Abstract
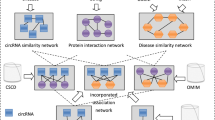
Circular RNAs (circRNAs) are a special class of non-coding RNAs with covalently closed-loop structures. Studies prove that circRNAs perform critical roles in various biological processes, and the aberrant expression of circRNAs is closely related to tumorigenesis. Therefore, identifying potential circRNA-disease associations is beneficial to understand the pathogenesis of complex diseases at the circRNA level and helps biomedical researchers and practitioners to discover diagnostic biomarkers accurately. However, it is tremendously laborious and time-consuming to discover disease-related circRNAs with conventional biological experiments. In this study, we develop an integrative framework, called iCDA-CMG, to predict potential associations between circRNAs and diseases. By incorporating multi-source prior knowledge, including known circRNA-disease associations, disease similarities and circRNA similarities, we adopt a collective matrix completion-based graph learning model to prioritize the most promising disease-related circRNAs for guiding laborious clinical trials. The results show that iCDA-CMG outperforms other state-of-the-art models in terms of cross-validation and independent prediction. Moreover, the case studies for several representative cancers suggest the effectiveness of iCDA-CMG in screening circRNA candidates for human diseases, which will contribute to elucidating the pathogenesis mechanisms and unveiling new opportunities for disease diagnosis and targeted therapy.




Similar content being viewed by others
References
Cai D, He X, Han J, Huang TS (2011) Graph regularized nonnegative matrix factorization for data representation. IEEE Trans Pattern Anal Mach Intell 33(8):1548–1560
Chen L, Huang C, Wang X, Shan G (2015) Circular RNAs in eukaryotic cells. Curr Genom 16(5):312–318
Couto FM, Silva MJ, Coutinho PM (2007) Measuring semantic similarity between Gene Ontology terms. Data Knowl Eng 61(1):137–152
Ding H, Takigawa I, Mamitsuka H, Zhu SF (2014) Similarity-based machine learning methods for predicting drug-target interactions: a brief review. Brief Bioinform 15(5):734–747
Fan C, Lei X, Fang Z, Jiang Q, Wu FX (2018) CircR2Disease: a manually curated database for experimentally supported circular RNAs associated with various diseases. Database 2018:1–8
Hansen TB, Kjems J, Damgaard CK (2013) Circular RNA and miR-7 in cancer. Cancer Res 73(18):5609–5612
He R, Liu P, **e X, Zhou Y, Liao Q, **ong W, Li X, Li G, Zeng Z, Tang H (2017) circGFRA1 and GFRA1 act as ceRNAs in triple negative breast cancer by regulating miR-34a. J Exp Clin Cancer Res 36(1):145
Hu B, **an Z, Zou Q, Zhang D, Su D, Yao J, Ren D (2020) CircFAT1 suppresses colorectal cancer development through regulating miR-520b/UHRF1 Axis or miR-302c-3p/UHRF1 Axis. Cancer Biotherapy Radiopharm. https://doi.org/10.1089/cbr.2019.3291
Huang XY, Zhang PF, Wei CY, Peng R, Lu JC, Gao C, Cai JB, Yang X, Fan J, Ke AW, Zhou J, Shi GM (2020) Circular RNA circMET drives immunosuppression and anti-PD1 therapy resistance in hepatocellular carcinoma via the miR-30-5p/snail/DPP4 axis. Mol Cancer 19(1):92
L′Abbate A, Tolomeo D, Cifola I, Severgnini M, Turchiano A, Augello B, Squeo G, D′Addabbo P, Traversa D, Daniele G, Lonoce A, Pafundi M, Carella M, Palumbo O, Dolnik A, Muehlematter D, Schoumans J, Van Roy N, De Bellis G, Martinelli G, Merla G, Bullinger L, Haferlach C and Storlazzi CT, (2018) MYC-containing amplicons in acute myeloid leukemia: genomic structures, evolution, and transcriptional consequences. Leukemia 32(10):2152–2166
Ledford H (2013) Circular RNAs throw genetics for a loop. Nature 494(7438):415
Lei X, Bian C (2020) Integrating random walk with restart and k-Nearest Neighbor to identify novel circRNA-disease association. Sci Rep 10(1):1943
