J, Ehinger M, Bjornsson JM, Wutz A, Sigvardsson M, Karlsson S (2007) HOXA10 is a critical regulator for hematopoietic stem cells and erythroid/megakaryocyte development. Blood 109(9):3687–3696. https://doi.org/10.1182/blood-2006-10-054676
Mathieu-Costello O (2001) Muscle adaptation to Altitude: tissue capillarity and capacity for aerobic metabolism. High Alt Med Biol 2(3):413–425. https://doi.org/10.1089/15270290152608598
Mukund K, Subramaniam S (2020) Skeletal muscle: a review of molecular structure and function, in health and disease. Wiley Interdiscip Rev Syst Biol Med 12(1):e1462. https://doi.org/10.1002/wsbm.1462
Neill T, Schaefer L, Iozzo RV (2012) Decorin: a guardian from the matrix. Am J Pathol 181(2):380–387. https://doi.org/10.1016/j.ajpath.2012.04.029
Neuber S, Mühmer M, Wratten D, Koch PJ, Moll R, Schmidt A (2010) The desmosomal plaque proteins of the plakophilin family. Dermatol Res Pract 101452. https://doi.org/10.1155/2010/101452
Oliveros JC (2007–2015) Venny. An interactive tool for comparing lists with Venn’s diagrams http://bioinfogp.cnb.csic.es/tools/venny/index.html
Olivieri J, Smaldone S, Ramirez F (2010) Fibrillin assemblies: extracellular determinants of tissue formation and fibrosis. Fibrogenesis Tissue Repair 3:1–8. https://doi.org/10.1186/1755-1536-3-24
Organ SL, Tsao MS (2011) An overview of the c-MET signaling pathway. Ther Adv Med Oncol 3(1):S7–S19. https://doi.org/10.1177/1758834011422556
Pareek CS, Sachajko M, Jaskowski JM, Herudzinska M, Skowronski M, Domagalski K, Szczepanek J, Czarnik U, Sobiech P, Wysocka D, Pierzchala M, Polawska E, Stepanow K, Ogłuszka M, Juszczuk-Kubiak E, Feng Y, Kumar D (2019) Comparative analysis of the liver transcriptome among cattle breeds using RNA-seq. Vet Sci 6(2):36. https://doi.org/10.3390/vetsci6020036
Pelletier J, Bellot G, Gounon P, Lacas-Gervais S, Pouysségur J, Mazure NM (2012) Glycogen synthesis is Induced in Hypoxia by the Hypoxia-Inducible factor and promotes Cancer Cell Survival. Front Oncol 2:18. https://doi.org/10.3389/fonc.2012.00018
Periasamy M, Kalyanasundaram A (2007) SERCA pump isoforms: their role in calcium transport and disease. Muscle Nerve Suppl 35(4):430–442
Rahman FA, Krause MP (2020) PAI-1, the Plasminogen System, and skeletal muscle. Int J Mol Sci 21(19):7066. https://doi.org/10.3390/ijms21197066
Sadkowski T, Jank M, Oprzadek J, Motyl T (2006) Age-dependent changes in bovine skeletal muscle transcriptomic profile. J Physiol Pharmacol 57(7):95
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Takada F, Vander Woude DL, Tong HQ, Thompson TG, Watkins SC, Kunkel LM, Beggs AH (2001) Myozenin: an alpha-actinin- and gamma-filamin-binding protein of skeletal muscle Z lines. Proc Natl Acad Sci 98(4):1595–1600. https://doi.org/10.1073/pnas.98.4.1595
Thomson J, Singh M, Eckersley A, Cain SA, Sherratt MJ, Baldock C (2018) Fibrillin microfibrils and elastic fibre proteins: functional interactions and extracellular regulation of growth factors. Semin Cell Dev Biol 89:109–117. https://doi.org/10.1016/j.semcdb.2018.07.016
Toth RK, Warfel NA (2017) Strange bedfellows: nuclear factor, erythroid 2-Like 2 (Nrf2) and hypoxia-inducible factor 1 (HIF-1) in Tumor Hypoxia. Antioxid (Basel) 6(2):27. https://doi.org/10.3390/antiox6020027
Wen W, Zhao Z, Li R, Guan J, Zhou Z, Luo X, Suman SP, Sun Q (2019) Skeletal muscle proteome analysis provides insights on high altitude adaptation of yaks. Mol Biol Rep 46(3):2857–2866. https://doi.org/10.1007/s11033-019-04732-8
Yadav DK, Arora R (2014) Genetic discrimination of Muzaffarnagri and Munjal sheep of northwestern semi-arid zone of India based on microsatellite markers and morphological traits. Indian J Anim Sci 84(5) https://epubs.icar.org.in/index.php/IJAnS/article/view/40666
Yoon D, Ponka P, Prchal JT (2011) Hypoxia. 5. Hypoxia and hematopoiesis. Am J Physiol Cell Physiol 300(6):C1215–C1222. https://doi.org/10.1152/ajpcell.00044.2011
Zhang W, Liu Y, Zhang H (2021) Extracellular matrix: an important regulator of cell functions and skeletal muscle development. Cell Biosci 11:65. https://doi.org/10.1186/s13578-021-00579-4
Zhu L, Li M, Li X et al (2009) Distinct expression patterns of genes associated with muscle growth and adipose deposition in tibetan pigs: a possible adaptive mechanism for high altitude conditions. High Alt Med Biol 10(1):45–55. https://doi.org/10.1089/ham.2008.1042
Zhu W, Lin Y, Liao H, Wang Y (2015) Selection of reference genes for gene expression studies related to intramuscular Fat Deposition in Capra hircus skeletal muscle. PLoS ONE 10(3):e0121280. https://doi.org/10.1371/journal.pone.0121280
Acknowledgements
We are grateful to ICAR-Consortium Research Platform-Genomics for financial support; Director, ICAR- National Bureau of Animal Genetic Resources (NBAGR), Karnal and Indian Council of Agricultural Research (ICAR), New Delhi for providing necessary facilities.
Funding
The research was supported by grant of funds under ICAR-Consortium Research Platform on Genomics.
Author information
Authors and Affiliations
Contributions
RA: conceptualization and supervision. MAM and MKS: resources. AK, MK and RG: data curation. PC, SA, RS and RA: formal analyses; MK, RA: Writing - Original Draft and RA: Writing - Review & Editing. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethical approval and consent to participate
All methods were carried out in accordance with guidelines of CPCSEA (Committee for the Purpose of Control and Supervision on Experiments on Animals), with due approval from the Institutional Animal Ethics Committee of ICAR-National Bureau of Animal Genetic Resources, Karnal (F.No. NBAGR/IAEC/2017, dated 21.01.2017).
Competing of interest
None of the authors’ have any competing interests, including financial and non-financial.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Arora, R., Kaur, M., Kumar, A. et al. Skeletal muscle transcriptomics of sheep acclimated to cold desert and tropical regions identifies genes and pathways accentuating their diversity. Int J Biometeorol (2024). https://doi.org/10.1007/s00484-024-02708-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00484-024-02708-3