Abstract
Key message
OsiCRY2 is involved in light-regulated plant development and plays a role in regulating photomorphogenesis, plant height, flowering and most strikingly partial photomorphogenesis in dark.
Abstract
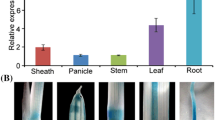
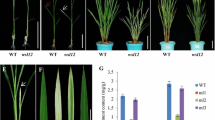
Cryptochrome 2 (CRY2), the blue/UV-A light photoreceptor in plants, has been reported to regulate photoperiod-dependent flowering and seedling photomorphogenesis (under low-intensity light). Among monocots, CRY2 has been reported from japonica rice, wheat, sorghum and barley. The two sub-species of rice, indica and japonica, exhibit a high degree of genetic variation and morphological and physiological differences. This article describes the characterization of CRY2 of indica rice (OsiCRY2). While the transcript levels of OsiCRY2 did not change significantly under blue light, its protein levels were found to decline with increased time duration under blue light. For phenotypic characterization, OsiCRY2 over-expression (OX) transgenics were generated in Oryza sativa Pusa Sugandh 2 (PS2) cultivar, a highly scented Basmati cultivar. The OsiCRY2OX transgenics displayed shorter coleoptiles and dwarfism than wild-type under blue light, white, and far-red light. Interestingly, even the dark-grown transgenics were shorter, concomitant with higher OsiCRY2 protein levels in transgenics than wild-type. Histological analysis revealed that the decrease in the length of the seedlings was due to a decrease in the length of the epidermal cells. The fully mature rice transgenics were shorter than the untransformed plants but flowered at the same time as wild-type. However, the OsiCRY2 Arabidopsis over-expressors exhibited early flowering by 10–15 days, indicating the potential and conservation of function of OsiCRY2. The whole-genome transcriptome profiling of rice transgenics revealed the differential up-regulation of several light-regulated genes in dark-grown coleoptiles. These data provide evidence that OsiCRY2 regulates photomorphogenesis, plant height, and flowering in indica rice.









Similar content being viewed by others
Data availability
The microarray data have been uploaded in Gene Expression Omnibus (GEO) database and its accession number is GSE200963.
References
Ahmad M, Cashmore AR (1993) HY4 gene of A. thaliana encodes a protein with characteristics of a blue-light photoreceptor. Nature 366:162–166. https://doi.org/10.1038/366162a0
Ahmad M, Lin C, Cashmore AR (1995) Mutations throughout an Arabidopsis blue-light photoreceptor impair blue-light-responsive anthocyanin accumulation and inhibition of hypocotyl elongation. Plant J 8:653–658
Ahmad M, Jarillo JA, Cashmore AR (1998) Chimeric proteins between cry1 and cry2 Arabidopsis blue light photoreceptors indicate overlap** functions and varying protein stability. Plant Cell 10:197–207. https://doi.org/10.1105/tpc.10.2.197
Barrero JM, Downie AB, Xu Q, Gubler F (2014) A role for barley CRYPTOCHROME1 in light regulation of grain dormancy and germination. Plant Cell 26:1094–1104. https://doi.org/10.1105/tpc.113.121830
Chatterjee M, Sharma P, Khurana JP (2006) Cryptochrome 1 from Brassica napus is up-regulated by blue light and controls hypocotyl/stem growth and anthocyanin accumulation. Plant Physiol 141:61–74. https://doi.org/10.1104/pp.105.076323
Chaves I, Pokorny R, Byrdin M, Hoang N, Ritz T, Brettel K, Essen LO, Van Der Horst GTJ, Batschauer A, Ahmad M (2011) The cryptochromes: blue light photoreceptors in plants and animals. Annu Rev Plant Biol 62:335–364. https://doi.org/10.1146/annurev-arplant-042110-103759
Chory J, Nagpal P, Peto CA (1991) Phenotypic and genetic analysis of det2, a new mutant that affects light-regulated seedling development in Arabidopsis. Plant Cell 3:445–459. https://doi.org/10.2307/3869351
Clough SJ, Bent AF (1998) Floral dip: A simplified method for agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313X.1998.00343.x
