Abstract
Main conclusion
After bud burst, a transcriptional reprogramming of the shikimate and phenylpropanoid pathways occurs in grapevine canes resulting in the accumulation of stilbenoids like resveratrol and viniferin.
Abstract
Stilbenoids are phenylpropanoid compounds with important biological properties and biotechnological applications that are synthesized in grapevine in response to different stresses. Although they are found in woody tissues, such as canes and buds, their biosynthesis and accumulation have been essentially described in berries. We have previously shown that transcripts encoding secondary metabolism enzymes accumulate in grapevine canes following the transition from dormancy (E-L 1) to bud burst (E-L 4) suggesting that secondary metabolites may accumulate in grapevine canes during this transition. In the present study, using UPLC-MS we demonstrate the accumulation of important metabolites such as ferulic acid and the stilbenoids E-resveratrol, E-piceatannol and E-ε-viniferin. Stilbenoids accumulation correlated with the increased expression of several stilbene synthase genes and of VviMYB14, encoding a transcription factor that regulates stilbene biosynthesis. In addition, a general stimulation of the plastidial shikimate pathway was observed. Taken together, results show that important secondary metabolites accumulate in the woody canes during bud burst. These findings may aid biotechnological approaches aimed at extracting biologically active phenolic compounds, including stilbenoids, from grapevine woody tissues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The grapevine secondary metabolism has been extensively studied, with a focus on the phenylpropanoid pathway due to its importance in fruit and wine quality (Matus 2016). Phenylpropanoid biosynthesis starts from the aromatic amino acid phenylalanine, which is produced in plastids by the shikimate pathway. Flavonoids, stilbenoids, and phenolic acids are subsequently generated from intermediates of the phenylpropanoid pathway through the action of specific enzymes (Fig. 1; Vogt 2010), including chalcone and stilbene synthases which play a fundamental role in directing the carbon flux into flavonoids or stilbenoids synthesis depending on environmental, hormonal, and biotic cues (Dao et al. 2011). Stilbenoids, especially resveratrol, are among the most studied plant secondary metabolites due to their recognized human health benefits. They possess potent antioxidant, antibacterial, antifungal, cardioprotective, neuroprotective, antiaging, and anticancer properties (Biais et al. 2017).
Simplified representation of the shikimate and phenylpropanoid pathways and their connection via the aromatic amino acid phenylalanine. CM chorismate mutase, PAT prephenate aminotransferase, ADT arogenate dehydratase, PAL phenylalanine ammonia lyase, C4H cinnamate 4-hydroxylase, 4CL 4-coumarate-CoA ligase, C3H coumarate 3-hydroxylase, CHS chalcone synthase, SS stilbene synthase
The grapevine secondary metabolism has been mainly studied in grape berries and leaves, and E-resveratrol has been shown to accumulate in the exocarp, seeds and infected leaves (Gatto et al. 2008; Németh et al. 2017; Billet et al. 2020). However, E-resveratrol and other stilbenoids, such as E-piceid and E-ε-viniferin, are known to accumulate at much higher levels in perennial tissues, including canes and roots and in winter buds (Németh et al. 2017). Additionally, different stilbenoids like ampelopsin A, E-miyabenol C, Z/E-vitisin B, hopeaphenol, and isohopeaphenol are also accumulated in grapevine canes (Lambert et al. 2013; Houillé et al. 2015a, 2015b). Stilbenoids accumulation in canes is genotype dependent (Lambert et al. 2013; Billet et al. 2021), and in winter-harvested canes its accumulation is modulated by storage temperatures (Houillé et al. 2015b), while downy mildew infections stimulate stilbenoids synthesis during the growing season (Houillé et al. 2015a).
The induction and release of dormancy in perennial woody fruit crops, for which grapevine (Vitis vinifera L.) stands as a model, is finely tuned by complex environmental and endogenous signals. Although the physiological modifications that occur in the buds have been well-characterized (Zheng et al. 2015), the understanding of processes initiated in woody tissues upon dormancy release is still limited (Noronha et al. 2018). In a recent transcriptomic analysis, we have shown that genes involved in secondary metabolic pathways, including those of the phenylpropanoid pathway, were highly upregulated in woody tissues located below the bud after inducing bud burst (Noronha et al. 2021). Here, we demonstrate that the transcriptional activation of these genes results in the accumulation of important metabolites including ferulic acid and the stilbenoids E-resveratrol, E-piceatannol and E-ε-viniferin. We analyzed the expression of genes encoding enzymes of the shikimate and stilbene pathways. Overall, results are consistent with a general stimulation of the different pathways necessary for phenylpropanoid synthesis, which suggests that during bud burst carbon is being fueled to the phenylpropanoid pathway.
