Abstract
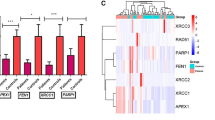
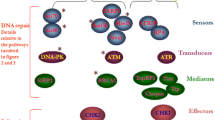
Hematological malignancies (HM) constitute a variety of cancers originating in blood, bone marrow (BM), and lymphatic systems. During the last two decades, the incidence of HM has dramatically increased worldwide. The etiology of HM is still debatable. Genetic instability is a major risk factor for HM. DDR network is a complex signal transduction cellular machinery that detects DNA damage and activates cellular repair factors, thus maintaining genomic integrity. DDR network detects a variety of DNA damage and triggers the activation of cell cycle control, DNA repair, senescence, and apoptosis. Among the DNA repairing pathways, the DNA damage response (DDR) pathway includes DNA damage signaling apparatus such as ATM and ATR genes. ATM tends to detect double-strand breaks (DSBs) while ATR detects single-strand DNA (ssDNA). The study was conducted to observe the expression deregulations of DNA damage response (DDR) pathway genes (ATM, ATR) at mRNA level in 200 blood cancer patients and 200 controls. The real-time PCR was used to analyze the expression of the target genes. The expression results showed statistically significant downregulation of ATM (p < 0.0001) and ATR (p < 0.0001) genes in blood cancer patients vs. controls. Moreover, a significant downregulation of ATM (p < 0.0001) and ATR (p < 0.0001) was obtained in chemotherapy-treated patients vs. healthy controls. The results suggest that dysregulation in ATM and ATR genes may be associated with increased blood cancer risk.








Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article.
References
Ait-Aissa K, Blaszak S, Beutner G et al (2019) Mitochondrial oxidative phosphorylation defect in the heart of subjects with coronary artery disease. nature.com
Angèle S, Treilleux I, Brémond A et al (2003) Altered expression of DNA double-strand break detection and repair proteins in breast carcinomas. Histopathology 43:347–353. https://doi.org/10.1046/J.1365-2559.2003.01713.X
Bakkenist C, Nature MK (2003) undefined DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. nature.com
Brown EJ, Baltimore D (2003) Essential and dispensable roles of ATR in cell cycle arrest and genome maintenance. Genes Dev 17:615–628. https://doi.org/10.1101/GAD.1067403
Cimprich KA, Cortez D (2008) ATR: an essential regulator of genome integrity. Nat Rev Mol Cell Biol 9:616–627. https://doi.org/10.1038/NRM2450
Coquerelle TM, Weibezahn KF, Lücke-Huhle C (1987) Rejoining of double strand breaks in normal human and ataxia-telangiectasia fibroblasts after exposure to 60 Co γ-rays, 241 Am α-particles or bleomycin. Int J Radiat Biol Relat Stud Phys Chem Med 51:209–218. https://doi.org/10.1080/09553008714550711
Cornforth MN, Bedford JS (1985) On the nature of a defect in cells from individuals with ataxia-telangiectasia. Science 227:1589–1591. https://doi.org/10.1126/SCIENCE.3975628
Dai Y, Grant S (2010) New insights into checkpoint kinase 1 in the DNA damage response signaling network. Clin Cancer Res 16:376–383. https://doi.org/10.1158/1078-0432.CCR-09-1029
Foray N, Priestley A, Alsbeih G et al (1997) Hypersensitivity of ataxia telangiectasia fibroblasts to ionizing radiation is associated with a repair deficiency of DNA double-strand breaks. Int J Radiat Biol 72:271–283. https://doi.org/10.1080/095530097143266
Grant RC, Al-Sukhni W, Borgida AE et al (2013) Exome sequencing identifies nonsegregating nonsense ATM and PALB2 variants in familial pancreatic cancer. Hum Genomics. https://doi.org/10.1186/1479-7364-7-11
Huntoon CJ, Flatten KS, Wahner Hendrickson AE et al (2013) ATR inhibition broadly sensitizes ovarian cancer cells to chemotherapy independent of BRCA status. Cancer Res 73:3683–3691. https://doi.org/10.1158/0008-5472.CAN-13-0110
Khanna KK, Chenevix-Trench G (2004) ATM and genome maintenance: defining its role in breast cancer susceptibility. J Mammary Gland Biol Neoplasia 9:247–262. https://doi.org/10.1023/B:JOMG.0000048772.92326.A1
Kovalev S, Mateen A, Zaika AI et al (2000) Lack of defective expression of the ATM gene in sporadic breast cancer tissues and cell lines. Int J Oncol 16:825–831. https://doi.org/10.3892/IJO.16.4.825
Kumar A, Mazzanti M, Mistrik M et al (2014) ATR mediates a checkpoint at the nuclear envelope in response to mechanical stress. Cell 158:633–646. https://doi.org/10.1016/J.CELL.2014.05.046
Lavin MF, Davidson M (1981) Repair of strand breaks in superhelical DNA of ataxia telangiectasia lymphoblastoid cells. J Cell Sci 48:383–391. https://doi.org/10.1242/JCS.48.1.383
Lavin MF, Shiloh Y (1997) The genetic defect in ataxia-telangiectasia. Annu Rev Immunol 15:177–202. https://doi.org/10.1146/ANNUREV.IMMUNOL.15.1.177
