Abstract
Aspergillus fumigatus is a ubiquitous pathogenic mold and causes several diseases, including mycotoxicosis, allergic reactions, and systemic diseases (invasive aspergillosis), with high mortality rates. In its ecological niche, the fungus has evolved and mastered many reply strategies to resist and survive against negative threats, including harsh environmental stress and deficiency of essential nutrients from natural environments, immunity responses and drug treatments in host, and competition from symbiotic microorganisms. Hence, treating A. fumigatus infection is a growing challenge. In this review, we summarized A. fumigatus reply strategies and escape mechanisms and clarified the main competitive or symbiotic relationships between A. fumigatus, viruses, bacteria, or fungi in host microecology. Additionally, we discussed the contemporary drug repertoire used to treat A. fumigatus and the latest evidence of potential resistance mechanisms. This review provides valuable knowledge which will stimulate further investigations and clinical applications for treating and preventing A. fumigatus infections.
Key points
• Harsh living environment was a great challenge for A. fumigatus survival.
• A. fumigatus has evolved multiple strategies to escape host immune responses.
• A. fumigatus withstands antifungal drugs via intrinsic escape mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aspergillus fumigatus is the main pathogenic fungus underlying aspergillosis and globally causes more than 300,000 life-threatening infections every year (Earl Kang and Celia 2021). Moreover, in recent years, during the Coronavirus 2019 (COVID-19) pandemic, pulmonary aspergillosis led to Aspergillus co-infections with COVID-19 developed into a clinical, major life-threatening fungal disease (Giacobbe et al. 2022; Hoenigl 2021). As a saprophytic fungal pathogen, A. fumigatus proliferates via abundant, small diameter (2–3 μm), air-borne asexual conidia. Although the fungus may encounter harsh natural conditions, such as high temperatures, poor carbon or nitrogen sources, ultraviolet light, and reduced metal ion levels, it has evolved adaptive survival systems. Generally, conidiophores generate thousands of conidia; indoor and outdoor airborne conidia concentrations can range from 1 to 100 conidia/m3 and even reach up to 108 conidia/m3 in certain environments (Latgé and Chamilos 2019). Moreover, abundant conidia are inhaled daily into human lungs (Furlong-Silva and Cook 2022). In immunocompetent hosts, approximately 90% of inhaled conidia are swiftly cleared at mucosal surfaces and ciliated cells in the respiratory tract. Residually activated and swollen conidia face hostile environments, including low carbon sources, low oxygen concentrations, and host immune responses (Margalit and Kavanagh 2015); therefore, A. fumigatus survival in healthy individuals poses challenges to the fungus. However, A. fumigatus can evade host defenses for the spores to germinate or proliferate and causes aspergillosis in immunocompromised and immunodeficient individuals (Verburg et al. 2022).
When A. fumigatus breaks through host airway defenses and accesses the lungs, a chain reaction of antifungal host responses is activated. For example, germinating conidia may be phagocytized by alveolar macrophages (AMs) while germ tubes are quickly and effectively targeted by neutrophils (Ortiz et al. 2022). However, several complex fungal regulatory systems, such as altering recognition receptors, releasing reactive oxygen species (ROS) detoxifying enzymes, as well as biofilm and aspergilloma development, have been evolved to cope with the host immune responses (Latgé and Chamilos 2019). A. fumigatus also appears to co-operate with viruses, bacteria, and fungi to facilitate multiple colocalization sites that augment A. fumigatus invasion and its seriousness. Currently, A. fumigatus is mainly treated using azoles, echinocandins, and polyenes. Especially the itraconazole, voriconazole, isavuconazole, and posaconazole were the primary treatment drugs (Ben-Ami 2023). However, with long-term drug use, drug-resistant strains can survive via targeted protein overexpression or mutation, efflux pump overexpression, and altered mitochondria-related the high-osmolarity glycerol (HOG)-mitogen-activated protein kinase (MAPK) signaling and Heat Shock Protein (Hsp)90-calcineurin pathways. Therefore, A. fumigatus infection requires serious clinical attention.
In this review, recent research progress on A. fumigatus and its ability to confront negative factors in the natural environment, host defenses, and other lung-colonizing microorganisms are outlined. Importantly, fungal adaptation mechanisms, incorporating perception, regulation, response, and adaptation processes, are examined. Also, current fungal therapies and major fungal drug resistance mechanisms are discussed. This review could provide valuable insights on A. fumigatus infection prevention and treatment measurements.
Fungal adaption in the natural environment
A. fumigatus is ubiquitous in external environments, including soil, decaying vegetation, and air, and has evolved many regulatory mechanisms to adapt growth to different factors (temperature, pH, carbon and nitrogen sources, and metallic ions). A. fumigatus grows and survives in extreme environments in temperatures up to 70 °C by exploiting multiple temperature regulatory systems (Hokken et al. 2023). In 2010, A. fumigatus proteome alterations during 30–48 ℃ shifts were examined by mass spectrometry and showed that Hsp30, Hsp42, and Hsp90 proteins were highly elevated after heat shock (Albrecht et al. 2010). Hsp90 is a conserved and essential eukaryotic molecular chaperone; it mediates proteostasis to avoid protein damage and misfolding during hyperthermy (Hervás and Oroz 2020). Hsp90 also interacts with cell wall integrity pathway (CWIP) components which are crucial signaling pathways maintaining cell viability under thermal stress (Crunden and Diezmann 2021). With the temperature increase, Hsp90 could interact with the CWIP component MAPK to prevent its aggregation (Rocha and Minari 2021). Furthermore, hsp expression is regulated by heat shock transcription factors (HSFs). Under stress, HSF factor 1 (Hsf1) homotrimers bind to specific DNA motifs or heat shock elements in target gene promoters to induce thermal adaptation gene expression, such as Hsp90 and Hsp70. Hsp90 also regulates Hsf1 activity by binding to the Hsf1 feedback regulatory loop to maintain its inactive state under normal conditions (Fabri et al. 2021). Additionally, many other genes are reportedly involved in A. fumigatus thermotolerance regulation, e.g., a putative α-1,2-mannosyltransferase deletion strain exhibited thinner hyphal cell walls when compared with the parental strain at 48 ℃. The thermotolerance gene thtA was essential for A. fumigatus growth at 48 ℃, while a thtA deletion mutant was more sensitive at this temperature when compared with the original strain (Chang et al. 2004; Wagener et al. 2008). Thermotolerance regulated genes from the literature are summarized (Table 1).
Nutrients are essential for A. fumigatus survival as the fungus has evolved highly sophisticated homeostatic mechanisms to respond to, take up, recycle, and use many varied nutrients. In the natural environment, nitrogen predominantly occurs in inorganic forms: atmospheric nitrogen, ammonia, nitrite, and nitrate. As a nitrogen sensor in A. fumigatus, rapamycin (TOR) protein was crucial for this fungal, and the gene missing mutant was more sensitivity and deficits in germination ability on poor nitrogen medium compared to wild strains. TOR promoted GLN3 (promoted GATA transcription factor) binding to cytoplasmic protein URE2 to control cytoplasmic protein synthesis and degradation under nitrogen-limited conditions (Baldin et al. 2015; Beck and Hall 1999). The Ras-related gene rhbA deletion mutant exhibited TOR kinase inhibitor rapamycin hypersensitivity and also had significantly reduced growth rates on poor nitrogen-sources (Panepinto et al. 2003). The brlA gene (C2H2 zinc finger transcription factor) deletion mutant exhibited downregulated gene expression encoding ribosomal proteins under nitrogen-limiting conditions; however, the mutant was insensitive to a TOR inhibitor, suggesting that brlA was not downstream of TOR signaling (Twumasi-Boateng et al. 2009). Furthermore, the stress-activated protein kinases (SakA)/HogA MAPK pathway was activated upon nitrogen starvation during vegetative growth (Ma and Li 2013). SakA is part of the nitrate and nitrite assimilation cascade during fungal germination processes. When compared with the wild-type stain, the sakA defective mutant showed increased germination rates on limited nitrogen medium and demonstrated that MAPK negatively regulated conidial germination (Perez-Cuesta and Guruceaga 2021). Other major uptake and metabolic gene regulators of non-favored nitrogen sources include the zinc finger transcription factor GATA-like genes areA and areB. AreA controls the expression of the glycosylphosphatidylinositol-anchored protein SwgA, which is localized to membranes and is involved in germination, growth, and morphogenesis (Samalova et al. 2023). On primary nitrogen sources, AreA interacts with NmrA protein (nitrogen metabolite repression compound) to inhibit the induction of secondary nitrogen sources (Andrianopoulos et al. 1998). However, when primary nitrogen sources are not present, tamA (Zn(II)2Cys6 transcription factor) interacts with areA to activate nitrogen catabolism (Downes et al. 2014). AreB is generally regarded as a negative nitrogen metabolism regulation factor, and AreB expression depends on AreA and AreB to negatively regulate AreA-dependent nitrogen catabolic gene expression under nitrogen-repressing or starvation conditions (Macios et al. 2012). Additionally, GcnE (acetyltransferase (Lin et al. 2020)), RgsC (G-protein signaling protein(Kim et al. 2017)), and Met6 (a bifunctional dehydrogenase/ferrochelatase enzyme (Dietl et al. 2018)) may also be involved in nitrogen metabolism; however, little is known about these processes. The regulatory genes responsible for A. fumigatus growth on different nitrogen sources are summarized (Table 1).