Material and methods
Plant material
Bud burst was induced following the protocol established by Noronha et al. (2021; Supplementary Fig. S1). Briefly, lignified grapevine canes from cv. Vinhão were collected from a commercial vineyard of the Controlled Appellation (DOC) region of Vinhos Verdes in the northwest region of Portugal (41°48′45.3″N 8°24′36.4″W) in 2018 and stored at 4 °C until the beginning of the experiment. Shoots of similar diameter were chosen and cut into single bud segments of approximately 15 cm and placed in the growth chamber (23 °C, 12/12 h photoperiod) in wet floral foam. When samples reached the E-L 4 stage, 3 cm segments just below the bud were cut, frozen in liquid N2, ground in an IKA A11 basic analytical mill, and stored at -80 ºC. Control samples were collected just before the beginning of the experiment (dormancy; E-L 1). At each sampling point, 3 or 4 independent cane segments were collected and pooled in 3 groups (biological replicates). The modified E-L system was used to classify the bud developmental stages (Pearce and Coombe 2004).
RNAseq analysis
Data treatment was previously performed on Noronha et al. (2021). Briefly, genes assigned as differentially expressed (P value < 0.05 and |log2FC|> 1.0) by DESeq (Anders and Huber 2010) were categorized according to the MapMan ontology (× 4.1) with Mercator tool (Lohse et al. 2014) and MapMan standalone software (v3.5.1) was used to explore the data.
Metabolomic analysis UPLC-MS
After lyophilization of ground samples, 50 mg were extracted in 1 mL ethanol/water solution (60:40; v/v), shaken for 30 min at 1400 rpm at 83 °C and centrifuged at 18 000 g for 5 min. The extracts were diluted 1:5 with 0.1% formic acid and stored at − 20 °C until analyses. UPLC-MS was performed using an ACQUITY™ Ultra Performance Liquid Chromatography system coupled to a photodiode array detector (PDA) and a Xevo TQD mass spectrometer (Waters, Milford, MA, USA) equipped with an electrospray ionization (ESI) source controlled by Masslynx 4.1 software (Waters). Analyte separation was achieved by using a Waters Acquity HSS T3 C18 column (150 mm × 2.1 mm, 1.8 µm) with a flow rate of 0.4 mL min−1 at 55 °C. The injection volume was 5 µL. The mobile phase consisted of solvent A (0.1% formic acid in water) and solvent B (0.1% formic acid in acetonitrile). Chromatographic separation was achieved using an 18-min linear gradient from 5 to 60% solvent B. MS detection was performed in both positive and negative modes. The capillary voltage was 3000 V and sample cone voltages were 30 and 60 V. The cone and desolvation gas flow rates were 60 and 800 L h−1. Identification of analytes was based on retention times, m/z values, and UV spectra as previously described (Billet et al. 2018a, b). Absolute quantifications were conducted using pure standards of tyrosine, tryptophan, phenylalanine, catechin, epicatechin, procyanidin B1, procyanidin B2, procyanidin B3, procyanidin B4, gallic acid, caffeic acid, ferulic acid, coutaric acid, caftaric acid, E-resveratrol, E-piceatannol, E-ε-viniferin, E-miyabenol C, ampelopsin A, Z/E-vitisin B and hopeaphenol using a five-point calibration curve (0–10 ppm). Z-ϵ-viniferin, ω-vinferin, δ-viniferin were expressed as E-ε-viniferin equivalent.
Analysis by HPLC–DAD
Methanolic extracts were prepared as indicated above. Samples were run at 0.8 mL/min, using 0.1% formic acid in water (eluent A) and 0.1% formic acid in methanol (eluent B) as mobile phase, through a Purospher Star RP-18 (25 × 0.4 mm, particle size of 5 µm, Merck) column connected to an HPLC (Hitachi—LaChrom Elite) equipped with a DAD (Diode Array Detector). The elution gradient start with 5% eluent B at 0 min to 95% B after 45 min. Detection was acquired in the range of 230–650 nm for obtention of compounds UV–Vis spectra. Principal Component Analysis Biplot (PCA) was performed in R Studio software version 4.1.0 using the FactoMineR package v1.34 (Lê et al. 2008).
Quantification of total phenolics and antioxidant activity in methanolic extracts
Phenolic compounds were extracted from 100 mg of lyophilized cane tissues by adding 1 mL of 80% methanol, mixing thoroughly, and an overnight incubation at 4 °C. Following this, the mixture was centrifuged, and the supernatants filtered. Total phenolics were quantified using the Folin-Ciocalteu method. Briefly, 10 µL of methanolic extracts were mixed with 100 µL of Folin reagent and 1.58 mL of H2O and, after 5 min incubation, 300 µL of 3 M sodium carbonate. After 2 h incubation in the dark, absorbance was measured at 765 nm and gallic acid was used as standard. Antioxidant capacity was assessed using the 2,2-diphenyl-1-picrylhydrazyl (DPPH) method. Briefly, 10 µL of methanolic extracts were incubated with 140 µL of 400 µM DPPH, the reaction incubated for 30 min, and the absorbance measured at 517 nm. Quercetin was used as standard.
