Abstract
Key message
An ICE-like transcription factor mediates jasmonate-regulated cold tolerance in the rubber tree (Hevea brasiliensis), and confers cold tolerance in transgenic Arabidopsis.
Abstract
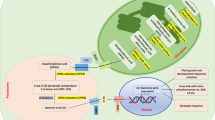
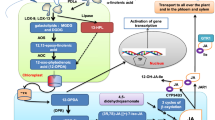
The rubber tree (Hevea brasiliensis) is susceptible to low temperatures, and understanding the mechanisms regulating cold stress is of great potential value for enhancing tolerance to this environmental variable. In this study, we find that treatment with exogenous methyl jasmonate (MeJA) could significantly enhance Hevea brasiliensis cold tolerance. In addition, yeast two-hybrid and bimolecular fluorescence complementation (BiFC) experiments show that JASMONATE ZIM-DOMAIN(JAZ) proteins, HbJAZ1 and HbJAZ12, key repressors of JA signaling pathway, interact with HbICE2, a novel ICE (Inducer of CBF Expression)-like protein. HbICE2 was nuclear-localised and bound to the MYC recognition (MYCR) sequence. The transcriptional activation activity of HbICE2 in yeast cells was dependent on the N-terminus, and overexpression of HbICE2 in Arabidopsis resulted in elevated tolerance to chilling stress. Furthermore, dual-luciferase transient assay reveals that HbJAZ1 and HbJAZ12 proteins inhibit the transcriptional function of HbICE2. The expression of C-repeat-binding factor (CBF) signalling pathway genes including HbCBF1, HbCBF2 and HbCOR47 were up-regulated by MeJA. Taken together, our data suggest that the new ICE-like transcription factor HbICE2 is involved in jasmonate-regulated cold tolerance in Hevea brasiliensis.










Similar content being viewed by others
Abbreviations
- ABA:
-
Abscisic acid
- ARRs:
-
Cold-regulated type-A response regulators
- BiFC:
-
Bimolecular fluorescence complementation
- CBF:
-
C-repeat-binding factor
- CBF/DREB1:
-
INDUCER OF CBF EXPRESSION (ICE)-C-REPEAT BINDING FACTOR/DRE BINDING FACTOR1
- COI1:
-
CORONATINE INSENSITIVE1
- EIN3:
-
ETHYLENE INSENSITIVE 3
- HOS1:
-
RING E3 ligase high expression of osmotically responsive gene 1
- ICE:
-
Inducer of CBF Expression
- JA:
-
Jasmonate
- JAZ:
-
JASMONATE ZIM-DOMAIN
- MAPK:
-
Mitogen-activated protein kinases
- MDA:
-
malondialdehyde
- MeJA:
-
methyl jasmonate
- MYCR:
-
MYC recognition
- OST1:
-
Open stomata1
- SIZ1:
-
SAP and Miz1
References
Agarwal M, Hao Y, Kapoor A, Dong CH, Fujii H, Zheng X, Zhu JK (2006) A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J Biol Chem 281:37636–37645
An XH, Hao YJ, Li EM, Xu K, Cheng CG (2017) Functional identification of apple MdJAZ2 in Arabidopsis with reduced JA-sensitivity and increased stress tolerance. Plant Cell Rep 36:255–265. https://doi.org/10.1007/s00299-016-2077-9
Cai W, Liu W, Wang WS, Fu ZW, Han TT, Lu YT (2015) Overexpression of rat neurons nitric oxide synthase in rice enhances drought and salt tolerance. PLoS One. https://doi.org/10.1371/journal.pone.0131599 (ARTN e0131599)
Cao SF, Zheng YH, Wang KT, ** P, Rui HJ (2009) Methyl jasmonate reduces chilling injury and enhances antioxidant enzyme activity in postharvest loquat fruit. Food Chem 115:1458–1463. https://doi.org/10.1016/j.foodchem.2009.01.082
Cheng H et al (2015) Functional characterization of hevea brasiliensis CRT/DRE binding factor 1 gene revealed regulation potential in the CBF pathway of tropical perennial tree. PLoS One 10:e0137634. https://doi.org/10.1371/journal.pone.0137634
Chinnusamy V, Ohta M, Kanrar S, Lee BH, Hong X, Agarwal M, Zhu JK (2003) ICE1: a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis. Genes Dev 17:1043–1054. https://doi.org/10.1101/gad.1077503
Chinnusamy V, Zhu J, Zhu JK (2007) Cold stress regulation of gene expression in plants. Trends Plant Sci 12:444–451. https://doi.org/10.1016/j.tplants.2007.07.002
Cutler SR, Ehrhardt DW, Griffitts JS, Somerville CR (2000) Random GFP::cDNA fusions enable visualization of subcellular structures in cells of Arabidopsis at a high frequency. Proc Natl Acad Sci U S A 97:3718–3723. https://doi.org/10.1073/pnas.97.7.3718