Environmental fungi carbon sources mainly include glucose, lactate, and acetate. Glucose is the most favored carbon source for fungal survival and niche colonization. It was reported that the G-protein coupled receptor system and the hexose transporter are required to sense and uptake glucose (Qadri et al. 2021; Ries et al. 2018). For example, a member of this family GprK was characterized as a carbon-sensing receptor in A. fumigatus. Gene loss mutants showed increasing germination rates under carbon starvation and growth restriction levels on a medium containing a sole carbon source (pentose) (Kim et al. 2017). Also, HOG/CWI pathways were implicated in carbohydrate metabolism. The Hog1p upstream signaling receptors Sho1p, Msb2p mucin, and Opy2p genetically interacted, while their null mutants showed altered trehalose and glycogen accumulation, suggesting regulated sugar storage by the HOG/CWI pathway (Silva and Frawley 2020). Another system called Carbon Catabolite Repression showed preferences for glucose or preferred sugars. The C2H2 zinc-finger transcription factor creA and the regulatory controller facB were available for extracellular glucose and acetate utilization (Ries et al. 2021). Genes related to carbon source regulation are summarized (Table 1).
Metal ion metabolism affects almost all A. fumigatus biological functions, including fungal virulence, cell wall integrity, azole susceptibility, protein phosphatases, antigen secretion, signal transduction, and even mitochondrial functions (Blatzer and Latgé 2017). During iron shortages, A. fumigatus generally increases expression of hapX (bZip CCAAT-binding transcription factor), sidA (ornithine monooxygenase), and mirB (siderophore transporter), while down-regulating sreA (GATA transcription factor), cccA (vacuole iron importer), and cycA (cytochrome C) during iron abundant conditions to maintain iron homeostasis (Matthaiou et al. 2018). Under iron starvation, HapX interacts with the CCAAT-binding core complex to activate iron acquisition and siderophore transporters, repress iron-consuming processes, and the vacuolar iron transporter pathway (Schrettl et al. 2010). During iron excess, SreA represses hapX expression, represses iron uptake, and promotes its use (Wiemann et al. 2014). Similarly, in the case of excess iron, the intracellular siderophore ferricrocin and vacuole was important to detoxification, and the vacuole iron importer encoding gene cccA deficiency decreases iron resistance (Gsaller et al. 2012). Also, the leucine biosynthetic and signal-transduction pathways, phosphatase Z and TOR kinases, are reportedly required during adverse iron conditions (Orasch et al. 2019).
Zinc is essential for fungal growth, and zinc homeostasis in A. fumigatus is regulated upon the external condition. During zinc affluent conditions and to transport redundant zinc to extracellular spaces or vacuoles for detoxification, aceA (transcription factor) induces crpA (Cu+ P-type ATPase) and zrcA (vacuolar zinc transporter) expression (Cai et al. 2018). Under zinc-deficient conditions, ZafA (transcriptional activator) up-regulates ZrfA and ZrfB (zinc transporters) in acidic or neutral conditions, while in the alkaline with calprotectin, it mainly up-regulated ZrfC to reduce zinc consumption (Amich and Calera 2014; Amich et al. 2009). Additionally, ZrfA, ZrfB, and ZrfC expression is further modulated by PacC (transcription factor) depending on ambient pH (Toledo et al. 2022). ZafA-mediated fungal growth regulation is also influenced by iron availability, which is enhanced in zinc- and iron-repleted media, but growth is restricted by reducing zinc intake under iron starvation (Vicentefranqueira et al. 2019). Metal ion metabolism genes are summarized (Table 1).
Thus, A. fumigatus has developed several effective mechanisms to survive adverse conditions and combat stress-related changes. It was reported that more than 80 A. fumigatus strains have been isolated from Arctic soils (Korfanty et al. 2021). Apart from normal regulatory mechanisms (temperature, carbon, nitrogen, and metal ion acquisition), several highly coordinated adaptation mechanisms are also used to exploit other external environmental conditions. For example, A. fumigatus can be grown in 5% carbon dioxide, very low pH (3.5), and under ultraviolet radiation. Proteome analyses have shown that stress responses, including cell wall reorganization, DNA repair, and oxidative stress responses during citric acid and itaconic acid production, are increased to overcome radiation effects (Alonso et al. 2017; Oliveira et al. 2021). However, these data constitute only a small component of A. fumigatus survival mechanisms. When A. fumigatus is inhaled into the human body, it faces a more complex internal environment.
A. fumigatus survival strategies in hosts
When A. fumigatus conidia invade organisms, they become metabolically active and swell. To quickly clear spores, host responses are physically initiated via mucosal surface barriers in the respiratory tract. On epithelial surfaces in upper and lower respiratory tracts, most conidia are trapped in mucus and removed via ciliated cell actions (Crossen and Ward 2022). In healthy individuals, mucociliary clearance and phagocytic defenses normally prevent fungal-associated diseases; thus, A. fumigatus isolation from respiratory secretions in normal hosts generally reflects colonization rather than infection (Gago et al. 2019). However, in immunocompromised hosts, A. fumigatus can attach to sinonasal epithelial cell monolayers to form three-dimensional biofilm structures with parallel-packed, cross-linked hyphae and channels to induce sinusitis or tracheobronchitis (Singhal et al. 2011). Consequently, conidia breakdown barriers to reach the lower respiratory tract, and compromised lung epithelia provide an entry portal for fungi (Bertuzzi et al. 2018). In alveolar epithelium, epithelial cells covering over 95% of inner alveolar surfaces function as efficient Aspergillus conidia neutralizers via actin-dependent phagocytosis in mature acidified phagolysosomes or by endocytosis induced by protein–protein interactions between the host and pathogen (Latgé and Chamilos 2019). For example, lung mucin glycoproteins contain twelve binding sites for fucosylated structures and avidly bind to FleA (lectin), which is expressed by A. fumigatus, while integrin α5β1 interacts with A. fumigatus CalA (thaumatin-like protein) (Liu et al. 2016; Richard et al. 2018). However, most pathogens control host innate immune responses at early stages, before infiltrating host immune cells arrive at infection sites. For example, A. fumigatus Aspf2 (factor H-binding protein) blocks host innate immune attack at early infection stages. Similarly, to avoid C3b complement system activation, A. fumigatus recruits several human plasma regulators (factor H, factor-H-like protein 1, and factor H-related protein 1). Aspf2 also recruits plasminogen to damage human lung epithelial cells, induce cell retraction, and expose the matrix. Therefore, when A. fumigatus is not phagocytosed, tissue is penetrated (Dasari et al. 2018). High-resolution live-cell confocal microscopy assays have indicated that A. fumigatus spores could survive from the maturation failure phagosome (about 60%), and the hyphae would fused to the host plasma membrane rather than rupture the phagolysosomal membrane to allow it growth better. Then, hyphae escape from epithelial lung cells in a non-lytic manner and elongate to adjacent cells without penetrating the host cytoplasm (Seidel et al. 2020). And the dihydroxynaphthalene (DHN)-melanin, the additional layers in the outer part of the conidia cell wall, was one of the interference factors for host endocytosis. Proteomics analyses of A. fumigatus conidia-containing phagolysosomes have shown that melanin inhibits phagolysosome acidification, Rab5- and Vamp8-mediated endocytic trafficking, and cathepsin Z (lysosomal cysteine protease) recruitment. Therefore, melanin promotes conidia germination and escapes from AMs via hyphal growth (Amin et al. 2014). Melanin is also involved in fungal adhesion and biofilm formation, enhanced fungal immune tolerance, and decreased exposure to pathogen-associated molecular patterns (PAMPs) to limit phagocyte phagocytosis. A recent study reported that melanin also removed chemokines (CXCL10 and CCL20) to suppress host inflammatory responses (Graf et al. 2023).