Phenylalanine ammonia-lyase (PAL) activity
Phenylalanine ammonia-lyase (PAL) activity was determined using the protocol of Conde et al. (2016). Total protein extracts were obtained by mixing approximately 200 mg of from cane tissues with 1 mL of extraction buffer [50 mM Tris–HCl pH 8.9, 5 mM MgCl2, 1 mM EDTA, 1 mM phenylmethylsulfonyl fluoride, 5 mM dithiothreitol, and 0.1% (v/v) Triton X-100] followed by thorough mixing. The homogenates were centrifuged for 20 min at 18,000 g and the supernatants were collected and maintained on ice until the PAL enzymatic determination. Total protein concentrations of the extracts were determined by the Bradford method using bovine serum albumin as a standard (Bradford 1976). PAL enzymatic activity was determined in crude cane protein extracts by continuously monitoring the conversion of L-phenylalanine to trans-cinnamic acid at 290 nm. The 2 mL reaction mixture contained 0.2 mL of enzyme extract, 3.6 mM NaCl, and 25 mM L-phenylalanine (a saturating concentration) as substrate in 50 mM Tris–HCl, pH 8.9. Enzyme activity was calculated using the trans-cinnamic extinction coefficient of 16,890 M cm−1.
Real-time PCR studies
Total RNA from cane tissues was isolated as previously described by Reid et al. (2006) following some adaptations (Noronha et al. 2021). For each condition, 500 mg of frozen tissue was mixed with 1 mL of extraction buffer containing 300 mm Tris HCl (pH 8.0), 25 mm EDTA, 2.0 M NaCl, 2% CTAB, 2% PVP (K-30), and 30 mm DTT. Samples were then incubated at 60 °C for 15 min and shaken every 5 min. Then, mixtures were extracted twice with 850 μL of chloroform: isoamyl alcohol (24:1, v/v) followed by centrifugation at 15,000 g for 15 min at 4 °C. Subsequently, 0.1 vol of 3 M NaOAc (pH 5.2) and 0.6 vol of isopropanol were added to the aqueous phase, followed by incubation at − 80 °C for 1 h. The mixture was centrifuged at 15,000g at 4 °C for 30 min and resuspended in 100 μL of H2O. To purify the samples, the GRS Total RNA Kit—Plant (GriSP, Lda., Porto, Portugal) was used following the manufacturer’s instructions. RNA concentration and purity were quantified spectrophotometrically in a NanoDrop ND-1000 (Termo Fisher Scientifc Inc.) and integrity checked in a 1% (w/v) agarose gel. First-strand cDNA synthesis was performed using the Xpert cDNA Synthesis Mastermix protocol (GriSP, Lda.).
Quantitative real-time PCRs were performed with Xpert Fast SYBR Blue (GriSP, Lda.) along with the conditions previously optimized in a CBX96 Real-Time Detection System (Bio-Rad). The amplification protocol included an initial denaturation step at 95 °C for 3 min, followed by an additional 40 cycles of denaturation for 3 s at 95 °C and 30 s at 60 °C. Experiments were done in biological replicates and then interpreted with the Bio-Rad CFX Manager (Bio-Rad) software, while VviGAPDH (glyceraldehyde-3-phosphate dehydrogenase) and VviACT1 (actin) were used as internal control. The primers used in this study are listed in Supplementary Table S1.
Statistical analysis
Results were analyzed in the Prism vs. 6 (GraphPad Software, Inc.) using Analysis of Variances tests (oneway ANOVA). Statistical differences between samples were marked with asterisks (*P ≤ 0.05; ***P < 0.001; ****P < 0.0001).
Results
Transcriptomic analysis of phenylpropanoid and shikimate pathways in grapevine woody tissues following bud burst
The transcriptome profile was analyzed in woody tissues before (E-L 1) and after bud burst (E-L 4) using previously generated RNAseq data (Noronha et al. 2021). The visualization of the phenylpropanoid pathway using MapMan showed a general stimulation of differentially expressed transcripts at E-L 4 (Fig. 2). Particularly, genes encoding three PAL (bin 16.2.1.1), the first enzyme of the phenylpropanoid pathway, and those coding for 13 stilbene synthases and 3 chalcone synthases (bin 16.8.2.1) were upregulated by up to fivefold (log2 fold expression). In grapevine, 48 putative VviSTS gene sequences were identified, and at least 31 of them encode full-length proteins (Ciaffi et al. 2019). To our knowledge, it has been shown that VviSTS4 (named Vst1; Hain et al 1993), VviSTS15 and VviSTS22 (named vst1 and vst2; Thomzik et al. 1997) and VviSTS2, -5, -7, -10, -16, -20, -21, -25 and -29 (Parage et al. 2012) are able to synthesize stilbenes. Due to their high sequence similarity, transcript validation by qPCR was limited because genes are highly homologous, and most of them cannot be analyzed separately (Ciaffi et al. 2019). However, results confirmed that VviSTS3/4, -7/8, -14 and -20 were upregulated in woody tissues at E-L 4 (Supplementary Fig. S2). In addition, VviMYB14, which codes for a transcription factor involved in the regulation of stilbene synthesis (Höll et al. 2013), was upregulated threefold from E-L 1 to E-L 4, while the expression of VviMYB15 was not detected by qPCR (Supplementary Fig. S2).