Deng YM, Chen SM, Chen FD, Cheng X, Zhang F (2011) The embryo rescue derived intergeneric hybrid between chrysanthemum and Ajania przewalskii shows enhanced cold tolerance. Plant Cell Rep 30:2177–2186. https://doi.org/10.1007/s00299-011-1123-x
Ding Y, Li H, Zhang X, **e Q, Gong Z, Yang S (2015) OST1 kinase modulates freezing tolerance by enhancing ICE1 stability in Arabidopsis. Dev Cell 32:278–289. https://doi.org/10.1016/j.devcel.2014.12.023
Dong CH, Agarwal M, Zhang Y, **e Q, Zhu JK (2006) The negative regulator of plant cold responses, HOS1, is a RING E3 ligase that mediates the ubiquitination and degradation of ICE1. Proc Natl Acad Sci U S A 103:8281–8286. https://doi.org/10.1073/pnas.0602874103
Fursova OV, Pogorelko GV, Tarasov VA (2009) Identification of ICE2, a gene involved in cold acclimation which determines freezing tolerance in Arabidopsis thaliana. Gene 429:98–103. https://doi.org/10.1016/j.gene.2008.10.016
Gonzalez-Aguilar GA, Fortiz J, Cruz R, Baez R, Wang CY (2000) Methyl jasmonate reduces chilling injury and maintains postharvest quality of mango fruit. J Agric Food Chem 48:515–519. https://doi.org/10.1021/jf9902806
Guo XY, Liu DF, Chong K (2018) Cold signaling in plants: Insights into mechanisms and regulation. J Integr Plant Biol 60:745–756. https://doi.org/10.1111/jipb.12706
Hao BZ, Ji-Lin WU (2000) Laticifer differentiation in Hevea brasiliensis: induction by exogenous jasmonic acid and linolenic acid. Ann Bot 85:37–43
Hu Y, Jiang L, Wang F, Yu D (2013) Jasmonate regulates the inducer of cbf expression-C-repeat binding factor/DRE binding factor1 cascade and freezing tolerance in Arabidopsis. Plant Cell 25:2907–2924. https://doi.org/10.1105/tpc.113.112631
Hu Y, Jiang Y, Han X, Wang H, Pan J, Yu D (2017) Jasmonate regulates leaf senescence and tolerance to cold stress: crosstalk with other phytohormones. J Exp Bot 68:1361–1369. https://doi.org/10.1093/jxb/erx004
Huang XS, Zhang Q, Zhu D, Fu X, Wang M, Moriguchi T, Liu JH (2015) ICE1 of Poncirus trifoliata functions in cold tolerance by modulating polyamine levels through interacting with arginine decarboxylase. J Exp Bot 66:3259–3274. https://doi.org/10.1093/jxb/erv138
Ito T, Ng KH, Lim TS, Yu H, Meyerowitz EM (2007) The homeotic protein AGAMOUS controls late stamen development by regulating a jasmonate biosynthetic gene in Arabidopsis. Plant Cell 19:3516–3529. https://doi.org/10.1105/tpc.107.055467
Laurens P, Alain G (2011) The JAZ proteins: a crucial interface in the jasmonate signaling cascade. Plant Cell 23:3089–3100. https://doi.org/10.1105/tpc.111.089300
Li C et al (2005) Role of beta-oxidation in jasmonate biosynthesis and systemic wound signaling in tomato. Plant Cell 17:971–986. https://doi.org/10.1105/tpc.104.029108
Li H, Ding Y, Shi Y, Zhang X, Zhang S, Gong Z, Yang S (2017) MPK3- and MPK6-mediated ICE1 phosphorylation negatively regulates ice1 stability and freezing tolerance in arabidopsis. Dev Cell 43(5):630–642.e4. https://doi.org/10.1016/j.devcel.2017.09.025
Liang C, Hui Z, Feng R, Qian-Qian G, Xu-Peng H, Xue-Bao L (2011) A novel cold-regulated gene, COR25, of Brassica napus is involved in plant response and tolerance to cold stress. Plant Cell Rep 30(4):463–471. https://doi.org/10.1007/s00299-010-0952-3
Liu JH, Inoue H, Moriguchi T (2008) Salt stress-mediated changes in free polyamine titers and expression of genes responsible for polyamine biosynthesis of apple in vitro shoots. Environ Exp Bot 62:28–35. https://doi.org/10.1016/j.envexpbot.2007.07.002
Lyons JM (1973) Chilling injury in plants annurevplant. Physiol 24:445–466
Mantyla E, Lang V, Palva ET (1995) Role of abscisic acid in drought-induced freezing tolerance, cold acclimation, and accumulation of LT178 and RAB18 proteins in arabidopsis thaliana. Plant Physiol 107:141–148. https://doi.org/10.1104/pp.107.1.141
