Abstract
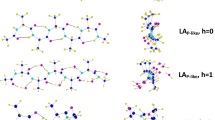
The sequence effects on the conformations and intra-molecular hydrogen bonds (NH···OC: H-bond) of collagen-like co-peptides (single chains) were analyzed using co-glycine (G)/L-proline (P) oligopeptides (6- and 9-mer, G/P = 1/2 mol), by QCC [B3LYP/6-31G(d,p)], with homo-oligopeptides. Each sequence was specified using three kinds of basic triplets of GPP, PGP, and PPG. The conformations (dihedral angles: φn/ψn) varied depending on the sequences. The conformation types of each unit were classified in the following three groups: φ/ψ of g−/g + (shorter polyproline II helical structure: P-like) with H-bond, g−/t + (polyproline II helical structure: PPII) with no H-bond, and φ/ψ of t−/g + with no H-bond. All H-bonds that have seven-membered ring were formed between units separated by one peptide. The positions and number (Nh) of H-bonds varied depending on the sequences. The H-bonds between CO of proline and NH of glycine unit were newly formed. The co-peptides that have no GPP triplet in N-end side (CH3CO-GPP) were found to be more stable than the corresponding blend models due to the formations of newly formed H-bonds. From the comparison with the structures observed for triple helix of collagen in other works and the findings (the existence of the CO group that have no H-bond), it was inferred that collagen may be formed by conformational rearrangement from P-like (single chain) to PPII (triple helix) and may be stabilized by inter-molecular hydrogen bonding between the residual CO groups and water molecules.






Similar content being viewed by others
References
Okuyama K, Xu X, Iguchi M, Noguchi K (2006) Revision of collagen molecular structure. Biopolymers 84:181–191
Ramachandran GN, Kartha G (1955) Structure of collagen. Nature (London) 176:593–595
Rich A, Crick FHC (1961) The molecular structure of collagen. J Mol Biol 3:483–506
Sakakibara S, Kishida Y, Okuyama K, Tanaka N, Ashida T, Kakudo M (1972) Single crystals of (Pro-Pro-Gly)10, a synthetic polypeptide model of collagen single crystals of (Pro-Pro-Gly)10. J Mol Biol 65:371–373
Okuyama K, Takayanagi M, Ashida T, Kakudo M (1977) A New structural model for collagen. Polymer J 9:341–343
Stimson ER, Zimmerman SS, Scheraga HA (1977) Conformational studies of oligopeptides containing proline and glycine. Macromolecules 10(5):1049–1060
Abagyan RA, Tumanian VG, Esipova NG (1984) Two types of tripeptide conformation in collagen: calculation of the structure of (Gly-Pro-Ser)n and (Gly-Val-Hyp)n polytripeptides. Bioorg Khim 10(4):476–482
Ramshaw JA, Shah NK, Brodsky B (1998) Gly-X-Y tripeptide frequencies in collagen: a context for host-guest triple-helical peptides. J Struct Biol 122:86–91
Krieger F, Möglich A, Kiefhaber T (2005) Effect of proline and glycine residues on dynamics and barriers of loop formation in polypeptide chains. J Am Chem Soc 127(10):3346–3352
Bella J, Liu J, Kramer R, Brodsky B, Berman HM (2006) Conformational effects of Gly-X-Gly interruptions in the collagen triple helix. J Mol Biol 362(2):298–311
Okuyama K, Narita H, Kawaguchi T, Noguchi T, Tanaka Y, Nishino N (2007) Collagen structure re-vised. Biopolymers 86:212–221
Okuyama K, Kawaguchi T (2010) Molecular and fibrillar Structures of collagen. Kobunshi Ronbunshu 67:229–247
Andrew H, Kangt & Jerome Gross, (1970) Relationship between the Intra and Intermolecular Cross-links of Collagen. PNAS 67(3):1307–1314
Kobayashi M, Sim JH, Sato H (2019) Conformational analyses for hydrated oligopeptides by quantum chemical calculation (QCC): effects of intra-molecular hydrogen bonds. Polym Bull 76:3247–3268
Kobayashi M, Sim JH, Sato H (2015) Conformational analyses for alanine oligomer during chain propagation by quantum chemical calculation. Polymer J 47:369–378
Graf J, Nguyen PH, Stock G, Schwalbe H (2007) Structure and dynamics of the homologous series of alanine peptides: a joint molecular dynamics/NMR study. J Am Chem Soc 129:1179–1189
Rigaudy J, Klesney SP (1979) Nomenclature of Organic Chemistry, Section E, 483. Pergamon Press, Oxford
“Gaussian 03 User’s Reference”, Gaussian Inc., Carnegie, PA, 2003.
Accessed May 2019 from https://en.wikipedia.org/wiki/Polyproline_helixProline:net
Balasubramanian R, Lakshminarayanan AV, Sabesan MN, Tegoni G, Venkatesan K, Ramachandran GN (1971) Studies on the conformation of amino acids, VI: conformation of the proline ring as observed in crystal structures of amino acids and peptides. Int J Protein Res 3(1):25–33
Hongo C, Noguchi K, Okuyama K, Tanaka Y, Nishino N (2005) Repeptive interactions observed in the crystal structure of a collagen-model peptide, [(Pro-Pro-Gly)9]3. J Biochem 138:135–144
Nagy PI (2014) Competing Intramolecular vs. Intermolecular Hydrogen Bonds in Solution. Int J Mol Sci 15:19562–19633
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kobayashi, M., Sim, J.H. & Sato, H. Conformational analyses of collagen-like Co -Glycine/L-proline oligopeptides by quantum chemical calculation (QCC): Sequence effects on conformations and intra-molecular hydrogen bonds . Polym. Bull. 79, 6627–6644 (2022). https://doi.org/10.1007/s00289-021-03805-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00289-021-03805-8