Lei X, Fang Z, Chen L, Wu FX (2018) PWCDA: path weighted method for predicting circRNA-disease associations. Int J Mol Sci 19(11):3410
Lei X, Fang Z, Guo L (2019) Predicting circRNA–disease associations based on improved collaboration filtering recommendation system with multiple data. Front Genet 10:897
Li G, Luo J, **ao Q, Liang C, Ding P (2018) Predicting microRNA-disease associations using label propagation based on linear neighborhood similarity. J Biomed Inform 82:169–177
Li X, Zhang Z, Jiang H, Li Q, Wang R, Pan H, Niu Y, Liu F, Gu H, Fan X, Gao J (2018) Circular RNA circPVT1 Promotes proliferation and invasion through sponging miR-125b and activating E2F2 signaling in non-small cell lung cancer. Cell Physiol Biochem 51(5):2324–2340
Lu H, Han X, Ren J, Ren K, Li Z, Sun Z (2020) Circular RNA HIPK3 induces cell proliferation and inhibits apoptosis in non-small cell lung cancer through sponging miR-149. Cancer Biol Ther 21(2):113–121
Luo J, **ao Q (2017) A novel approach for predicting microRNA-disease associations by unbalanced bi-random walk on heterogeneous network. J Biomed Inform 66:194–203
Luo JW, **ao Q, Liang C, Ding PJ (2017) Predicting MicroRNA-disease associations using Kronecker regularized least squares based on heterogeneous Omics data. Ieee Access 5:2503–2513
Meng X, Hu D, Zhang P, Chen Q, Chen M (2019) CircFunBase: a database for functional circular RNAs. Database. https://doi.org/10.1093/database/baz003
Pan B, Qin J, Liu X, He B, Wang X, Pan Y, Sun H, Xu T, Xu M, Chen X, Xu X, Zeng K, Sun L, Wang S (2019) Identification of serum exosomal hsa-circ-0004771 as a novel diagnostic biomarker of colorectal cancer. Frontiers in genetics 10:1096
Qiu M, **a W, Chen R, Wang S, Xu Y, Ma Z, Xu W, Zhang E, Wang J, Fang T, Hu J, Dong G, Yin R, Wang J, Xu L (2018) The Circular RNA circPRKCI promotes tumor growth in lung adenocarcinoma. Cancer Res 78(11):2839–2851
Shen Z, Lin Y, Zou Q (2020) Transcription factors-DNA interactions in rice: identification and verification. Brief Bioinform 21(3):946–956
Su R, Liu X, Wei L, Zou Q (2019) Deep-Resp-Forest: A deep forest model to predict anti-cancer drug response. Methods 166:91–102
Tian F, Wang Y, **ao Z, Zhu X (2017) Circular RNA CircHIPK3 promotes NCI-H1299 and NCI-H2170 Cell proliferation through miR-379 and its target IGF1. Chin J Lung Cancer 20(7):459–467
Vicens Q, Westhof E (2014) Biogenesis of Circular RNAs. Cell 159(1):13–14
Vo JN, Cieslik M, Zhang Y, Shukla S, **ao L, Zhang Y, Wu YM, Dhanasekaran SM, Engelke CG, Cao X, Robinson DR, Nesvizhskii AI, Chinnaiyan AM (2019) The landscape of circular RNA in cancer. Cell 176(4):869–881
Wang D, Wang J, Lu M, Song F, Cui Q (2010) Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 26(13):1644–1650
Wang Z, Su M, **ang B, Zhao K, Qin B (2019) Circular RNA PVT1 promotes metastasis via miR-145 sponging in CRC. Biochem Biophys Res Commun 512(4):716–722
**ao Q, Dai JH, Luo JW, Fujita H (2019) Multi-view manifold regularized learning-based method for prioritizing candidate disease miRNAs. Knowl-Based Syst 175:118–129
**ao Q, Luo J, Dai J (2019) Computational prediction of human disease- associated circRNAs based on manifold regularization learning framework. IEEE J Biomed Health Inform 23(6):2661–2669
**ao Q, Luo J, Liang C, Cai J, Li G, Cao B (2019) CeModule: an integrative framework for discovering regulatory patterns from genomic data in cancer. BMC Bioinform 20(1):67–67
**ao Q, Luo J, Liang C, Li G, Cai J, Ding P, Liu Y (2020) Identifying lncRNA and mRNA co-expression modules from matched expression data in ovarian cancer. IEEE/ACM Trans Comput Biol Bioinform 17(2):623–634
**ao Q, Luo JW, Liang C, Cai J, Ding PJ (2018) A graph regularized non-negative matrix factorization method for identifying microRNA-disease