MapMan pathway for phenylpropanoid pathway with DEGs. Each square represents a gene. In blue, genes more expressed at E-L 4 (bud burst); in red, genes more expressed at E-L 1 (dormancy). Bin numbers can be found in additional files with gene details. PAL phenylalanine ammonia lyase, C4H cinnamate 4-hydroxylase, 4CL 4-coumarate-CoA ligase, C3H coumarate 3-hydroxylase, COMT catechol-O-methyltransferase, 4CL 4-coumarate-CoA ligase, CHS chalcone synthase, SS stilbene synthase
RNA seq analysis also shows an increase in the transcript levels of key genes of the shikimate pathway when comparing E-L 1 to E-L 4 (Fig. 3). This is true for DAHPs (3-deoxy-d-arabinoheptulosonate 7-phosphate synthase) (except VviDAHPS3), which codes the first enzyme of the shikimate pathway, and for the ADT gene encoding arogenate dehydratases, which convert arogenate into phenylalanine. Also, the expression of several plastidial aromatic amino acid transporters was upregulated at E-L 4, namely VviCAT6 (Vitvi02g01754). (Fig. 3).
MapMan pathway for shikimate pathway with DEGs. Each square represents a gene. In blue, genes more expressed at E-L 4 (bud burst); in red, genes more expressed at E-L 1 (dormancy). When MapMan bin not available, grapevine VCost.v3 accession numbers are shown. Bin numbers can be found in additional files with gene details. DAHPS 3-deoxy-d-arabinoheptulosonate 7-phosphate synthase, DHQS 3-dehydroquinate synthase, SDH shikimate dehydrogenase, SK shikimate kinase, EPSPS 5-enolpyruvylshikimate-3-phosphate synthase, CM chorismate synthase, CM chorismate mutase, AS anthranilate synthase, PAT prephenate aminotransferase, ADT arogenate dehydratase, ADH arogenate dehydrogenase
Metabolomic analysis of grapevine canes by HPLC–DAD and UPLC-MS
The PCA of the HPLC–DAD analysis showed a clear separation between phenolic extracts from E-L 1 and E-L 4 (3-cm woody canes below the emerging bud), with PC1 accounting for 63% of the variation and PC2 for 16% (Supplementary Fig. S3). The targeted metabolomic analysis by UPLC-MS performed in the present study identified 25 metabolites, including phenolic acids, flavan-3-ols, and stilbenoids. Figure 4 shows the profile of variation of phenolic acids and flavan-3-ols from E-L 1 to E-L 4. While no changes were observed in the amounts of flavan-3-ols (Fig. 4a), some key phenolic acids changed following the bud burst. This is the case of ferulic acid that increased tenfold after bud burst (from 0.2 at E-L 1 to 2.2 µg g DW−1 at E-L 4; Fig. 4b). Conversely, coutaric and caftaric acids levels in woody tissues were reduced by up to 35% from E-L 1 to E-L 4.
Ten stilbenoids were identified and quantified by UPLC-MS in woody tissues at E-L 1 and E-L 4 (Fig. 5; chemical structures can be found in Supplementary Fig. S4). The content of monomeric stilbenoids like E-resveratrol significantly increased 6.3-fold from E-L 1 to E-L 4 (69.1 to 436.5 µg g DW−1) while E-piceatannol content increased 12.3-fold (9.2 to 113.18 µg g DW−1) in the same period (Fig. 5a). Dimeric stilbenoids were more abundant than monomeric ones at both developmental stages, but only δ-viniferin, which was not found at E-L 1, abruptly accumulated at bud burst up to 1.5 mg g DW−1. No changes were identified in the amounts of trimeric (Fig. 5c) and tetrameric stilbenoids following bud burst (Fig. 5d).
Despite the observed changes in the metabolomic profile of grapevine woody tissues following bud burst, the amount of total phenolics, measured by the Folin–Ciocalteau method, was not significantly modified. Nonetheless, the antioxidant capacity of the methanolic extracts increased by 48% from 14.0 at E-L 1 to 20.7 mg quercetin eq. g DW−1 at E-L 4 (Supplementary Fig. S5), most likely reflecting the above-referred qualitative changes.