Due to continuous inflammatory response activation during A. fumigatus infection, pulmonary would appear local tissue hypoxia (Gago et al. 2019). As oxygen is essential for A. fumigatus biochemical processes, the fungus adapts to oxygen limitations. Transcriptomic and proteomic analyses have shown that during glycolysis induction, the transcriptional down-regulation of the tricarboxylic acid cycle and oxidative phosphorylation processes are major hypoxia-response measures. Transcripts were associated with iron and sterol metabolism, the cell wall, and GABA shunts, which were significantly increased to cope with stress (Barker et al. 2012). It was reported that TcsC (Group III two-component sensor kinase) was required for adapting fungi to low oxygen levels. Low oxygen caused TcsC-dependent phosphorylation of SakA, and the ΔtcsC mutant was susceptible to increased morphogenetic changes (McCormick et al. 2012). Similarly, mitochondrial respiration is also critical for fungal pathogenesis during hypoxia. In a mouse model, the mitochondrial respiration chain component cycA (cytochrome C gene) and alcC (alcohol dehydrogenase gene) deleted strains have been come out the defect virulence (Grahl et al. 2012, 2011). Chromatin immunoprecipitation followed by parallel DNA sequencing showed that SrbA (sterol regulatory element-binding protein gene) helped regulate ergosterol biosynthesis and iron uptake during hypoxic conditions or iron limitation (Zhang et al. 2021a). Critically, the lung is a “sponge,” and there are not enough carbon or nitrogen sources on lung surfaces to limit A. fumigatus growth. Therefore, A. fumigatus generates proteases (serine proteases, metalloproteinases, and aspartic proteases) to decompose organic components (Abad et al. 2010). Besides these genes above-mentioned about the carbon and nitrogen metabolism, the transcription factors are also important for A. fumigatus survival in vivo. For example, facB (transcription regulatory factor required for acetate utilization) is essential for carbon metabolism in vivo. A facB deficient strain showed significantly reduced virulence in both Galleria mellonella and murine invasive pulmonary aspergillosis (IPA) models (Ries et al. 2021). During mold infection, amino acid biosynthesis is required for nitrogen metabolism. The cpcA gene encodes the transcriptional activator of the cross-pathway control system (CPC) of amino acid biosynthesis. Indeed, a cpcA deletion strain not only impaired the CPC system in terms of amino acid starvation, but also attenuated virulence in pulmonary aspergillosis mice (Krappmann et al. 2004).
Esca** immune responses
After A. fumigatus evades the host’s upper respiratory tract, it can survive on lung surfaces where conidia germinate and form invasive hyphae which penetrate pulmonary tissues and enter alveoli. AMs are first-line innate host defenses and use pathogen-recognition receptors (PRR) and PAMPs. Toll-like receptors (TLRs) are a major PRR class responsible for activated innate immune responses, especially TLR2 and TLR4, which recognize fungal PAMPs, including, peptidoglycans, RNAs, zymosan, lipopolysaccharide, and HSPs (Kumar 2022). However, the TLR4 induces signals were responding just in the stimulation of conidia, while A. fumigatus germinates into hyphae, the TLR4-mediated signaling would be loosed (Netea et al. 2003). Thus, the main effects of proinflammatory cytokines come from TLR2-activated non-protective Th2 (T-helper 2) responses (Buckland et al. 2008). Meanwhile, the Aspergillus gliotoxin produced by A. fumigatus could target the host cell phosphatidylinositol 3,4,5-trisphosphate [PtdIns(3,4,5)P3] metabolism to break the phagocytes protective functions, so that this pathogen could escape the macrophage recognition and downregulating phagocytic immune defenses (Schlam et al. 2016). Macrophages and neutrophils also generate ROS to combat A. fumigatus conidia and hyphae (Henriet et al. 2011; Shlezinger and Hohl 2021), e.g., if nicotinamide adenine dinucleotide phosphate hydrogen oxidase is blocked and the ROS generation by neutrophils was disturbed, this could significantly decrease the damage of A. fumigatus swollen conidia (Idol et al. 2022). Additionally, A. fumigatus has evolved an efficient ROS detoxification system which provides protection in high-ROS environments. Cat1 and Cat2 are known catalase peroxidases; a cat1 deletion mutant was found to increase conidia susceptibility to hydrogen peroxide in vitro, and delayed infection is observed in rats treated with double cat1 and cat2 mutant strains (Paris et al. 2003; Shibuya et al. 2006). A study showed that an oxrA deficient strain decreased inflammation, cytokine secretion, and markedly reduced neutrophil influx into the lungs. Furthermore, cat1 or cat2 overexpression rescued phenotypes associated with oxrA deficiency (Zhai et al. 2021). Additionally, superoxide dismutase enzymes, Sod1 (cytoplasmic Cu/ZnSOD), Sod2 (mitochondrial MnSOD), Sod3 (cytoplasmic MnSOD), and Sod4 are important detoxifying superoxide anions. Δsod1 and Δsod2 mutants inhibited hypersensitive growth to menadione, while triple sod1/sod2/sod3 mutants delayed conidial germination and increased AM sensitivity to killing in immunocompetent mice; however, no significant virulence differences were recorded in immunocompromised murine aspergillosis models when compared with wild-type strains (Lambou et al. 2010). A. fumigatus oxidative stress response genes (ppoA, ppoB, and ppoC), non-ribosomal peptide synthetase (pes1), and transcription factors controlling responses to external reactive oxidants (yap1 and skn7) have also been reported for the resisting to the host ROS reaction so as to protect the mold from the host defense (Lamarre et al. 2007; Reeves et al. 2006; Schlam et al. 2016; Tsitsigiannis et al. 2005). Furthermore, human and murine neutrophils release neutrophil extracellular traps (NETs), which eliminate extracellular A. fumigatus; however, this only decreases polar A. fumigatus germ tubes rather than killing the fungi (McCormick et al. 2010). Hence, NET evasion appears to be a strategy permitting pathogen survival and dissemination. Currently, precise A. fumigatus evasion mechanisms from NETs are unclear. However, it is reported that degradation of NETs with the DNases, inhibition of NETs release by down-regulating host inflammatory responses, or withstanding the NETs encapsulation are the main escape strategy for respiratory pathogens, including Bordetella pertussis, Haemophilus influenzae, and group A Streptococcus (Storisteanu et al. 2017). In interactions between A. fumigatus and neutrophils, interacting hyphae may generate new hyphal branches at de novo tips to avoid neutrophil interactions; therefore, increased branch induction may result in the more aggressive of A. fumigatus for the limited number of neutrophils (Ellett et al. 2017).
Monocytes, dendritic cells (DCs), and natural killer (NK) cells are required to control A. fumigatus infections. Conidia germination and hyphal growth may be inhibited by monocytes (Schiefermeier-Mach and Haller 2020). Also, NK cells are directly activated against A. fumigatus hyphae rather than resting conidia, and if NK cells prestimulated by interleukin (1L)-2, high levels of interferon-γ and granulocyte macrophage colony-stimulating factor were produced, so A. fumigatus significantly reduced (Schmidt et al. 2011). NK cells also secrete numerous tumor necrosis factor-α, 1L-18, and galectin-9 molecules to induce macrophage polarization to M1 phenotypes after stimulation by A. fumigatus (Zhang et al. 2019). DCs can phagocytose A. fumigatus conidia and hyphae to stimulate Th1 cytokine (IL-1A, IL-1B, IL-12B, and TNF-α) and Th2 cytokine production (IL-6 and IL-10) (Bozza et al. 2002; Mezger et al. 2008). When frequent or high fungal burdens occur, acquired immunity is activated. T cells, induced by A. fumigatus, protect against invasive aspergillosis (IA) (Hebart et al. 2002). Th2 cells predominate in response to cystic fibrosis (CF) allergic bronchopulmonary aspergillosis (ABPA) patients infected with A. fumigatus (Jat et al. 2021). A. fumigatus secreted proteins may promote Th2 cell activation (Bozza et al. 2009). Currently, there was still little knowledge about immune evasion for A. fumigatus, just to know that after phagocytosis, A. fumigatus conidia rapidly escaped from DCs. And NK cell failed to release full granule, NK cell surface activatory receptors NKG2D and NKp46 were contact-dependent down-regulation (Santiago et al. 2018). Thus, more research is required to identify A. fumigatus escape strategies from host immune defense systems.