Melotto M et al (2008) A critical role of two positively charged amino acids in the Jas motif of Arabidopsis JAZ proteins in mediating coronatine- and jasmonoyl isoleucine-dependent interactions with the COI1 F-box protein. Plant J Cell Mol Biol 55:979–988. https://doi.org/10.1111/j.1365-313X.2008.03566.x
Meng XH, Han J, Wang Q, Tian SP (2009) Changes in physiology and quality of peach fruits treated by methyl jasmonate under low temperature stress. Food Chem 114:1028–1035. https://doi.org/10.1016/j.foodchem.2008.09.109
Miura K et al (2007) SIZ1-mediated sumoylation of ICE1 controls CBF3/DREB1A expression and freezing tolerance in Arabidopsis. Plant Cell 19:1403–1414. https://doi.org/10.1105/tpc.106.048397
Schutze K, Harter K, Chaban C (2009) Bimolecular fluorescence complementation (BiFC) to study protein-protein interactions in living plant cells. Methods Mol Biol 479:189–202. https://doi.org/10.1007/978-1-59745-289-2_12
Shi Y, Tian S, Hou L, Huang X, Zhang X, Guo H, Yang S (2012) Ethylene signaling negatively regulates freezing tolerance by repressing expression of CBF and type-A ARR genes in Arabidopsis Plant
Shi Y, Ding Y, Yang S (2015) Cold signal transduction and its interplay with phytohormones during cold acclimation. Plant Cell Physiol 56:7–15. https://doi.org/10.1093/pcp/pcu115
Spoel SH, Dong X (2008) Making sense of hormone crosstalk during plant immune responses. Cell Host Microbe 3:348–351. https://doi.org/10.1016/j.chom.2008.05.009
Thines B et al (2007) JAZ repressor proteins are targets of the SCF(COI1) complex during jasmonate signalling. Nature 448:661–665. https://doi.org/10.1038/nature05960
Thomashow MF (1999) Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Annu Rev Plant Physiol Plant Mol Biol 50:571–599. https://doi.org/10.1146/annurev.arplant.50.1.571
Toledo-Ortiz G, Huq E, Quail PH (2003) The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 15:1749–1770. https://doi.org/10.1105/tpc.013839
Walter M et al (2004) Visualization of protein interactions in living plant cells using bimolecular fluorescence complementation. The Plant Journal: Cell Mol Biol 40:428–438. https://doi.org/10.1111/j.1365-313X.2004.02219.x
Wasternack C, Strnad M (2017) Jasmonates are signals in the biosynthesis of secondary metabolites—pathways, transcription factors and applied aspects—a brief review. New Biotechnol 25:48:1–11. https://doi.org/10.1016/j.nbt.2017.09.007
**e DX, Feys BF, James S, Nieto-Rostro M, Turner JG (1998) COI1: an Arabidopsis gene required for jasmonate-regulated defense and fertility. Science 280:1091–1094. https://doi.org/10.1126/science.280.5366.1091
Xu W, Jiao Y, Li R, Zhang N, **ao D, Ding X, Wang Z (2014) Chinese wild-growing Vitis amurensis ICE1 and ICE2 encode MYC-type bHLH transcription activators that regulate cold tolerance in Arabidopsis. PLoS One 9:e102303. https://doi.org/10.1371/journal.pone.0102303
Yuan HM, Sheng Y, Chen WJ, Lu YQ, Tang X, Ou-Yang M, Huang X (2017) Overexpression of Hevea brasiliensis HbICE1 enhances cold tolerance in arabidopsis. Front Plant Sci https://doi.org/10.3389/Fpls.2017.01462
Zhao ML et al (2013) Induction of jasmonate signalling regulators MaMYC2s and their physical interactions with MaICE1 in methyl jasmonate-induced chilling tolerance in banana fruit. Plant Cell Environ 36:30–51. https://doi.org/10.1111/j.1365-3040.2012.02551.x
Zhu J et al (2015) Low temperature inhibits root growth by reducing auxin accumulation via ARR1/12. Plant Cell Physiol 56:727–736. https://doi.org/10.1093/pcp/pcu217
Funding
This work was supported by the National Natural Science Foundation of China [#31872045 and #31560197], and Youth Foundation of Hainan University [# hdkyxj201703].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Ying-Tang Lu.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, WJ., Wang, X., Yan, S. et al. The ICE-like transcription factor HbICE2 is involved in jasmonate-regulated cold tolerance in the rubber tree (Hevea brasiliensis). Plant Cell Rep 38, 699–714 (2019). https://doi.org/10.1007/s00299-019-02398-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-019-02398-x