Abstract
Background
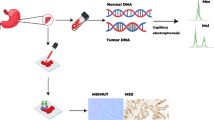
Microsatellite instability (MSI) is one of the essential tumor biomarkers for cancer treatment and prognosis. The presence of more significant PD-L1 expression on the surface of tumor cells in endometrial cancer with MSI suggests that MSI may be a promising biomarker for anti-PD-1/PD-L1 immunotherapy. However, the conventional testing methods are labor-intensive and expensive for patients.
Methods
Inspired by classifiers for MSI based on fast and low-cost deep-learning methods in previous investigations, a new architecture for MSI classification based on an attention module is proposed to extract features from pathological images. Especially, slide-level microsatellite status will be obtained by the bag of words method to aggregate probabilities predicted by the proposed model. The H&E-stained whole slide images (WSIs) from The Cancer Genome Atlas endometrial cohort are collected as the dataset. The performances of the proposed model were primarily evaluated by the area under the receiver-operating characteristic curve, accuracy, sensitivity, and F1-Score.
Results
On the randomly divided test dataset, the proposed model achieved an accuracy of 0.80, a sensitivity of 0.857, a F1-Score of 0.826, and an AUROC of 0.799. We then visualize the results of the microsatellite status classification to capture more specific morphological features, hel** pathologists better understand how deep learning performs the classification.
Conclusions
This study implements the prediction of microsatellite status in endometrial cancer cases using deep-learning methods directly from H&E-stained WSIs. The proposed architecture can help the model capture more valuable features for classification. In contrast to current laboratory testing methods, the proposed model creates a more convenient screening tool for rapid automated testing for patients. This method can potentially be a clinical method for detecting the microsatellite status of endometrial cancer.










Similar content being viewed by others
Data availability
The public datasets to support the results of this subject can be gained from TCGA (https://portal.gdc.cancer.gov/). The original training images for tissue classification are available at https://zenodo.org/record/2530789.
References
Albarqouni S, Baur C, Achilles F et al (2016) AggNet: deep learning from crowds for mitosis detection in breast cancer histology images. IEEE Trans Med Imaging 35:1313–1321
Astley J, Wild J, Tahir B (2021) Deep learning in structural and functional lung image analysis. Br J Radiol 95:20201107
Bonneville R, Krook MA, Kautto EA et al (2017) Landscape of microsatellite instability across 39 cancer types. JCO Precis Oncol. https://doi.org/10.1200/po.17.00073
Bray F, Ferlay J et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424
Cao R, Yang F, Ma SC et al (2020) Development and interpretation of a pathomics-based model for the prediction of microsatellite instability in colorectal cancer. Theranostics 10:11080–11091. https://doi.org/10.7150/thno.49864
Concin N, Matias-Guiu X, Vergote I et al (2021) ESGO/ESTRO/ESP guidelines for the management of patients with endometrial carcinoma. Int J Gynecol Cancer 31:12–39. https://doi.org/10.1136/ijgc-2020-002230
Coudray N, Moreira AL, Sakellaropoulos T et al (2017) Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat Med 24:1559–1567
Crosbie EJ, Ryan N, Arends MJ et al (2019) The Manchester International Consensus Group recommendations for the management of gynecological cancers in Lynch syndrome. Genet Med 21:2390–2400
Cui M, Zhang DY (2021) Artificial intelligence and computational pathology. Lab Investig 101:412–422
Dabass M, Vashisth S, Vig R (2022) A convolution neural network with multi-level convolutional and attention learning for classification of cancer grades and tissue structures in colon histopathological images. Comput Biol Med 147:105680
Dijk B, Santos BF, Pita JP (2016) The recoverable robust stand allocation problem: a GRU airport case study. Or Spectrum 41:615–639
Doll KM, Tseng J, Denslow SA et al (2014) High-grade endometrial cancer: revisiting the impact of tumor size and location on outcomes. Gynecol Oncol 132:44–49. https://doi.org/10.1016/j.ygyno.2013.10.023
Echle A, Grabsch HI, Quirke P et al (2020) Clinical-grade detection of microsatellite instability in colorectal tumors by deep learning. Gastroenterology 159:1406-1416.e11. https://doi.org/10.1053/j.gastro.2020.06.021
Farajzadeh N, Sadeghzadeh N, Hashemzadeh M (2022) A fully-convolutional residual encoder-decoder neural network to localize breast cancer on histopathology images. Comput Biol Med 147:105698
Fleming M, Ravula S, Tatishchev SF, Wang HL (2012) Colorectal carcinoma: pathologic aspects. J Gastrointest Oncol 3:153
Gabriel SB, Cibulskis K, Lander E et al (2013) Integrated genomic characterization of endometrial carcinoma. Nature 497:67–73
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: 2016 IEEE conference on computer vision and pattern recognition (CVPR), pp 770–778
Hu J, Shen L, Albanie S et al (2020) Squeeze-and-excitation networks. IEEE Trans Pattern Anal Mach Intell 42:2011–2023
Jaderberg M, Simonyan K, Zisserman A, Kavukcuoglu K (2015) Spatial transformer networks. MIT Press, Cambridge
Jia D, Wei D, Socher R et al (2009) ImageNet: a large-scale hierarchical image database, pp 248–255