Forming special structures which resist host defenses is a common strategy in pathogenic fungi. Biofilms or aspergilloma are major mechanisms which inhibit host defenses and usually contain numerous hyphae and extracellular matrix (ECM) components which contain virulent factors, including β-D-glucan, galactomannan, and other proteins. Virulence factor release activates immune responses, causes mucus plugging, and eosinophilic pneumonia by generating intense inflammatory reactions (Agarwal et al. 2020). If the lungs are damaged via chronic inflammatory and fibrotic processes, A. fumigatus mycelia can successfully grow in abnormal lung mucus, which increases fungal colonization, stimulates Th2-based responses, and favors ABPA development (Kraemer et al. 2006). Fungal biofilms and aspergilloma have significant roles in antifungal resistance (Borghi et al. 2016; Kashyap et al. 2023), while the ECM adsorbs antifungal drugs and prevents their diffusion (Wei et al. 2022). Therefore, fungal cells cannot contact the high concentrations of drugs so they can survive (Wuyts et al. 2018).
Typically, A. fumigatus invasion and host defenses are dynamic and complex processes. The host immune system recognizes distinct A. fumigatus morphological forms to control growth and prevent tissue invasion, whereas fungi require nutrients and must adapt to hostile environments by esca** immune recognition and counteracting host responses. Understanding these highly dynamic interactions is essential to fully understand aspergillosis pathogenesis and facilitate new therapeutic drug design to overcome morbidity and mortality caused by A. fumigatus (Schweer et al. 2014).
Microecological relationships between symbiotic microorganisms in the lung
As the lungs are an open, interconnective environmental system, A. fumigatus and complex microbial mixtures contribute to dynamic microecological homeostasis and are undoubtedly reciprocal in nature (Liu et al. 2021). If a mixed infection, rather than a single infection, occurs, tissue inflammation and damage may be more serious (Neupane et al. 2020). Polymicrobial biofilms are simultaneously formed in the lung, increase antifungal resistance, and are difficult to eradicate using therapeutics (Wang et al. 2020).
Viruses
Aspergillus and viruses are common pathogens, and their inter-microbial interactions occur naturally in clinical settings. Viruses damaging the immune system and infecting the respiratory tract are commonly reported in co-infections with Aspergillus. COVID-19 is a novel virus and induces severe acute respiratory syndrome. A. fumigatus co-infection rates with COVID-19 in intensive care unit patients in the UK, Netherlands, Germany, Italy, and France were 14.1%, 19.4%, 26.0%, 27.7%, and 33.3%, respectively (Arastehfar et al. 2021; White et al. 2021). When treating COVID-19, the long-term and continual administration of high corticosteroid doses probably limited phosphoinositide 3-kinase/Akt signaling, which upregulated proinflammatory cytokines and promoted FleA signaling activation, thereby facilitating Aspergillus spore entry into cells (Banerjee et al. 2021; Steenwyk et al. 2021). Moreover, epithelial lung damage stemming from COVID-19 immunopathology may have facilitated Aspergillus superinfections (Marr et al. 2021). Additionally, post-respiratory viral Th-2 immune responses, mediated by increased IL-10 levels, followed by temporary Th1 immune depression and down-regulated macrophage responses, may have facilitated aspergillosis invasion (Lai and Yu 2021; Tavakoli et al. 2020). However, no significant differences were identified between COVID-19 patients and healthy controls in terms of immune response (Moser et al. 2021).
HIV (human immunodeficiency virus) may cause abnormalities that affect immune system components. IA often occurs in advanced AIDS (acquired immune deficiency syndrome) conditions and mainly affects the respiratory tract (Singh et al. 1991). In the lung, radiological data have shown vascular invasion, pulmonary abscesses, bilateral nodular infiltrates, and cavitary lesions in the upper lobes (Hakkouni and Mansouri 2018). Also, chronic A. fumigatus colonization in CF is accompanied by CXCR4 (HIV coreceptor) accumulation in airway granulocytes (Carevic et al. 2015). The A. fumigatus Asp f1 protein possesses an amino acid domain that resembles the HIV-1 gp41 heptad repeat 2 (Becker 2007). Additionally, influenza is a common respiratory virus, and mice post-influenza PR/8/34 H1N1 and challenged with A. fumigatus had increased fungal and viral burdens, inflammation, and mortality rates. The influenza A-induced signal transducer and activator of transcription 1 molecule inhibited neutrophil recruitment and increased susceptibility to post-influenza IPA (Tobin and Nickolich 2020). Moreover, some viruses occur in co-infections, including Cetacean morbillivirus, parainfluenza virus type 3, cytomegalovirus, and respiratory syncytial virus (Cassle et al. 2016; Hassantoufighi et al. 2007; Lee et al. 2010), and which are summarized in Table 2.
Bacteria
Concomitant colonization by A. fumigatus and bacteria is reportedly widespread in immunocompromised or respiratory disease patients. Co-infection pathogens usually exert synergistic effects and mutual interference which depend on airway microenvironmental factors. Interactions between A. fumigatus and Pseudomonas aeruginosa represent major fungal-bacterial co-operation between pulmonary microbiota (Ostapska et al. 2022). High levels of pyochelin (P. aeruginosa metabolite) can transfer iron to the fungal siderophore triacetylfusarinine C for fungal growth (Briard et al. 2019). Pyochelin and phenazines may also kill A. fumigatus by inducing oxidative and nitrosative stress and iron starvation; however, sub-inhibitory phenazine, pyocyanin, phenazine-1-carboxamide, and phenazine-1-carboxylic acid concentrations can stimulate fungal growth via iron acquisition (Briard et al. 2015, 2019). When iron levels are high, 2-heptyl-3-hydroxy-4-quinolone enhances fungal metabolism and growth. Furthermore, volatile metabolic by-products containing sulfur groups and released by P. aeruginosa benefit fungal growth via interactions with hyphal cell walls (Briard et al. 2016). A. fumigatus and P. aeruginosa biofilms share a similar chemical composition such that both organisms can generate partially cationic de-N-acetylated exopolysaccharides which are important in biofilm formation. Similarly, cationic exopolysaccharide Pel-containing bacterial culture supernatants may augment A. fumigatus biofilm adherence. Similarly, P. aeruginosa adhered to A. fumigatus hyphae in a cationic exopolysaccharide galactosaminogalactan (GAG)-dependent manner on GAG-coated A. fumigatus biofilm coverslips (Ostapska et al. 2022). P. aeruginosa also possesses multiple mechanisms that attenuate NET production and resist NET-mediated killing (Block and Zarbock 2021). Further analyses of P. aeruginosa strains, isolated from CF patients at early and late disease stages, showed that resistance to NET-mediated killing evolved over time. Thus, A. fumigatus and P. aeruginosa co-operation is favorable for fungal evasion. Nevertheless, competition between P. aeruginosa and A. fumigatus has also been reported. Indeed, P. aeruginosa was shown to generate siderophore pyoverdine to induce iron starvation and inhibit A. fumigatus biofilm metabolism; however, A. fumigatus may be self-protected by siderophore production (Sass et al. 2019). In CF, invasive aspergillosis, or chronic obstructive pulmonary disease patients, A. fumigatus co-localization with other pathogens (Streptococcus pneumonia, Staphylococcus aureus, Klebsiella pneumonia, Mycobacterium tuberculosis, and Mycoplasma pneumoniae) has been reported (Iwahashi et al. 2020; Moodley et al. 2014; Nogueira et al. 2019; Peccini et al. 2019; Ramírez Granillo et al. 2015). Most of these pathogens also produce biofilms, and inhibitory interactions have been reported depending on the residing microenvironment. S. pneumoniae and S. aureus are known to disrupt preformed A. fumigatus biofilms. Scanning electron microscopy data have shown that A. fumigatus mycelial networks are fragmented, hypha are markedly reduced, and short and thin abortive hyphae are formed. Moreover, conidia are scarce, and their surfaces and presented lyses have modified, and finally preformed fungal biofilm ECM had disappeared (Iwahashi et al. 2020; Ramírez Granillo et al. 2015). K. pneumoniae renders A. fumigatus sensitive to cell wall stress and upregulates cell wall-related genes (Nogueira et al. 2019). Overall, interactions between pulmonary microbiota are complicated, and more studies are required to explore this phenomenon. Other important pathogens are outlined (Table 2).