Deng XW, Caspar T, Quail PH (1991) cop1: a regulatory locus involved in light-controlled development and gene expression in Arabidopsis. Genes Dev 5:1172–1182. https://doi.org/10.1101/gad.5.7.1172
El-Din El-Assal S, Alonso-Blanco C, Peeters AJM, Wagemaker C, Weller JL, Koornneef M (2003) The role of cryptochrome 2 in flowering in Arabidopsis. Plant Physiol 133:1504–1516. https://doi.org/10.1104/pp.103.029819
Giliberto L, Perrotta G, Pallara P, Weller JL, Fraser PD, Bramley PM, Fiore A, Tavazza M, Giuliano G (2005) Manipulation of the blue light photoreceptor cryptochrome 2 in tomato affects vegetative development, flowering time, and fruit antioxidant content. Plant Physiol 137:199–208. https://doi.org/10.1104/pp.104.051987
Guo H, Yang H, Mockler TC, Lin C (1998) Regulation of flowering time by Arabidopsis photoreceptors. Science. https://doi.org/10.1242/dev.02340
Guo H, Duong H, Ma N, Lin C (1999) The Arabidopsis blue light receptor cryptochrome 2 is a nuclear protein regulated by a blue light-dependent post-transcriptional mechanism. Plant J 19:279–287. https://doi.org/10.1046/j.1365-313X.1999.00525.x
Guo J, Li W, Shang L, Wang Y, Yan P, Bai Y, Da X, Wang K, Guo Q, Jiang R, Mao C, Mo X (2021) OsbHLH98 regulates leaf angle in rice through transcriptional repression of OsBUL1. New Phytol 230:1953–1966. https://doi.org/10.1111/nph.17303
Hirose F, Shinomura T, Tanabata T, Shimada H, Takano M (2006) Involvement of rice cryptochromes in De-etiolation responses and flowering. Plant Cell Physiol 47:915–925. https://doi.org/10.1105/tpc.113.121830
Jang S, An G, Li HY (2017) Rice leaf angle and grain size are affected by the OsBUL1 transcriptional activator complex. Plant Physiol 173:688–702. https://doi.org/10.1104/pp.16.01653
Jiao Y, Lau OS, Deng XW (2007) Light-regulated transcriptional networks in higher plants. Nat Rev Genet 8(3):217–230. https://doi.org/10.1038/nrg2049
Koornneef M, Rolff E, Spruit CJP (1980) Genetic control of light-inhibited hypocotyl elongation in Arabidopsis thaliana. Z Pflanzenphysiol 100:147–160. https://doi.org/10.1016/S0044-328X(80)80208-X
Kumar DK, Khurana JP (2000) Isolation and characterization of OsCRY2 cDNA from rice (Oryza sativa L.) encoding a sensory blue light receptor, cryptochrome 2. M Phil thesis, University of Delhi, India
Lee J, Kang MH, Kim JY, Lim PO (2021) The role of light and circadian clock in regulation of leaf senescence. Front Plant Sci. https://doi.org/10.3389/fpls.2021.669170
Li YY, Mao K, Zhao C, Zhao XY, Zhang RF, Zhang HL, Shu HR, Hao YJ (2013) Molecular cloning and functional analysis of a blue light receptor gene MdCRY2 from apple (Malus domestica). Plant Cell Rep 32:555–566. https://doi.org/10.1007/s00299-013-1387-4
Lin C (1996) CRY2, a second member of the Arabidopsis cryptochrome gene family. Plant Physiol 110:1047
Lin C, Ahmad M, Gordon D, Cashmore AR (1995) Expression of an Arabidopsis cryptochrome gene in transgenic tobacco results in hypersensitivity to blue, UV-A, and green light. Proc Natl Acad Sci U S A 92:8423–8427. https://doi.org/10.1073/pnas.92.18.8423
Lin C, Yang H, Guo H, Mockler T, Chen J, Cashmore AR (1998) Enhancement of blue-light sensitivity of Arabidopsis seedlings by a blue light receptor cryptochrome 2. Proc Natl Acad Sci USA 95:2686–2690. https://doi.org/10.1073/pnas.95.5.2686
Liu H, Yu X, Li K, Klejnot J, Yang H, Lisiero D, Lin C (2008a) Photoexcited CRY2 Interacts with CIB1 to regulate transcription and floral initiation in Arabidopsis. Science. https://doi.org/10.1126/science.1163927
Liu LJ, Zhang YC, Li QH, Sang Y, Mao J, Lian HL, Wang L, Yang HQ (2008b) COP1-mediated ubiquitination of CONSTANS is implicated in cryptochrome regulation of flowering in Arabidopsis. Plant Cell 20:292–306. https://doi.org/10.1105/tpc.107.057281
Liu H, Liu B, Zhao C, Pepper M, Lin C (2011) The action mechanisms of plant cryptochromes. Trends Plant Sci 16(12):684–691. https://doi.org/10.1016/j.tplants.2011.09.002
