Abstract
Purpose
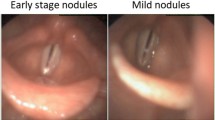
To develop and validate a deep learning model for distinguishing healthy vocal folds (HVF) and vocal fold polyps (VFP) on laryngoscopy videos, while demonstrating the ability of a previously developed informative frame classifier in facilitating deep learning development.
Methods
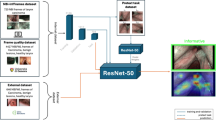
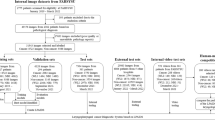
Following retrospective extraction of image frames from 52 HVF and 77 unilateral VFP videos, two researchers manually labeled each frame as informative or uninformative. A previously developed informative frame classifier was used to extract informative frames from the same video set. Both sets of videos were independently divided into training (60%), validation (20%), and test (20%) by patient. Machine-labeled frames were independently verified by two researchers to assess the precision of the informative frame classifier. Two models, pre-trained on ResNet18, were trained to classify frames as containing HVF or VFP. The accuracy of the polyp classifier trained on machine-labeled frames was compared to that of the classifier trained on human-labeled frames. The performance was measured by accuracy and area under the receiver operating characteristic curve (AUROC).
Results
When evaluated on a hold-out test set, the polyp classifier trained on machine-labeled frames achieved an accuracy of 85% and AUROC of 0.84, whereas the classifier trained on human-labeled frames achieved an accuracy of 69% and AUROC of 0.66.
Conclusion
An accurate deep learning classifier for vocal fold polyp identification was developed and validated with the assistance of a peer-reviewed informative frame classifier for dataset assembly. The classifier trained on machine-labeled frames demonstrates improved performance compared to the classifier trained on human-labeled frames.
Level of evidence
4.





Similar content being viewed by others
References
Esteva A, Robicquet A, Ramsundar B et al (2019) A guide to deep learning in healthcare. Nat Med 25(1):24–29. https://doi.org/10.1038/s41591-018-0316-z
Wang P, **ao X, Glissen Brown JR et al (2018) Development and validation of a deep-learning algorithm for the detection of polyps during colonoscopy. Nat Biomed Eng 2(10):741–748. https://doi.org/10.1038/s41551-018-0301-3
Lee JY, Jeong J, Song EM et al (2020) Real-time detection of colon polyps during colonoscopy using deep learning: systematic validation with four independent datasets. Sci Rep 10(1):8379. https://doi.org/10.1038/s41598-020-65387-1
Urban G, Tripathi P, Alkayali T et al (2018) Deep learning localizes and identifies polyps in real time with 96% accuracy in screening colonoscopy. Gastroenterology 155(4):1069-1078.e8. https://doi.org/10.1053/j.gastro.2018.06.037
Ren J, **g X, Wang J et al (2020) Automatic recognition of laryngoscopic images using a deep-learning technique. Laryngoscope 130(11):E686–E693. https://doi.org/10.1002/lary.28539
**ong H, Lin P, Yu JG et al (2019) Computer-aided diagnosis of laryngeal cancer via deep learning based on laryngoscopic images. EBioMedicine 48:92–99. https://doi.org/10.1016/j.ebiom.2019.08.075
Yao P, Witte D, Gimonet H, German A, Andreadis K, Cheng M, Sulica L, Elemento O, Barnes J, Rameau A (2022) Automatic classification of informative laryngoscopic images using deep learning. Laryngoscope Investig Otolaryngol 7(2):460–466. https://doi.org/10.1002/lio2.754
Rosen CA, Gartner-Schmidt J, Hathaway B et al (2012) A nomenclature paradigm for benign midmembranous vocal fold lesions. Laryngoscope 122(6):1335–1341. https://doi.org/10.1002/lary.22421
Dunham ME, Kong KA, McWhorter AJ, Adkins LK (2022) Optical biopsy: automated classification of airway endoscopic findings using a convolutional neural network. Laryngoscope 132(Suppl 4):S1–S8. https://doi.org/10.1002/lary.28708
He K, Zhang X, Ren S, Sun J (2021) Deep Residual Learning for Image Recognition. Ar**v151203385 Cs. Published online December 10, 2015. http://arxiv.org/abs/1512.03385. Accessed January 22, 2021
Kingma DP, Ba J (2021) Adam: A Method for Stochastic Optimization. Ar**v14126980 Cs. Published online January 29, 2017. http://arxiv.org/abs/1412.6980. Accessed January 22, 2021
Pandey R, Purohit H, Castillo C, Shalin VL (2022) Modeling and mitigating human annotation errors to design efficient stream processing systems with human-in-the-loop machine learning. Int J Human-Comput Stud. 160:102772. https://doi.org/10.1016/j.ijhcs.2022.102772
Burghardt K, Hogg T, Lerman K (2018) Quantifying the impact of cognitive biases in question-answering systems. In: Proceedings of the International AAAI Conference on Web and Social Media 12(1)
Zhang L, Tanno R, Xu MC, ** C, Jacob J, Cicarrelli O, Barkhof F, Alexander D (2020) Disentangling human error from ground truth in segmentation of medical images. Adv Neural Inf Process Syst 33:15750–15762
Cheplygina V, de Bruijne M, Pluim JPW (2019) Not-so-supervised: A survey of semi-supervised, multi-instance, and transfer learning in medical image analysis. Med Image Anal 54:280–296. https://doi.org/10.1016/j.media.2019.03.009
Zhang L, Wu L, Wei L, Wu H, Lin Y (2023) A novel framework of manifold learning cascade-clustering for the informative frame selection. Diagnostics (Basel) 13(6):1151. https://doi.org/10.3390/diagnostics13061151
Kuo CFJ, Lai WS, Barman J, Liu SC (2021) Quantitative laryngoscopy with computer-aided diagnostic system for laryngeal lesions. Sci Rep 11(1):10147. https://doi.org/10.1038/s41598-021-89680-9
Funding
This project was supported by the American Laryngological Voice and Research Education Grant. Anaïs Rameau was supported by a Paul B. Beeson Emerging Leaders Career Development Award in Aging (K76 AG079040) from the National Institute on Aging and by the Bridge2AI award (OT2 OD032720) from the NIH Common Fund. Anaïs Rameau is a medical advisor for Perceptron Health, Inc. Dan Witte is a co-founder of Perceptron Health, Inc.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All conflicts of interest are disclosed in the Title Page.
Ethical approval
Weill Cornell Medical College IRB approval was obtained for this study, Protocol # 19-05020151. Ethical Standards were upheld by all authors.
Informed consent
Informed consent was not required by the IRB due to the retrospective nature of the clinical data used in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yao, P., Witte, D., German, A. et al. A deep learning pipeline for automated classification of vocal fold polyps in flexible laryngoscopy. Eur Arch Otorhinolaryngol 281, 2055–2062 (2024). https://doi.org/10.1007/s00405-023-08190-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00405-023-08190-8