Abstract
Objectives
Deep learning is in this study used through convolutional neural networks (CNN) to the determination of vocal fold nodules. Through high-speed video (HSV) images and computer-assisted tools, a comparison of convolutional neural network models and their accuracy will be presented.
Methods
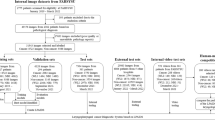
The data have been collected by an Ear Nose Throat (ENT) specialist with a 90° rigid scope in the years from 2007 to 2019, where 15.732 high-speed videos have been collected from 7909 patients. A total of 4000 images have been carefully selected, 2000 images were of normal vocal folds and 2000 images were of vocal folds with varying degrees of vocal fold nodules. These images were then split into training-, validation-, and testing-data set, for use with a CNN model with 5 layers (CNN5) and compared to other models: VGG19, MobileNetV2, and Inception-ResNetV2. To compare the neural network models, the following evaluation metrics have been calculated: accuracy, sensitivity, specificity, precision, and negative predictive values.
Results
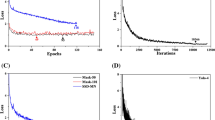
All the trained CNN models have shown high accuracy when applied to the test set. The accuracy is 97.75%, 83.5%, 91.5%, and 89.75%, for CNN5, VGG19, MobileNetV2, and InceptionResNetV2, respectively.
Conclusions
Precision was identified as the most relevant performance metric for a study that focuses on the classification of vocal fold nodules. The highest performing model was MobilNetV2 with a precision of 97.7%. The average accuracy across all 4 neural networks was 90.63% showing that neural networks can be used for classifying vocal fold nodules in a clinical setting.



Similar content being viewed by others
References
Nagata K, Kurita S, Yasumoto S, Maeda T, Kawasaki H, Hirano M (1983) Vocal fold polyps and nodules. A 10-year review of 1,156 patients. Auris Nasus Larynx 10(Suppl):S27-35. https://doi.org/10.1016/s0385-8146(83)80003-0
Pedersen M, McGlashan J (2012) Surgical versus non-surgical interventions for vocal cord nodules. Cochrane Database Syst Rev. https://doi.org/10.1002/14651858.CD001934.pub2
Pedersen M, Jønsson AO, Akbulut S, Oguz H, Nawka T (2020) Benign organic voice disorders. In: am Zehnhoff-Dinnesen A, Wiskirska-Woznica B, Neumann K, Nawka T (eds) Phoniatrics 1, 1st edn. Springer, Berlin, pp 257–263
Oates J, Dacakis G (1997) Voice change in transsexuals. Venereology 10:178
Yao P, Usman M, Chen YH, German A, Andreadis K, Mages K, Rameau A (2021) Applications of artificial intelligence to office laryngoscopy: a sco** review. Laryngoscope. https://doi.org/10.1002/lary.29886
Kist AM, Gómez P, Dubrovskiy D, Schlegel P, Kunduk M, Echternach M, Patel R, Semmler M, Bohr C, Dürr S, Schützenberger A, Döllinger M (2021) A deep learning enhanced novel software tool for laryngeal dynamics analysis. J Speech Lang Hear Res 64(6):1889–1903. https://doi.org/10.1044/2021_JSLHR-20-00498
Unger J, Lohscheller J, Reiter M, Eder K, Betz CS, Schuster M (2015) A noninvasive procedure for early-stage discrimination of malignant and precancerous vocal fold lesions based on laryngeal dynamics analysis. Cancer Res 75(1):31–39. https://doi.org/10.1158/0008-5472.CAN-14-1458
Azam MA, Sampieri C, Ioppi A, Africano S, Vallin A, Mocellin D, Fragale M, Guastini L, Moccia S, Piazza C, Mattos LS, Peretti G (2021) Deep learning applied to white light and narrow band imaging videolaryngoscopy: toward real-time laryngeal cancer detection. Laryngoscope. https://doi.org/10.1002/lary.29960
Parker F, Brodsky MB, Akst LM, Ali H (2021) Machine learning in laryngoscopy analysis: a proof-of-concept observational study for the identification of post-extubation ulcerations and granulomas. Ann Otol Rhinol Laryngol 130(3):286–291. https://doi.org/10.1177/0003489420950364
Ren J, **g X, Wang J, Ren X, Xu Y, Yang Q, Ma L, Sun Y, Xu W, Yang N, Zou J, Zheng Y, Chen M, Gan W, **ang T, An J, Liu R, Lv C, Lin K, Zheng X, Lou F, Rao Y, Yang H, Liu K, Liu G, Lu T, Zheng X, Zhao Y (2020) Automatic recognition of laryngoscopic images using a deep-learning technique. Laryngoscope 130(11):E686–E693. https://doi.org/10.1002/lary.28539
Cho WK, Lee YJ, Joo HA, Jeong IS, Choi Y, Nam SY, Kim SY, Choi SH (2021) Diagnostic accuracies of laryngeal diseases using a convolutional neural network-based image classification system. Laryngoscope 131(11):2558–2566. https://doi.org/10.1002/lary.29595
Crowson MG, Ranisau J, Eskander A, Babier A, Xu B, Kahmke RR, Chen JM, Chan TCY (2020) A contemporary review of machine learning in otolaryngology-head and neck surgery. Laryngoscope 130(1):45–51. https://doi.org/10.1002/lary.27850
Keras.io, Keras Applications, https://keras.io/api/applications/. Accessed 12 May 2022
Wikipedia.org, The Company – ImageNet, https://en.wikipedia.org/wiki/ImageNet. Accessed 10 May 2022
Müller AC, Guido S (2017) Introduction to machine learning with Python: a guide for data scientists. O’reilly, Bei**g
Geekymedics.com, Sensitivity, specificity, PPV and NPV, https://geekymedics.com/sensitivity-specificity-ppv-and-npv/. Accessed 14 May 2022
Pedersen M, Larsen CF (2021) Accuracy of laryngoscopy for quantitative vocal fold analysis in combination with AI, A cohort study of manual artefacts. Sch J Otolaryngol. https://doi.org/10.32474/SJO.2021.06.000237
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Larsen, C.F., Pedersen, M. Comparison of convolutional neural networks for classification of vocal fold nodules from high-speed video images. Eur Arch Otorhinolaryngol 280, 2365–2371 (2023). https://doi.org/10.1007/s00405-022-07736-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00405-022-07736-6