Kather JN, Krisam J, Charoentong P et al (2019a) Predicting survival from colorectal cancer histology slides using deep learning: a retrospective multicenter study. PLoS Med 16:e1002730
Kather JN, Pearson AT, Halama N et al (2019b) Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat Med 25:1054–1056. https://doi.org/10.1038/s41591-019-0462-y
Kong J, Huang J, Yu H et al (2019) RNN-based default logic for route planning in urban environments. Neurocomputing 338:307–320
Krizhevsky A, Sutskever I, Hinton GE (2012) ImageNet classification with deep convolutional neural networks. In: Proceedings of the 25th international conference on neural information processing systems, vol 1. Curran Associates Inc., Red Hook, pp 1097–1105
Lee SH, Song IH, Jang HJ (2021) Feasibility of deep learning-based fully automated classification of microsatellite instability in tissue slides of colorectal cancer. Int J Cancer 149:728–740. https://doi.org/10.1002/ijc.33599
Li W, Li J, Sarma KV et al (2019) Path R-CNN for prostate cancer diagnosis and Gleason grading of histological images. IEEE Trans Med Imaging 38:945–954
Luchini C, Bibeau F, Ligtenberg MJL et al (2019) ESMO recommendations on microsatellite instability testing for immunotherapy in cancer, and its relationship with PD-1/PD-L1 expression and tumour mutational burden: a systematic review-based approach. Ann Oncol 30:1232–1243. https://doi.org/10.1093/annonc/mdz116
Meyer LA, Broaddus RR, Lu KH (2009) Endometrial cancer and lynch syndrome: clinical and pathologic considerations. Cancer Control 16:14–22. https://doi.org/10.1177/107327480901600103
Paula A, Delair DF, Ferrando L et al (2021) Genetic and molecular subtype heterogeneity in newly diagnosed early- and advanced-stage endometrial cancer. Gynecol Oncol 161:535–544
Raffone A, Troisi J, Boccia D et al (2020) Metabolomics in endometrial cancer diagnosis: a systematic review. Acta Obstet Gynecol Scand 99:1135–1146
Reinhard E, Ashikhmin M, Gooch B, Shirley P (2002) Color transfer between images. IEEE Comput Graph Appl 21:34–41
Saeed OAM, Mann SA, Luchini C et al (2021) Evaluating mismatch repair deficiency for solid tumor immunotherapy eligibility: immunohistochemistry versus microsatellite molecular testing. Hum Pathol 115:10–18
Saillard C, Dehaene O, Marchand T et al (2021) Self supervised learning improves dMMR/MSI detection from histology slides across multiple cancers, pp 1–16
Sapitri AI, Nurmaini S, Rachmatullah MN et al (2023) Deep learning-based real time detection for cardiac objects with fetal ultrasound video. Inform Med Unlocked 36:101150
Simonyan K, Zisserman A (2015) Very deep convolutional networks for large-scale image recognition
Sinicrope FA, Sargent DJ (2012) Molecular pathways: microsatellite instability in colorectal cancer: prognostic, predictive, and therapeutic implications. Clin Cancer Res 18:1506–1512. https://doi.org/10.1158/1078-0432.CCR-11-1469
Song J, Chai YJ, Masuoka H et al (2019) Ultrasound image analysis using deep learning algorithm for the diagnosis of thyroid nodules. Medicine (baltimore) 98:e15133
Tan X, Su AT, Hajiabadi H et al (2021) Applying machine learning for integration of multi-modal genomics data and imaging data to quantify heterogeneity in tumour tissues. Methods Mol Biol 2190:209–228
Troisi J, Mollo A, Lombardi M et al (2022) The metabolomic approach for the screening of endometrial cancer: validation from a large cohort of women scheduled for gynecological surgery. Biomolecules. https://doi.org/10.3390/biom12091229
Wang T, Lu W, Yang F et al (2020) Microsatellite instability prediction of uterine corpus endometrial carcinoma based on H&E histology whole-slide imaging. In: IEEE international symposium on biomedical imaging, ISBI 2020
Weiss JM, Gupta S, Burke CA et al (2021) NCCN Guidelines®Insights: genetic/familial high-risk assessment: colorectal, version 1.2021. J Natl Compr Cancer Netw 19:1122–1132
Yamashita R, Long J, Longacre T et al (2021) Deep learning model for the prediction of microsatellite instability in colorectal cancer: a diagnostic study. Lancet Oncol 22:132–141. https://doi.org/10.1016/S1470-2045(20)30535-0
Zhao X, Wu Y, Song G et al (2017) A deep learning model integrating FCNNs and CRFs for brain tumor segmentation. Med Image Anal 43:98–111
Zou S, Li C, Sun H et al (2022) TOD-CNN: an effective convolutional neural network for tiny object detection in sperm videos. Comput Biol Med 146:105543. https://doi.org/10.1016/j.compbiomed.2022.105543
Acknowledgements
The authors gratefully thank Doctor Li for her reviews and suggestions on pathology in this work. The authors also thank Xuzhou Medical University, The Affiliated Hospital of Xuzhou Medical University, and National Yang Ming Chiao Tung University for their support and help.
Funding
This work is funded by the General Program of the China Postdoctoral Science Foundation under Grant No. 2019M651974.
Author information
Authors and Affiliations
Contributions
YZ and SC made equal contributions to this work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Y., Chen, S., Wang, Y. et al. Deep learning-based methods for classification of microsatellite instability in endometrial cancer from HE-stained pathological images. J Cancer Res Clin Oncol 149, 8877–8888 (2023). https://doi.org/10.1007/s00432-023-04838-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04838-4