associations. Bioinformatics 34(2):239–248
**ao Q, Yu H, Zhong J, Liang C, Luo J (2020) An in-silico method with graph-based multi-label learning for large-scale prediction of circRNA-disease associations. Genomics 112(5):3407–3415
**ao Q, Zhang N, Luo J, Dai J, Tang X (2020) Adaptive multi-source multi-view latent feature learning for inferring potential disease-associated miRNAs. Brief Bioinform. https://doi.org/10.1093/bib/bbaa028
Yan C, Wang J, Wu FX (2018) DWNN-RLS: regularized least squares method for predicting circRNA-disease associations. BMC Bioinformatics 19(Suppl 19):520
Yang X, Liu L, Zou H, Zheng YW, Wang KP (2019) circZFR promotes cell proliferation and migration by regulating miR-511/AKT1 axis in hepatocellular carcinoma. Digest Liver Dis 51(10):1446–1455
Yu SP, Liang C, **ao Q, Li GH, Ding PJ, Luo JW (2018) GLNMDA: a novel method for miRNA-disease association prediction based on global linear neighborhoods. RNA Biol 15(9):1215–1227
Zeng X, Liu L, Lü L, Zou Q (2018) Prediction of potential disease-associated microRNAs using structural perturbation method. Bioinformatics 34(14):2425–2432
Zhang P, Zuo Z, Shang W, Wu A, Bi R, Wu J, Li S, Sun X, Jiang L (2017) Identification of differentially expressed circular RNAs in human colorectal cancer. Tumour Biol 39(3):1010428317694546
Zhang W, Yu C, Wang X, Liu F (2019) Predicting CircRNA-disease associations through linear neighborhood label propagation method. IEEE Access 7:83474–83483
Zhang Y, Zhao H, Zhang L (2018) Identification of the tumor-suppressive function of circular RNA FOXO3 in non-small cell lung cancer through sponging miR-155. Mol Med Rep 17(6):7692–7700
Zhao J, Li L, Wang Q, Han H, Zhan Q, Xu M (2017) CircRNA expression profile in early-stage lung adenocarcinoma patients. Cellul Physiol Biochem 44(6):2138–2146
Zhao Z, Wang K, Wu F, Wang W, Zhang K, Hu H, Liu Y, Jiang T (2018) circRNA disease: a manually curated database of experimentally supported circRNA-disease associations. Cell Death Dis 9(5):475
Zheng K, You Z-H, Li J-Q, Wang L, Guo Z-H, Huang Y-A (2020) iCDA-CGR: identification of circRNA-disease associations based on Chaos Game Representation. Plos Comput Biol 16(5):e1007872
Zhu X, Wang X, Wei S, Chen Y, Chen Y, Fan X, Han S, Wu G (2017a) hsa_circ_0013958: a circular RNA and potential novel biomarker for lung adenocarcinoma. FEBS J 284(14):2170
Zhu X, Wang X, Wei S, Chen Y, Chen Y, Fan X, Han S, Wu G (2017b) hsa_circ_0013958: a circular RNA and potential novel biomarker for lung adenocarcinoma. FEBS J 284(14):2170–2182
Zou Q, Li J, Song L, Zeng X, Wang G (2016) Similarity computation strategies in the microRNA-disease network: a survey. Brief Funct Genomics 15(1):55–64
Acknowledgements
The authors are grateful to the members of Yide Yang’s group (School of Medicine, Hunan Normal University) for helpful discussions and valuable comments.
Funding
This work was supported in part by the National Natural Science Foundation of China (Grant No. 62002116), the Hunan Provincial Natural Science Foundation of China (2020JJ5373), the Scientific Research Fund of Hunan Provincial Education Department (No.20B348), and the Hunan Provincial Science & Technology Project Foundation (2018TP1018, 2018RS3065).
Author information
Authors and Affiliations
Contributions
QX and JWL conceived the model, prepared the data sets, performed and analyzed experiments, and wrote the manuscript. JCZ and XWT analyzed the prediction results. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
**ao, Q., Zhong, J., Tang, X. et al. iCDA-CMG: identifying circRNA-disease associations by federating multi-similarity fusion and collective matrix completion. Mol Genet Genomics 296, 223–233 (2021). https://doi.org/10.1007/s00438-020-01741-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-020-01741-2