Currently, there is limited information on the relationships between A. fumigatus and other fungi or parasites in the lung. Mixed A. fumigatus and Candida albicans infections have been identified from the histopathological examination of patient’s lungs (Nasri and Fakhim 2019). A combined infection with A. fumigatus and the parasite Pneumocystis carinii was also found in a disease case, but no further information about their coinfection (Lee et al. 2010). Detailed A. fumigatus co-infections with other pathogens are described (Table 2). Future studies are required to examine co-operative interactions between A. fumigatus and microbes and show how their interactions facilitate escape from host immune systems in in vitro and in vivo models.
Drug therapy resistance
In immunodeficient or immunocompromised individuals, opportunistic fungal pathogen infections are now primary factors associated with mortality (Jenks et al. 2020). Therefore, new drug therapies must be explored to treat these pathogens. Currently, echinocandins, azoles, and polyenes are the main conventional therapies used to clinically treat A. fumigatus. In contrast, and thanks to widespread clinical-drug use, fungi have evolved several escape mechanisms to withstand antifungal drug damage. For example, Candida auris is a multidrug-resistant fungus and resistant to almost all antifungal agents (Chowdhary et al. 2020).
Antifungal drug use against A. fumigatus in clinical settings
The fungal cell wall provides structural integrity, protection, and physical defenses against adverse environments. More importantly, cell wall is absent in human cells and has been successively targeted using innovative drug design (Zhou et al. 2022). β-1,3-D-glucan synthase is encoded by fks1, is required for cell wall assembly, and has been used as a target for antifungal agents for antifungal agent echinocandins (Satish et al. 2019), Caspofungin, anidulafungin, and micafungin are the representatives of this group (Revie et al. 2018). Caspofungin is fungistatic against A. fumigatus and reduces the increase growth of mold by caspofungin paradoxical effect (CPE) means. The proteins involved in CPE responses contain the basal modulation of the RNA polymerase II initiation sites, calcium metabolism, and cell wall remodeling (Mattos et al. 2020; Valero et al. 2020). Micafungin is an effective prophylactic antifungal agent and has been used in patients with hematological diseases (e.g., acute leukemia) who are at high risk of invasive mold infections (Park et al. 2019; Siopi et al. 2021). The drug significantly up-regulates the conidiophore brlA gene and alters Aspergillus nidulans morphology (Reese et al. 2021). Importantly, antifungal prophylaxis success rates are up to 80% in hematopoietic stem cell transplant recipients when administered micafungin (50 mg/day) (Jarvis et al. 2004). Additionally, anidulafungin therapy increases survival and improves pulmonary infarct scores for pulmonary or disseminated aspergillosis in animal models (Pfaller 2004).
The cell membrane is also an important drug target for protecting the cell interior. Specifically, ergosterol is the most important membrane sterol in fungal cells, but is not found in human cell membranes. Two agents related to ergosterol are widely used in clinical practice: azoles and amphotericin B (AMB) (representative polyene agent). Azoles have excellent antifungal activity and are the only oral drugs with anti-Aspergillus activity. Azoles inhibit the cytochrome P450-dependent enzyme 14α demethylase (Cyp51) to block the channel of lanosterol conversion into ergosterol (Emami et al. 2017). A. fumigatus demonstrated intrinsic resistance to fluconazole, while the triazole antifungals itraconazole, voriconazole, isavuconazole, and posaconazole are favored treatment options against aspergillosis. In an immunosuppressed IPA rabbit model, 40 mg/kg itraconazole had in vivo antifungal activity; however, near-peak itraconazole plasma concentrations varied from 0.5 to 16.8 μg/mL. And the antifungal activity in an inhibitory sigmoid maximum-effect model was strongly correlated with itraconazole plasma concentrations (Berenguer et al. 1994). Previous studies also reported that itraconazole effectively treated ABPA by decreasing immunological severity (Pasqualotto et al. 2009). Voriconazole and posaconazole are effective therapies for asthma in ABPA and severe asthma (Agarwal 2012). AMB binds to sterols in lipid bilayers to form large, extra-membranous aggregates which cause membrane leakage and fungal death (Agarwal 2012). When treated with 0.5 mg/mL AMB, approximately 90% of A. fumigatus protoplast permeability was lost and fungal death ensued (Mousavi and Robson 2004). In clinical settings, AMB administration exhibited dose-limiting toxicity, nephrotoxicity, and infusion-related reactions; therefore, several AMB formulations have been generated: amphotericin B deoxycholate (DAMB), liposomal amphotericin B (LAmB), AmpB lipid complex, and AmpB colloid dispersions (Monk and Goffeau 2008). In 2008, Infectious Diseases Society of America aspergillosis guidelines recommended that LAmB could be used for patients with IA, where voriconazole was not appropriate; thus, a 3 mg/kg/day LAmB dose was advocated (Lanternier and Lortholary 2008). Moreover, monocytes, exposed to a combination of A. fumigatus and DAMB, expressed decreased cytokine (IL-10, IL-2, and IL-3) and up-regulated IL-1β levels (Simitsopoulou et al. 2011). Furthermore, polyene and azole combinations increased survival and demonstrated a significantly greater reduction in tissue burden when compared with monotherapies (Martin-Vicente et al. 2016).
Drug resistance in A. fumigatus
Azole resistance is the most common phenomenon in A. fumigatus. It is reported that patients with chronic pulmonary aspergillosis, when treated with voriconazole, show 5% resistance rates (Bongomin et al. 2018). Also, 11% of itraconazole-treated patients were resistant to A. fumigatus, and PCR data identified A. fumigatus resistance rates of up to 55%, suggesting that increased azole-resistance burdens were limiting factors in aspergillosis treatment (Bongomin et al. 2018; Singh et al. 2020). Mechanistically, cyp51- and non-cyp51-mediated azole resistance actions are implicated in these phenomena. Underlying cyp51-mediated resistance mechanisms are primarily linked to structural changes or the up-regulated azole target lanosterol 14-α-demethylase, especially the cyp51A gene with various of tandem repeat (TR) fragments in the promoter region, causing significant overexpression of the cyp51A. The cyp51-mediated resistance mechanisms mainly contain TR integrations alone or combined with amino acid substitution in the coding gene: TR34/L98H, TR34/L98H/S297T/F495I, TR46/Y121F/T289A, TR53, and TR120 (Garcia-Rubio et al. 2017; Snelders et al. 2008). Another important mechanism underpinning azole resistance is efflux pump gene overexpression which increases multidrug resistance, whereas deletion shows multidrug sensitivity (Meneau et al. 2016). Currently, approximately 49 putative ATP-binding cassette transporters and 278 major facilitator superfamily members have been identified in A. fumigatus genomic sequences. Efflux pumps genes, including abcD, abcE, atrB, atrC, atrF, atrI, cdr1B, mdr1, mdr2, mdr3, and mdr4, are related to azole resistance (Pérez-Cantero et al. 2020; Rivero-Menendez and Alastruey-Izquierdo 2016). Additionally, cell membrane homeostasis, calcium signaling, cell wall integrity, Hsp90-calcineurin pathway, HOG-MAPK signaling, and iron balance are reportedly involved in azole stress responses (Chen et al. 2020).