Mas P, Devlin PF, Panda S, Kay SA (2000) Functional interaction of phytochrome B and cryptochrome 2. Nature 408:207–211. https://doi.org/10.1038/35041583
Matsumoto N, Hirano T, Iwasaki T, Yamamoto N (2003) Functional analysis and intracellular localization of rice cryptochromes. Plant Physiol 133:1494–1503. https://doi.org/10.1104/pp.103.025759.Shalitin
Matsumura H, Kitajima H, Akada S, Abe J, Minaka N, Takahashi R (2009) Molecular cloning and linkage map** of cryptochrome multigene family in soybean. Plant Genome 2:271–281. https://doi.org/10.3835/plantgenome.2009.06.0018
Mishra S, Khurana JP (2017) Emerging roles and new paradigms in signaling mechanisms of plant cryptochromes. CRC Crit Rev Plant Sci 36:89–115
Mockler TC, Guo H, Yang H, Duong H, Lin C (1999) Antagonistic actions of Arabidopsis cryptochromes and phytochrome B in the regulation of floral induction. Development 126:2073–2082. https://doi.org/10.1242/dev.126.10.2073
Mohanty A, Sarma NP, Tyagi AK (1999) Agrobacterium-mediated high frequency transformation of an elite indica rice variety Pusa Basmati 1 and transmission of the transgenes to R2 progeny. Plant Sci 147:127–137. https://doi.org/10.1016/S0168-9452(99)00103-X
Mouradov A, Cremer F, Coupland G (2002) Control of flowering time: Interacting pathways as a basis for diversity. Plant Cell 14:111–130. https://doi.org/10.1105/tpc.001362
Perrotta G, Ninu L, Flamma F, Weller JL, Kendrick RE, Nebuloso E, Giuliano G (2000) Tomato contains homologues of Arabidopsis cryptochromes 1 and 2. Plant Mol Biol 42:765–773. https://doi.org/10.1023/A:1006371130043
Platten JD, Foo E, Foucher F, Hecht V, Reid JB, Weller JL (2005) The cryptochrome gene family in pea includes two differentially expressed CRY2 genes. Plant Mol Biol 59:683–696. https://doi.org/10.1007/s11103-005-0828-z
Rascio N, Mariani P, Casadoro G (1984) Etioplast-chloroplast transformation in maize leaves: effects of tissue age and light intensity. Protoplasma 119:110–120. https://doi.org/10.1007/BF01287823
Sang Y, Li QH, Rubio V, Zhang YC, Mao J, Deng XW, Yang HQ (2005) N-terminal domain-mediated homodimerization is required for photoreceptor activity of Arabidopsis cryptochrome 1. Plant Cell 17:1569–1584. https://doi.org/10.1105/tpc.104.029645
Shalitin D, Yang H, Mockler TC, Maymon M, Guo H, Whitelam GC, Lin C (2002) Regulation of Arabidopsis cryptochrome 2 by blue-light-dependent phosphorylation. Nature 417:763–767. https://doi.org/10.1038/nature00815
Shalitin D, Yu X, Maymon M, Mockler T, Lin C (2003) Blue light-dependent in vivo and in vitro phosphorylation of Arabidopsis cryptochrome 1. Plant Cell 15:2421–2429. https://doi.org/10.1105/tpc.013011
Sharma R, Agarwal P, Ray S, Deveshwar P, Sharma P, Sharma N, Nijhawan A, Jain M, Singh AK, Singh VP, Khurana JP, Tyagi AK, Kapoor S (2012) Expression dynamics of metabolic and regulatory components across stages of panicle and seed development in indica rice. Funct Integr Genomics 12:229–248. https://doi.org/10.1007/s10142-012-0274-3
Sharma P, Chatterjee M, Burman N, Khurana JP (2014) Cryptochrome 1 regulates growth and development in Brassica through alteration in the expression of genes involved in light, phytohormone and stress signalling. Plant, Cell Environ 37:961–977. https://doi.org/10.1111/pce.12212
Sharma P, Mishra S, Burman N, Chatterjee M, Singh S, Pradhan AK, Khurana P, Khurana JP (2022) Characterization of Cry2 genes (CRY2a and CRY2b ) of B. napus and comparative analysis of BnCRY1 and BnCRY2a in regulating seedling photomorphogenesis. Plant Mol Biol. https://doi.org/10.1007/s11103-022-01293-6
Singh V, Singh AK, Mohapatra T, Gopala Krishnan S, Ellur RK (2018) Pusa Basmati 1121–a rice variety with exceptional kernel elongation and volume expansion after cooking. Rice. https://doi.org/10.1186/s12284-018-0213-6