Quantification of aromatic amino acids in grapevine canes
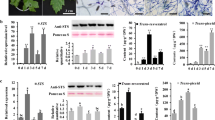
The content of aromatic amino acids in grapevine woody tissues below the bud determined by UPLC-MS showed that phenylalanine, which accumulates to 13.4 mg g DW−1, is by far more abundant (up to150-fold) than tyrosine or tryptophan (Fig. 6a). Following bud burst, while the contents in phenylalanine decreased by 40%, tyrosine levels increased by 450% (from 0.09 at E-L 1 to 0.41 mg g DW−1 at E-L 4) and tryptophan did not change. In parallel, in crude protein extracts of the woody canes, the biochemical activity of phenylalanine ammonia-lyase (PAL) increased by 60% (from 0.10 at E-L 1 to 0.16 µmol trans-cinnamic acid h−1 mg−1 protein at E-L 4) during the transition from dormancy to bud burst (Fig. 6b), which most likely accounted for the observed decrease in phenylalanine levels.
Discussion
Bud burst stimulates the shikimate and phenylpropanoid pathways in grapevine woody tissues
In this work, we explored the expression profiles of the phenylpropanoid and shikimate pathways and performed a metabolomic analysis using UPLC-MS in grapevine woody tissues at two developmental stages. Most genes of the shikimate pathway, which controls the synthesis of aromatic amino acids, are stimulated upon bud burst, particularly those encoding the ADT enzymes that catalyze the production of phenylalanine. Similarly, the expression level of genes involved in the transport of l-phenylalanine was also stimulated. Moreover, the increase of the enzymatic activity of PAL in woody canes protein extracts from E-L 1 to E-L 4 correlates with previous transcriptomic data showing the regulation of several VviPAL genes (Noronha et al. 2021). In agreement, both are consistent with stimulation of the phenylpropanoid pathway following bud burst, which most likely contributes to the decrease in the concentration of its precursor phenylalanine. A similar pattern with an increase in the expression of the PAL associated with a higher accumulation level of phenylpropanoids and a decrease in phenylalanine content was already observed in cold stored grape berries (Maoz et al. 2019). These findings strongly support the idea that, following bud burst, plastids located in grapevine woody tissues can de novo synthesize aromatic amino acids which are subsequently exported to the cytosol to fuel the phenylpropanoid pathway.
The stimulation of the phenylpropanoid pathway did not lead to a significant change in the total amount of phenolics but resulted in an enhanced antioxidant capacity of the woody tissue. This effect was associated with significant changes in the abundance of specific metabolites, including some phenolic acids and stilbenoids (discussed below). Of note, the observed increase in woody canes of ferulic acid at E-L 4, may be related to the increased production of cell wall components at this stage, including hemicelluloses or lignin, when secondary growth in grapevine woody tissues is resumed during spring (Noronha et al. 2021).
Stilbenoids are synthesized and accumulate following bud burst in grapevine woody tissues
In grapevine, most studies on stilbenoids are focused on berries, due to their relevance to the wine industry, or in leaves, due to their role as phytoalexins following pathogen infection (Billet et al. 2020), but they are also constitutively synthesized and accumulated in lignified grapevine organs (Gatto et al. 2008). In this regard, in the present study we demonstrate that at bud burst de novo synthesis and accumulation of the stilbenoids E-resveratrol, E-piceatannol, and δ-viniferin occurs in woody tissues close to the bud. Furthermore, the correlation between the accumulation of these compounds and the regulation of genes that control their synthesis suggests that stilbenoid accumulation is primarily controlled at the transcriptional level in woody tissues at bud burst. Consistently, stilbene synthase (STS) genes were upregulated/derepressed during the transition from E-L 1 to E-L 4. In addition, it has been described that VviMYB14 and VviMYB15 are fundamental in regulating stilbene accumulation in grapevine in response to several biotic and abiotic stresses (Höll et al. 2013) and that each of the transcription factors may activate the expression of different STS genes (Ciaffi et al. 2019). In the present study, the upregulation of 13 stilbene synthases genes (VviSTS3, -4, -7, -8, -13, -14, -15, -16, -17, -18, -19, -20; see Supplementary Table S2 for details) correlates with the increased expression of VviMYB14. Thus, we anticipate that VviMYB14 might be involved in the induction of the VviSTS genes leading to the accumulation of stilbenoids, especially considering that some of them (VviSTS4, -7, -15 and -16) have been shown to synthesize stilbenes (Hain et al. 1993; Thomsik et al. 1997; Parage et al. 2012). Interestingly, even though no increase in flavonoid content was detected by UPLC-MS, an upregulation of chalcone synthase expression, which competes with stilbene synthase for the same substrates, was also observed, suggesting that other undetected compounds may be synthesized in grapevine canes following bud burst.
Possible role for increased stilbenoids accumulation in grapevine woody tissues following bud burst
Taken together, the present results suggest that specific secondary compounds, including biologically relevant stilbenoids, are actively synthesized when young woody tissues may require additional protection against stress. Interestingly, besides the accumulation of monomeric stilbenoids (piceatannol and resveratrol) found at E-L 4, the levels of E-ε-viniferin also increased. The role of dimeric stilbenoids in grapevine has been shown previously, and viniferin production is considered a good indicator of plant resistance against fungal attacks (Pezet et al. 2004). Also, grapevine cv. Vinhão cultured cells accumulate stilbenoids (including viniferin-type compounds) following their inoculation with Phaeomoniella chlamydospore autoclaved extracts, a pathogen associated with esca disease (Lima et al. 2012). Thus, it is likely that the stilbenoids accumulated in grapevine woody tissues promote protection against fungal pathogens following bud burst.