AMB has broad activity against pathogenic fungi and is associated with lower antifungal resistance rates. AMB-resistant Aspergillus spp. are related to cell wall composition rather than ergosterol content and possess α-1,3-glucan and protein complex alterations in the outermost wall layer (Seo et al. 1999). However, protoplasts and conidia in AMB-resistant A. terreus have no impact on cell walls (Posch et al. 2018). Currently, echinocandin resistance mechanisms in Aspergillus remain under-characterized. For A. fumigatus, the target of echinocandin, glucan synthetase encoding gene fks1 site-directed mutation increased 16-fold resistance to caspofungin. Interestingly, the spontaneous mutants digested cell wall were sensitive to low levels of drug but displayed nearly normal growth above 0.5 μg/ml, and those mutations without significant differences in the chitin and β-1,3-glucan distributions for the mutant and wild-type strains (Gardiner et al. 2005). Together, target gene mutations or alterations in cell wall components are important reasons for the emergence of increased drug resistance to Aspergillus.
Conclusions
In summary, A. fumigatus escape strategies from adverse living conditions are varied and complex. To survive natural environmental factors, including temperature and nutritional stress, and after entering the host for the growing challenge from the immune response, A. fumigatus has evolved several signal receptors, transmitters, effectors, and mutant target proteins to respond to these pressures. Moreover, A. fumigatus hyphae can penetrate lung epithelial tissue, invade blood vessels, and spread through the circulation to other organs (Latgé and Chamilos 2019). Although several therapies have been developed to treat this mold, new drugs are required against drug-resistant strains (Zhang et al. 2021b).
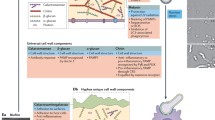
Apart from A. fumigatus, other Aspergillus species, including A. flavus, A. terreus, and A. niger, are common mold infections in humans and cause several serious diseases in both immunocompetent and immunocompromised patients (Stemler et al. 2023). Thus, are there differences between other Aspergillus spp. infections and A. fumigatus processes? How about the negative threats that other Aspergillus spp. encountered in the natural environment and the immune response in host? To answer these questions, future research on fungal epidemiology or secondary infections caused by Aspergillus, especially A. fumigatus, are required (Fig. 1).
The negative threats encountered by A. fumigatus and putative molecules/pathways required for fungal escape. The orange boxes represent the negative threats encountered by A. fumigatus, and the putative molecules/pathways required for fungal escape have been demonstrated in rectangle box text in gray area
Identifying more effective antifungal drug targets and drug repurposing strategies is required to combat A. fumigatus infections. However, antifungal studies have only focused on in vitro outcomes. More in vivo animal model and clinical studies are required to evaluate antifungal effectiveness in vivo (Zhou et al. 2023).
Data Availability
Data sharing is not applicable to this review article as no new data were created in this review article.
References
Abad A, Fernández-Molina JV, Bikandi J, Ramírez A, Margareto J, Sendino J, Hernando FL, Pontón J, Garaizar J, Rementeria A (2010) What makes Aspergillus fumigatus a successful pathogen? Genes and molecules involved in invasive aspergillosis. Rev Iberoam Micol 27(4):155–182. https://doi.org/10.1016/j.riam.2010.10.003
Agarwal R (2012) What is the current place of azoles in allergic bronchopulmonary aspergillosis and severe asthma with fungal sensitization. Expert Rev Respir Med 6(4):363–371. https://doi.org/10.1586/ers.12.35
Agarwal R, Sehgal IS, Dhooria S, Muthu V, Prasad KT, Bal A, Aggarwal AN, Chakrabarti A (2020) Allergic bronchopulmonary aspergillosis. Indian J Med Res 151(6):529–549. https://doi.org/10.4103/ijmr.IJMR_1187_19
Albrecht D, Guthke R, Brakhage AA, Kniemeyer O (2010) Integrative analysis of the heat shock response in Aspergillus fumigatus. BMC Genomics 11:32. https://doi.org/10.1186/1471-2164-11-32
Alonso V, Cavaglieri L, Ramos AJ, Torres A, Marin S (2017) Modelling the effect of pH and water activity in the growth of Aspergillus fumigatus isolated from corn silage. J Appl Microbiol 122(4):1048–1056. https://doi.org/10.1111/jam.13395
Amich J, Calera JA (2014) Zinc acquisition: a key aspect in Aspergillus fumigatus virulence. Mycopathologia 178(5–6):379–385. https://doi.org/10.1007/s11046-014-9764-2
Amich J, Leal F, Calera JA (2009) Repression of the acid ZrfA/ZrfB zinc-uptake system of Aspergillus fumigatus mediated by PacC under neutral zinc-limiting conditions. Int Microbiol 12(1):39–47
Amich J, Vicentefranqueira R, Leal F, Calera JA (2010) Aspergillus fumigatus survival in alkaline and extreme zinc-limiting environments relies on the induction of a zinc homeostasis system encoded by the zrfC and aspf2 genes. Eukaryot Cell 9(3):424–437. https://doi.org/10.1128/ec.00348-09
Amin S, Thywissen A, Heinekamp T, Saluz HP, Brakhage AA (2014) Melanin dependent survival of Apergillus fumigatus conidia in lung epithelial cells. Int J Med Microbiol 304(5–6):626–636. https://doi.org/10.1016/j.ijmm.2014.04.009
Andrianopoulos A, Kourambas S, Sharp JA, Davis MA, Hynes MJ (1998) Characterization of the Aspergillus nidulans nmrA gene involved in nitrogen metabolite repression. J Bacteriol 180(7):1973–1977. https://doi.org/10.1128/jb.180.7.1973-1977.1998
Arastehfar A, Carvalho A, Houbraken J, Lombardi L, Garcia-Rubio R, Jenks JD, Rivero-Menendez O, Aljohani R, Jacobsen ID, Berman J, Osherov N, Hedayati MT, Ilkit M, Armstrong-James D, Gabaldón T, Meletiadis J, Kostrzewa M, Pan W, Lass-Flörl C, Perlin DS, Hoenigl M (2021) Aspergillus fumigatus and aspergillosis: from basics to clinics. Stud Mycol 100:100115. https://doi.org/10.1016/j.simyco.2021.100115
Baldin C, Valiante V, Krüger T, Schafferer L, Haas H, Kniemeyer O, Brakhage AA (2015) Comparative proteomics of a tor inducible Aspergillus fumigatus mutant reveals involvement of the Tor kinase in iron regulation. Proteomics 15(13):2230–2243. https://doi.org/10.1002/pmic.201400584
Banerjee S, Kar A, Mukherjee PK, Haldar PK, Sharma N, Katiyar CK (2021) Immunoprotective potential of Ayurvedic herb Kalmegh (Andrographis paniculata) against respiratory viral infections - LC-MS/MS and network pharmacology analysis. Phytochem Anal 32(4):629–639. https://doi.org/10.1002/pca.3011
Barker BM, Kroll K, Vödisch M, Mazurie A, Kniemeyer O, Cramer RA (2012) Transcriptomic and proteomic analyses of the Aspergillus fumigatus hypoxia response using an oxygen-controlled fermenter. BMC Genomics 13:62. https://doi.org/10.1186/1471-2164-13-62
Beck T, Hall MN (1999) The TOR signalling pathway controls nuclear localization of nutrient-regulated transcription factors. Nature 402(6762):689–692. https://doi.org/10.1038/45287
Becker Y (2007) HIV-1 gp41 heptad repeat 2 (HR2) possesses an amino acid domain that resembles the allergen domain in Aspergillus fumigatus Asp f1 protein: review, hypothesis and implications. Virus Genes 34(3):233–240. https://doi.org/10.1007/s11262-007-0082-3