Sperling U, Franck F, Van Cleve B, Frick G, Apel K, Armstrong GA (1998) Etioplast differentiation in Arabidopsis: both PORA and PORB restore the prolamellar body and photoactive protochlorophyllide-F655 to the cop1 photomorphogenic mutant. Plant Cell 10:283–296. https://doi.org/10.1105/tpc.10.2.283
Wang Q, Lin C (2020) Mechanisms of cryptochrome-mediated photoresponses in plants. Annu Rev Plant Biol 71:1–27. https://doi.org/10.1146/annurev-arplant-050718-100300
Wang H, Ma LG, Li JM, Zhao HY, Deng XW (2001) Direct interaction of Arabidopsis cryptochromes with COP1 in light control development. Science. https://doi.org/10.1126/science.1063630
Wei N, Deng XW (1996) The role of the COP/DET/FUS genes in light control of arabidopsis seedling development. Plant Physiol 112:871–878. https://doi.org/10.1104/pp.112.3.871
Wei FJ, Tsai YC, Wu HP, Huang LT, Chen YC, Chen YF, Wu CC, Tseng YT, Hsing Y, Ie C (2016) Both Hd1 and Ehd1 are important for artificial selection of flowering time in cultivated rice. Plant Sci 242:187–194. https://doi.org/10.1016/j.plantsci.2015.09.005
**e XZ, Chen ZP, Wang XJ (2005) Cloning and expression analysis of CRY2 gene in Sorghum bicolor. J Plant Physiol Mol Biol 31:261–268
Xu P, **ang Y, Zhu H, Xu H, Zhang Z, Zhang C, Zhang L, Ma Z (2009) Wheat cryptochromes: subcellular localization and involvement in photomorphogenesis and osmotic stress responses. Plant Physiol 149:760–774. https://doi.org/10.1104/pp.108.132217
Yang HQ, Wu YJ, Tang RH, Liu D, Liu Y, Cashmore AR (2000) The C termini of Arabidopsis cryptochromes mediate a constitutive light response. Cell 103:815–827. https://doi.org/10.1016/S0092-8674(00)00184-7
Yang Y, Fu D, Zhu C, He Y, Zhang H, Liu T, Li X, Wu C (2015) The ring-finger ubiquitin ligase HAF1 mediates heading date 1 degradation during photoperiodic flowering in rice. Plant Cell 27:2455–2468. https://doi.org/10.1105/tpc.15.00320
Yu X, Klejnot J, Zhao X, Shalitin D, Maymon M, Yang H, Lee J, Liu X, Lopez J, Lina C (2007) Arabidopsis cryptochrome 2 completes its posttranslational life cycle in the nucleus. Plant Cell 19:3146–3156. https://doi.org/10.1105/tpc.107.053017
Zhang YC, Gong SF, Li QH, Sang Y, Yang HQ (2006) Functional and signaling mechanism analysis of rice CRYPTOCHROME 1. Plant J 46:971–983. https://doi.org/10.1111/j.1365-313X.2006.02753.x
Zhang Q, Li H, Li R, Hu R, Fan C, Chen F, Wang Z, Liu X, Fu Y, Lin C (2008) Association of the circadian rhythmic expression of GmCRY1a with a latitudinal cline in photoperiodic flowering of soybean. Proc Natl Acad Sci USA 105:21028–21033. https://doi.org/10.1073/pnas.0810585105
Acknowledgements
This research was funded by the Department of Biotechnology, Government of India (BT/PR12394/AGIII/103/891/2014) and the J.C. Bose National Fellowship to JPK by the Science and Engineering Research Board (SERB), Government of India. The authors also acknowledge the infrastructural support provided by the Department of Science and Technology (FIST and PURSE programmes), Government of India, and the University Grants Commission (UGC-SAP), New Delhi. The award of Senior Research Fellowship from the Council of Scientific and Industrial Research (CSIR), New Delhi, to PS, SM and SS is duly acknowledged.
Funding
Department of Biotechnology, Government of India, BT/PR12394/AGIII/103/891/2014, Council for Scientific and Industrial Research, India, J.C. Bose National Fellowship, SB/SR/JCB-13/2013, Jitendra P Khurana.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Additional information
Communicated by Amit Dhingra.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Singh, S., Sharma, P., Mishra, S. et al. CRY2 gene of rice (Oryza sativa subsp. indica) encodes a blue light sensory receptor involved in regulating flowering, plant height and partial photomorphogenesis in dark. Plant Cell Rep 42, 73–89 (2023). https://doi.org/10.1007/s00299-022-02937-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-022-02937-z