Despite the vast body of reports on stilbenoids, their true subcellular localization remains puzzling. In grapevine, several reports have shown that the STS enzyme is localized in the cell wall (Fornara et al. 2008) together with its product stilbenoids (Bellow et al. 2012). The cell wall is a barrier that fungal pathogens must penetrate to access intracellular components and lignin deposition, as well as other phenolics, is recognized as a defense mechanism against these attacks (Ninkuu et al. 2023). Thus, the synthesis and accumulation of stilbenoids in the cell wall is consistent with its antimicrobial and protective properties and may be an effective way to protect the plant from undesired toxic effects (Fornara et al. 2008). Indeed, overexpressing STS genes leads to an increased resistance against pathogens in Arabidopsis (Huang et al. 2016), tobacco (He et al. 2018) and grapevine 41B rootstock (Coutos-Thévenot et al. 2001). Strikingly, it was also shown that the overexpression of an STS from Vitis quinquangularis in Arabidopsis thaliana produces plants that are more tolerant of osmotic stress conditions, possibly via the stimulation of the plant antioxidant machinery (Huang et al. 2016). Thus, it is possible that stilbenoids accumulated in cell walls could help mitigating oxidative stress that occurs during secondary growth.
Besides the scientific dimension of the present study, results also hold biotechnological interest since grapevine woody tissues are currently being used as raw material for the extraction of biologically active phenolic compounds and to obtain extracts with antimicrobial activities (Billet et al. 2018a, b; El Khawand et al. 2020).
Data availability
The data that support the findings of this study are available in the European Nucleotide Archive with the ENA study accession PRJEB43358.
Abbreviations
- PAL:
-
Phenylalanine ammonia lyase
- STS:
-
Stilbene synthase
References
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:R106. https://doi.org/10.1186/gb-2010-11-10-r106
Bellow S, Latouche G, Brown SC, Poutaraud A, Cerovic ZG (2012) In vivo localization at the cellular level of stilbene fluorescence induced by Plasmopara viticola in grapevine leaves. J Exp Bot 63(10):3697–3707. https://doi.org/10.1093/jxb/ers060
Biais B, Krisa S, Cluzet S, Da Costa G, Waffo-Teguo P, Mérillon JM, Richard T (2017) Antioxidant and cytoprotective activities of grapevine stilbenes. J Agric Food Chem 65(24):4952–4960. https://doi.org/10.1021/acs.jafc.7b01254
Billet K, Delanoue G, Arnault I, Besseau S, Oudin A, Courdavault V, Marchand PA, Giglioli-Guivarc’h N, Guérin L, Lanoue A (2018a) Vineyard evaluation of stilbenoid-rich grape cane extracts against downy mildew: a large-scale study. Pest Manag Sci 75:1252–1257. https://doi.org/10.1002/ps.5237
Billet K, Houillé B, de Bernonville TD, Besseau S, Oudin A, Courdavault V, Delanoue G, Guérin L, Clastre M, Giglioli-Guivarc’h N, Lanoue A (2018b) Field-based metabolomics of Vitis vinifera L. stems provides new insights for genotype discrimination and polyphenol metabolism structuring. Front Plant Sci 9:798. https://doi.org/10.3389/fpls.2018.00798
Billet K, Malinowska MA, Munsch T, Unlubayir M, Adler S, Delanoue G, Lanoue A (2020) Semi-targeted metabolomics to validate biomarkers of grape downy mildew infection under field conditions. Plants 9:1008. https://doi.org/10.3390/plants9081008
Billet K, Unlubayir M, Munsch T, Malinowska MA, de Bernonville TD, Oudin A, Courdavault V, Besseau S, Giglioli-Guivarc’h N, Lanoue A (2021) Postharvest treatment of wood biomass from a large collection of European grape varieties: impact on the selection of polyphenol-rich byproducts. ACS Sustain Chem Eng 9(9):3509–3517. https://doi.org/10.1021/acssuschemeng.0c07875
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 7(72):248–254. https://doi.org/10.1006/abio.1976.9999
Ciaffi M, Paolacci AR, Paolocci M, Alicandri E, Bigini V, Badiani M, Muganu M (2019) Transcriptional regulation of stilbene synthases in grapevine germplasm differentially susceptible to downy mildew. BMC Plant Biol 19(1):404. https://doi.org/10.1186/s12870-019-2014-5