Ben-Ami R (2023) Systemic antifungal therapy for invasive pulmonary infections. J Fungi 9(2):144. https://doi.org/10.3390/jof9020144
Berenguer J, Ali NM, Allende MC, Lee J, Garrett K, Battaglia S, Piscitelli SC, Rinaldi MG, Pizzo PA, Walsh TJ (1994) Itraconazole for experimental pulmonary aspergillosis: comparison with amphotericin B, interaction with cyclosporin A, and correlation between therapeutic response and itraconazole concentrations in plasma. Antimicrob Agents Chemother 38(6):1303–1308. https://doi.org/10.1128/aac.38.6.1303
Bertuzzi M, Hayes GE, Icheoku UJ, van Rhijn N, Denning DW, Osherov N, Bignell EM (2018) Anti-Aspergillus activities of the respiratory epithelium in health and disease. J Fungi 4(1):8. https://doi.org/10.3390/jof4010008
Bhabhra R, Zhao W, Rhodes JC, Askew DS (2006) Nucleolar localization of Aspergillus fumigatus CgrA is temperature-dependent. Fungal Genet Biol 43(1):1–7. https://doi.org/10.1016/j.fgb.2005.07.005
Blatzer M, Latgé JP (2017) Metal-homeostasis in the pathobiology of the opportunistic human fungal pathogen Aspergillus fumigatus. Curr Opin Microbiol 40:152–159. https://doi.org/10.1016/j.mib.2017.11.015
Blatzer M, Barker BM, Willger SD, Beckmann N, Blosser SJ, Cornish EJ, Mazurie A, Grahl N, Haas H, Cramer RA (2011) SREBP coordinates iron and ergosterol homeostasis to mediate triazole drug and hypoxia responses in the human fungal pathogen Aspergillus fumigatus. PLoS Genet 7(12):e1002374. https://doi.org/10.1371/journal.pgen.1002374
Block H, Zarbock A (2021) A fragile balance: does neutrophil extracellular trap formation drive pulmonary disease progression? Cells 10(8):1932. https://doi.org/10.3390/cells10081932
Bongomin F, Harris C, Hayes G, Kosmidis C, Denning DW (2018) Twelve-month clinical outcomes of 206 patients with chronic pulmonary aspergillosis. PLoS ONE 13(4):17. https://doi.org/10.1371/journal.pone.0193732
Borghi E, Borgo F, Morace G (2016) Fungal biofilms: update on resistance. Adv Exp Med Biol 931:37–47. https://doi.org/10.1007/5584_2016_7
Bozza S, Gaziano R, Spreca A, Bacci A, Montagnoli C, di Francesco P, Romani L (2002) Dendritic cells transport conidia and hyphae of Aspergillus fumigatus from the airways to the draining lymph nodes and initiate disparate Th responses to the fungus. J Immunol 168(3):1362–1371. https://doi.org/10.4049/jimmunol.168.3.1362
Bozza S, Clavaud C, Giovannini G, Fontaine T, Beauvais A, Sarfati J, D’Angelo C, Perruccio K, Bonifazi P, Zagarella S, Moretti S, Bistoni F, Latgé JP, Romani L (2009) Immune sensing of Aspergillus fumigatus proteins, glycolipids, and polysaccharides and the impact on Th immunity and vaccination. J Immunol 183(4):2407–2414. https://doi.org/10.4049/jimmunol.0900961
Briard B, Bomme P, Lechner BE, Mislin GL, Lair V, Prévost MC, Latgé JP, Haas H, Beauvais A (2015) Pseudomonas aeruginosa manipulates redox and iron homeostasis of its microbiota partner Aspergillus fumigatus via phenazines. Sci Rep 5:8220. https://doi.org/10.1038/srep08220
Briard B, Mislin GLA, Latgé JP, Beauvais A (2019) Interactions between Aspergillus fumigatus and pulmonary bacteria: current state of the field, new data, and future perspective. J Fungi 5(2):48. https://doi.org/10.3390/jof5020048
Briard B, Heddergott C, Latgé JP (2016) Volatile compounds emitted by Pseudomonas aeruginosa stimulate growth of the fungal pathogen Aspergillus fumigatus. mBio 7(2):e00219. https://doi.org/10.1128/mBio.00219-16
Buckland KF, O’Connor E, Murray LA, Hogaboam CM (2008) Toll like receptor-2 modulates both innate and adaptive immune responses during chronic fungal asthma in mice. Inflamm Res 57(8):379–387. https://doi.org/10.1007/s00011-008-8004-y
Cabaret O, Bonnal C, Canoui-Poitrine F, Emirian A, Bizouard G, Levesque E, Maitre B, Fihman V, Decousser JW, Botterel F (2016) Concomitant presence of Aspergillus fumigatus and Stenotrophomonas maltophilia in the respiratory tract: a new risk for patients with liver disease? J Med Microbiol 65(5):414–419. https://doi.org/10.1099/jmm.0.000233
Cai Z, Du W, Zhang Z, Guan L, Zeng Q, Chai Y, Dai C, Lu L (2018) The Aspergillus fumigatus transcription factor AceA is involved not only in Cu but also in Zn detoxification through regulating transporters CrpA and ZrcA. Cell Microbiol 20(10):e12864. https://doi.org/10.1111/cmi.12864
Carevic M, Singh A, Rieber N, Eickmeier O, Griese M, Hector A, Hartl D (2015) CXCR4+ granulocytes reflect fungal cystic fibrosis lung disease. Eur Respir J 46(2):395–404. https://doi.org/10.1183/09031936.00173514
Cassle SE, Landrau-Giovannetti N, Farina LL, Leone A, Wellehan JF Jr, Stacy NI, Thompson P, Herring H, Mase-Guthrie B, Blas-Machado U, Saliki JT, Walsh MT, Waltzek TB (2016) Coinfection by Cetacean morbillivirus and Aspergillus fumigatus in a juvenile bottlenose dolphin (Tursiops truncatus) in the Gulf of Mexico. J Vet Diagn Invest 28(6):729–734. https://doi.org/10.1177/1040638716664761
Chang YC, Tsai HF, Karos M, Kwon-Chung KJ (2004) THTA, a thermotolerance gene of Aspergillus fumigatus. Fungal Genet Biol 41(9):888–896. https://doi.org/10.1016/j.fgb.2004.06.004
Chen P, Liu J, Zeng M, Sang H (2020) Exploring the molecular mechanism of azole resistance in Aspergillus fumigatus. J Mycol Med 30(1):100915. https://doi.org/10.1016/j.mycmed.2019.100915
Chen Y, Ai L, Zhou Y, Zhao Y, Huang J, Tang W, Liang Y (2021) Rapid and precise diagnosis of pneumonia coinfected by Pneumocystis jirovecii and Aspergillus fumigatus assisted by next-generation sequencing in a patient with systemic lupus erythematosus: a case report. Ann Clin Microbiol Antimicrob 20(1):47. https://doi.org/10.1186/s12941-021-00448-5
Chowdhary A, Tarai B, Singh A, Sharma A (2020) Multidrug-resistant Candida auris infections in critically ill coronavirus disease patients, India, April-July 2020. Emerg Infect Dis 26(11):2694–2696. https://doi.org/10.3201/eid2611.203504
Crossen AJ, Ward RA (2022) Human airway epithelium responses to invasive fungal infections: a critical partner in innate immunity. J Fungi 9(1):40. https://doi.org/10.3390/jof9010040
Crunden JL, Diezmann S (2021) Hsp90 interaction networks in fungi-tools and techniques. FEMS Yeast Res 21(7):foab054. https://doi.org/10.1093/femsyr/foab054
Dasari P, Shopova IA, Stroe M, Wartenberg D, Martin-Dahse H, Beyersdorf N, Hortschansky P, Dietrich S, Cseresnyés Z, Figge MT, Westermann M, Skerka C, Brakhage AA, Zipfel PF (2018) Aspf2 from Aspergillus fumigatus recruits human immune regulators for immune evasion and cell damage. Front Immunol 9:1635. https://doi.org/10.3389/fimmu.2018.01635
Dichtl K, Helmschrott C, Dirr F, Wagener J (2012) Deciphering cell wall integrity signalling in Aspergillus fumigatus: identification and functional characterization of cell wall stress sensors and relevant Rho GTPases. Mol Microbiol 83(3):506–519. https://doi.org/10.1111/j.1365-2958.2011.07946.x
Dietl AM, Binder U, Shadkchan Y, Osherov N, Haas H (2018) Siroheme is essential for assimilation of nitrate and sulfate as well as detoxification of nitric oxide but dispensable for murine virulence of Aspergillus fumigatus. Front Microbiol 9:2615. https://doi.org/10.3389/fmicb.2018.02615