Lehle S, Hildebrand D, Merz B et al (2014) LORD-Q: a long-run real-time PCR-based DNA-damage quantification method for nuclear and mitochondrial genome analysis. academic.oup.com
Li Y, Carty MP, Oakley GG et al (2001) Expression of ATM in ataxia telangiectasia fibroblasts rescues defects in DNA double-strand break repair in nuclear extracts. Environ Mol Mutagen 37:128–140. https://doi.org/10.1002/EM.1020
Lieber MR (2010) The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem 79:181–211. https://doi.org/10.1146/ANNUREV.BIOCHEM.052308.093131
Lin K, Adamson J, Johnson G et al (2021) Functional analysis of the ATM-p53-p21 pathway in the LRF CLL4 trial: blockade at the level of p21 is associated with short response duration. AACR
Liu S, Kurzrock R (2014) Toxicity of targeted therapy: implications for response and impact of genetic polymorphisms. Cancer Treat Rev 40:883–891. https://doi.org/10.1016/J.CTRV.2014.05.003
Mehmood A, Kayani MA, Ahmed MW et al (2020) Association between single nucleotide polymorphisms of DNA damage response pathway genes and increased risk in breast cancer. Future Oncol 16:1977–1995. https://doi.org/10.2217/FON-2020-0086
Mochan T, Venere M, Jr RD et al (2003) 53BP1 and NFBD1/MDC1-Nbs1 function in parallel interacting pathways activating ataxia-telangiectasia mutated (ATM) in response to DNA damage. AACR
Mrozik KM, Blaschuk OW, Cheong CM et al (2018) N-cadherin in cancer metastasis, its emerging role in haematological malignancies and potential as a therapeutic target in cancer. BMC Cancer. https://doi.org/10.1186/S12885-018-4845-0
Royo H, Prosser H et al (2013) ATR acts stage specifically to regulate multiple aspects of mammalian meiotic silencing. genesdev.cshlp.org. https://doi.org/10.1101/gad.219477.113
Saldivar JC, Cortez D, Cimprich KA (2017) The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat Rev Mol Cell Biol 18:622–636. https://doi.org/10.1038/NRM.2017.67
Shechter D, Costanzo V, Gautier J (2004) ATR and ATM regulate the timing of DNA replication origin firing. Nat Cell Biol 6(7):648–655
Sirbu BM and Cortez D (2013) In DC-CSH perspectives, 2013 DNA damage response: three levels of DNA repair regulation. cshperspectives.cshlp.org. https://doi.org/10.1101/cshperspect.a012724
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/CAAC.21660
Taylor AMR, Byrd PJ (2005) Molecular pathology of ataxia telangiectasia. J Clin Pathol 58:1009–1015. https://doi.org/10.1136/JCP.2005.026062
Vo QN, Kim WJ, Cvitanovic L et al (2004) The ATM gene is a target for epigenetic silencing in locally advanced breast cancer. Oncogene 23:9432–9437. https://doi.org/10.1038/SJ.ONC.1208092
Wang H, Wang H, Powell SN et al (2004) ATR affecting cell radiosensitivity is dependent on homologous recombination repair but independent of nonhomologous end joining. Cancer Res 64:7139–7143. https://doi.org/10.1158/0008-5472.CAN-04-1289
Wang H, Wang S, Shen L et al (2010) CHK2 down-regulation by promoter hypermethylation in human bulk gliomas. Life Sci 86:185–191. https://doi.org/10.1016/J.LFS.2009.11.023
Wright JA, Keegan KS, Herendeen DR et al (1998) Protein kinase mutants of human ATR increase sensitivity to UV and ionizing radiation and abrogate cell cycle checkpoint control. Proc Natl Acad Sci USA 95:7445–7450. https://doi.org/10.1073/PNAS.95.13.7445
Ye C, Dai Q, Lu W et al (2007) Two-stage case-control study of common ATM gene variants in relation to breast cancer risk. Breast Cancer Res Treat 106:121–126. https://doi.org/10.1007/S10549-006-9473-8
Zou L, Elledge SJ (2003) Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science 300:1542–1548. https://doi.org/10.1126/SCIENCE.1083430
Acknowledgements
Higher Education Commission of Pakistan (HEC) and CAMSATS University, Islamabad, provided financial and infrastructural support to the authors. The authors thank the patients and the personnel of the Pakistan Institute of Medical Sciences (PIMS) in Islamabad for their participation in study.
Funding
There was no specific grant for this research from any government, commercial or non-profit funding body.
Author information
Authors and Affiliations
Contributions
All of the authors read and approved the final version of manuscript. AP, AUK, and AM collected and isolated the RNA samples. AP and AN performed the expression analysis study. AP and IM performed statistical analyses of the data and draft of the manuscript. MAK and IM supervised the project and provided critical revisions. All the authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
There are no competing interests for the study.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Parvez, A., Mahjabeen, I., Mehmood, A. et al. Expression variations of DNA damage response genes ATM and ATR in blood cancer patients. Mol Genet Genomics 298, 1173–1183 (2023). https://doi.org/10.1007/s00438-023-02043-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-023-02043-z