Conde A, Pimentel D, Neves A, Dinis LT, Bernardo S, Correia CM, Gerós H, Moutinho-Pereira J (2016) Kaolin foliar application has a stimulatory effect on phenylpropanoid and flavonoid pathways in grape berries. Front Plant Sci 7:1150. https://doi.org/10.3389/fpls.2016.01150
Coutos-Thévenot P, Poinssot B, Bonomelli A, Yean H, Breda C, Buffard D, Esnault R, Hain R, Boulay M (2001) In vitro tolerance to Botrytis cinerea of grapevine 41B rootstock in transgenic plants expressing the stilbene synthase Vst1 gene under the control of a pathogen-inducible PR 10 promoter. J Exp Bot 52(358):901–910. https://doi.org/10.1093/jexbot/52.358.901
Dao TT, Linthorst HJ, Verpoorte R (2011) Chalcone synthase and its functions in plant resistance. Phytochem Rev 10(3):397–412. https://doi.org/10.1007/s11101-011-9211-7
El Khawand T, Gabaston J, Taillis D, Iglesias ML, Pedrot E, Palos Pinto A, Fonayet JV, Merillon JM, Decendit A, Cluzet S, Richard T (2020) A dimeric stilbene extract produced by oxidative coupling of resveratrol active against plasmopara viticola and botrytis cinerea for vine treatments. OENO One 54(1):157–164. https://doi.org/10.20870/oeno-one.2020.54.1.2529
Fornara V, Onelli E, Sparvoli F, Rossoni M, Aina R, Marino G, Citterio S (2008) Localization of stilbene synthase in Vitis vinifera L. during berry development. Protoplasma 233:83–93. https://doi.org/10.1007/s00709-008-0309-8
Gatto P, Vrhovsek U, Muth J, Segala C, Romualdi C, Fontana P, Pruefer D, Stefanini M, Moser C, Mattivi F, Velasco R (2008) Ripening and genotype control stilbene accumulation in healthy grapes. J Agric Food Chem 56(24):11773–11785. https://doi.org/10.1021/jf8017707
Hain R, Reif HJ, Krause E, Langebartels R, Kindl H, Vornam B, Wiese W, Schmelzer E, Schreier PH, Stöcker RH, Stenzel K (1993) Disease resistance results from foreign phytoalexin expression in a novel plant. Nature 361(6408):153–156. https://doi.org/10.1038/361153a0
He X, Xue F, Zhang L, Guo H, Ma L, Yang M (2018) Overexpressing fusion proteins of 4-coumaroyl-CoA ligase (4CL) and stilbene synthase (STS) in tobacco plants leading to resveratrol accumulation and improved stress tolerance. Plant Biotechnol Rep 12:295–302. https://doi.org/10.1007/s11816-018-0494-7
Höll J, Vannozzi A, Czemmel S, D’Onofrio C, Walker AR, Rausch T, Lucchin M, Boss PK, Dry IB, Bogs J (2013) The R2R3-MYB transcription factors MYB14 and MYB15 regulate stilbene biosynthesis in Vitis vinifera. Plant Cell 25(10):4135–4149. https://doi.org/10.1105/tpc.113.117127
Houillé B, Besseau S, Courdavault V, Oudin A, Glévarec G, Delanoue G, Guérin L, Simkin AJ, Papon N, Clastre M, Giglioli-Guivarc’h N, Lanoue A (2015a) Biosynthetic origin of E-resveratrol accumulation in grape canes during postharvest storage. J Agric Food Chem 63(5):1631–1638. https://doi.org/10.1021/jf505316a
Houillé B, Besseau S, Delanoue G, Oudin A, Papon N, Clastre M, Simkin AJ, Guérin L, Courdavault V, Giglioli-Guivarc’h N, Lanoue A (2015b) Composition and tissue-specific distribution of stilbenoids in grape canes are affected by downy mildew pressure in the vineyard. J Agric Food Chem 63(38):8472–8477. https://doi.org/10.1021/acs.jafc.5b02997
Huang L, Zhang S, Singer SD, Yin X, Yang J, Wang Y, Wang X (2016) Expression of the grape VqSTS21 gene in Arabidopsis confers resistance to osmotic stress and biotrophic pathogens but not Botrytis cinerea. Front Plant Sci 7:1379. https://doi.org/10.3389/fpls.2016.01379
Lambert C, Richard T, Renouf E, Bisson J, Waffo-Téguo BL, Ollat N, Mérillon JM, Cluzet S (2013) Comparative analyses of stilbenoids in canes of major Vitis vinifera L. cultivars. J Agric Food Chem 61:11392–11399. https://doi.org/10.1021/jf403716y
Lê S, Josse J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Soft 25(1):1–18
Lima MRM, Ferreres F, Dias ACP (2012) Response of Vitis vinifera cell cultures to Phaeomoniella chlamydospora: changes in phenolic production, oxidative state and expression of defence-related genes. Eur J Plant Pathol 132:133–146. https://doi.org/10.1007/s10658-011-9857-4
Lohse M, Nagel A, Herter T, May P, Schroda M, Zrenner R, Tohge T, Fernie AR, Stitt M, Usadel B (2014) Mercator: a fast and simple web server for genome scale functional annotation of plant sequence data. Plant Cell Environ 37:1250–1258. https://doi.org/10.1111/pce.12231
Maoz I, De Rosso M, Kaplunov T, Vedova AD, Sela N, Flamini R, Lewinsohn E, Lichter A (2019) Metabolomic and transcriptomic changes underlying cold and anaerobic stresses after storage of table grapes. Sci Rep 9(1):2917. https://doi.org/10.1038/s41598-019-39253-8
Matus JT (2016) Transcriptomic and metabolomic networks in the grape berry illustrate that it takes more than flavonoids to fight against ultraviolet radiation. Front Plant Sci 7:1337. https://doi.org/10.3389/fpls.2016.01337
Németh G, Hegyi O, Dunai A, Kocsis L (2017) Stilbenes in the different organs of Vitis vinifera cv. Merlot grafted on TK5BB rootstock. OENO One 51(3):323–328. https://doi.org/10.20870/oeno-one.2016.50.4.1068
Ninkuu V, Yan J, Fu Z, Yang T, Ziemah J, Ullrich MS, Kuhnert N, Zeng H (2023) Lignin and its pathway-associated phytoalexins modulate plant defense against fungi. J Fungi 9:52. https://doi.org/10.3390/jof9010052
Noronha H, Silva A, Dai Z, Gallusci P, Rombolà AD, Gerós H (2018) A molecular perspective on starch metabolism in woody tissues. Planta 248:559–568. https://doi.org/10.1007/s00425-018-2954-2
Noronha H, Garcia V, Silva A, Delrot S, Gallusci P, Gerós H (2021) Molecular reprogramming in grapevine woody tissues at bud burst. Plant Sci 311:110984. https://doi.org/10.1016/j.plantsci.2021.110984
Parage C, Tavares R, Réty S, Baltenweck-Guyot R, Poutaraud A, Renault L, Heintz D, Lugan R, Marais GA, Aubourg S, Hugueney P (2012) Structural, functional, and evolutionary analysis of the unusually large stilbene synthase gene family in grapevine. Plant Physiol 160(3):1407–1419. https://doi.org/10.1104/pp.112.202705
Pearce I, Coombe BG (2004) Grapevine phenology. In: Dry P, Coombe BG (eds) Viticulture 1—resources, 2nd edn. Winetitles, Adelaide, pp 150–166
Pezet R, Gindro K, Viret O, Spring JL (2004) Glycosylation and oxidative dimerization of resveratrol are respectively associated to sensitivity and resistance of grapevine cultivars to downy mildew. Physiol Mol Plant Pathol 65:297–303. https://doi.org/10.1016/j.pmpp.2005.03.002
Reid KE, Olsson N, Schlosser J, Peng F, Lund ST (2006) An optimized grapevine RNA isolation procedure and statistical determination of reference genes for real-time RT-PCR during berry development. BMC Plant Biol 6:27. https://doi.org/10.1186/1471-2229-6-27
Thomzik JE, Stenzel K, Stöcker R, Schreier PH, Hain R, Stahl DJ (1997) Synthesis of a grapevine phytoalexin in transgenic tomatoes (Lycopersicon esculentum Mill.) conditions resistance against Phytophthora infestans. Physiol Mol Plant Pathol 51:265–278. https://doi.org/10.1006/pmpp.1997.0123
Vogt T (2010) Phenylpropanoid biosynthesis. Mol Plant 3(1):2–20. https://doi.org/10.1093/mp/ssp106
Zheng C, Halaly T, Acheampong AK, Takebayashi Y, Jikumaru Y, Kamiya Y, Or E (2015) Abscisic acid (ABA) regulates grape bud dormancy, and dormancy release stimuli may act through modification of ABA metabolism. J Exp Bot 66(5):1527–1542. https://doi.org/10.1093/jxb/eru519
Funding
Open access funding provided by FCT|FCCN (b-on). This work was supported by Fundação para a Ciência e Tecnologia (FCT) under the strategic program UIDB/BIA/04050/2020. The work was also supported by FCT, CCDR-N (Norte Portugal Regional Coordination and Development Commission) and European Funds (FEDER/POCI/COMPETE2020) through the project AgriFood XXI (NORTE-01–0145-FEDER-000041) and the research project GrapeMicrobiota (PTDC/BAA-AGR/2691/2020). AS and HN were supported by FCT grants SFRH/BD/135782/2018 and SFRH/BPD/115518/2016, respectively. This work benefited from the networking activities within the CoLAB Vines & Wines.
Author information
Authors and Affiliations
Contributions
HN, PG, AL and HG conceptualized the work. HN, AS, VG, AD and KB conducted the experiments. HN, AL, AD, PG and HG contributed to the analysis of the results. HN, AS, VG, KB, AD, AL, PG and HG wrote and reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to disclose.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Noronha, H., Silva, A., Garcia, V. et al. Grapevine woody tissues accumulate stilbenoids following bud burst. Planta 258, 118 (2023). https://doi.org/10.1007/s00425-023-04270-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-023-04270-5