Dirr F, Echtenacher B, Heesemann J, Hoffmann P, Ebel F, Wagener J (2010) AfMkk2 is required for cell wall integrity signaling, adhesion, and full virulence of the human pathogen Aspergillus fumigatus. Int J Med Microbiol 300(7):496–502. https://doi.org/10.1016/j.ijmm.2010.03.001
Downes DJ, Davis MA, Wong KH, Kreutzberger SD, Hynes MJ, Todd RB (2014) Dual DNA binding and coactivator functions of Aspergillus nidulans TamA, a Zn(II)2Cys6 transcription factor. Mol Microbiol 92(6):1198–1211. https://doi.org/10.1111/mmi.12620
Earl Kang S, Celia BN (2021) Sporulation environment drives phenotypic variation in the pathogen Aspergillus fumigatus. G3 (Bethesda) 11(8):jkab208. https://doi.org/10.1093/g3journal/jkab208
Ellett F, Jorgensen J, Frydman GH, Jones CN, Irimia D (2017) Neutrophil interactions stimulate evasive hyphal branching by Aspergillus fumigatus. PLoS Pathog 13(1):e1006154. https://doi.org/10.1371/journal.ppat.1006154
Emami S, Tavangar P, Keighobadi M (2017) An overview of azoles targeting sterol 14α-demethylase for antileishmanial therapy. Eur J Med Chem 135:241–259. https://doi.org/10.1016/j.ejmech.2017.04.044
Fabri J, Rocha MC, Fernandes CM, Persinoti GF, Ries LNA, da Cunha AF, Goldman GH, Del Poeta M, Malavazi I (2021) The heat shock transcription factor HsfA is essential for thermotolerance and regulates cell wall integrity in Aspergillus fumigatus. Front Microbiol 12:656548. https://doi.org/10.3389/fmicb.2021.656548
Furlong-Silva J, Cook PC (2022) Fungal-mediated lung allergic airway disease: the critical role of macrophages and dendritic cells. PLoS Pathog 18(7):e1010608. https://doi.org/10.1371/journal.ppat.1010608
Gago S, Denning DW, Bowyer P (2019) Pathophysiological aspects of Aspergillus colonization in disease. Med Mycol 57(Supplement_2):S219–S227. https://doi.org/10.1093/mmy/myy076
Garcia-Rubio R, Cuenca-Estrella M, Mellado E (2017) Triazole resistance in Aspergillus species: an emerging problem. Drugs 77(6):599–613. https://doi.org/10.1007/s40265-017-0714-4
Gardiner RE, Souteropoulos P, Park S, Perlin DS (2005) Characterization of Aspergillus fumigatus mutants with reduced susceptibility to caspofungin. Med Mycol 43(Suppl 1):S299–S305. https://doi.org/10.1080/13693780400029023
Giacobbe DR, Prattes J, Wauters J, Dettori S, Signori A, Salmanton-García J, Maertens J, Bourgeois M, Reynders M, Rutsaert L, Van Regenmortel N, Lormans P, Feys S (2022) Prognostic impact of bronchoalveolar lavage fluid galactomannan and Aspergillus culture results on survival in COVID-19 intensive care unit patients: a post hoc analysis from the European confederation of medical mycology (ECMM) COVID-19-associated pulmonary aspergillosis study. J Clin Microbiol 60(4):e0229821. https://doi.org/10.1128/jcm.02298-21
Graf KT, Liu H, Filler SG (2023) Depletion of extracellular chemokines by Aspergillus melanin. mBio e0019423. https://doi.org/10.1128/mbio.00194-23
Grahl N, Puttikamonkul S, Macdonald JM, Gamcsik MP, Ngo LY, Hohl TM, Cramer RA (2011) In vivo hypoxia and a fungal alcohol dehydrogenase influence the pathogenesis of invasive pulmonary aspergillosis. PLoS Pathog 7(7):e1002145. https://doi.org/10.1371/journal.ppat.1002145
Grahl N, Dinamarco TM, Willger SD, Goldman GH, Cramer RA (2012) Aspergillus fumigatus mitochondrial electron transport chain mediates oxidative stress homeostasis, hypoxia responses and fungal pathogenesis. Mol Microbiol 84(2):383–399. https://doi.org/10.1111/j.1365-2958.2012.08034.x
Gsaller F, Eisendle M, Lechner BE, Schrettl M, Lindner H, Müller D, Geley S, Haas H (2012) The interplay between vacuolar and siderophore-mediated iron storage in Aspergillus fumigatus. Metallomics 4(12):1262–1270. https://doi.org/10.1039/c2mt20179h
Hakkouni AE, Mansouri N (2018) Invasive pulmonary aspergillosis in a patient with human immunodeficiency virus (HIV). Pan Afr Med J 31:40. https://doi.org/10.11604/pamj.2018.31.40.16637
Hassantoufighi A, Oglesbee M, Richter BW, Prince GA, Hemming V, Niewiesk S, Eichelberger MC (2007) Respiratory syncytial virus replication is prolonged by a concomitant allergic response. Clin Exp Immunol 148(2):218–229. https://doi.org/10.1111/j.1365-2249.2007.03341.x
Hebart H, Bollinger C, Fisch P, Sarfati J, Meisner C, Baur M, Loeffler J, Monod M, Latgé JP, Einsele H (2002) Analysis of T-cell responses to Aspergillus fumigatus antigens in healthy individuals and patients with hematologic malignancies. Blood 100(13):4521–4528. https://doi.org/10.1182/blood-2002-01-0265
Henriet SS, Hermans PW, Verweij PE, Simonetti E, Holland SM, Sugui JA, Kwon-Chung KJ, Warris A (2011) Human leukocytes kill Aspergillus nidulans by reactive oxygen species-independent mechanisms. Infect Immun 79(2):767–773. https://doi.org/10.1128/iai.00921-10
Hervás R, Oroz J (2020) Mechanistic insights into the role of molecular chaperones in protein misfolding diseases: from molecular recognition to amyloid disassembly. Int J Mol Sci 21(23):9186. https://doi.org/10.3390/ijms21239186
Hoenigl M (2021) Invasive fungal disease complicating coronavirus disease 2019: when it rains, it spores. Clin Infect Dis 73(7):e1645–e1648. https://doi.org/10.1093/cid/ciaa1342
Hokken MWJ, Coolen JPM, Steenbreker H, Zoll J, Baltussen TJH, Verweij PE, Melchers WJG (2023) The transcriptome response to azole compounds in Aspergillus fumigatus shows differential gene expression across pathways essential for azole resistance and cell survival. J Fungi 9(8):807. https://doi.org/10.3390/jof9080807
Idol RA, Bhattacharya S, Huang G, Song Z, Huttenlocher A, Keller NP, Dinauer MC (2022) Neutrophil and macrophage NADPH oxidase 2 differentially control responses to inflammation and to Aspergillus fumigatus in mice. J Immunol 209(10):1960–1972. https://doi.org/10.4049/jimmunol.2200543
Iwahashi J, Kamei K, Watanabe H (2020) Disruption of Aspergillus fumigatus biofilm by Streptococcus pneumoniae: mycelial fragmentation by hydrogen peroxide. J Infect Chemother 26(8):831–837. https://doi.org/10.1016/j.jiac.2020.03.015
Jarvis B, Figgitt DP, Scott LJ (2004) Micafungin Drugs 64:969–982. https://doi.org/10.2165/00003495-200464090-00004
Jat KR, Walia DK, Khairwa A (2021) Anti-IgE therapy for allergic bronchopulmonary aspergillosis in people with cystic fibrosis. Cochrane Database Syst Rev 9(9):Cd010288. https://doi.org/10.1002/14651858.CD010288.pub5
Jenks JD, Cornely OA, Chen SC, Thompson GR 3rd, Hoenigl M (2020) Breakthrough invasive fungal infections: who is at risk? Mycoses 63(10):1021–1032. https://doi.org/10.1111/myc.13148
Jung MG, Kim SS, Yu JH, Shin KS (2016) Characterization of gprK encoding a putative hybrid G-protein-coupled receptor in Aspergillus fumigatus. PLoS ONE 11(9):e0161312. https://doi.org/10.1371/journal.pone.0161312
Kashyap VH, Mishra A, Bordoloi S, Varma A, Joshi NC (2023) Exploring the intersection of Aspergillus fumigatus biofilms, infections, immune response and antifungal resistance. Mycoses 66(9):737–754. https://doi.org/10.1111/myc.13619
Kim Y, Heo IB, Yu JH, Shin KS (2017) Characteristics of a regulator of G-protein signaling (RGS) rgsC in Aspergillus fumigatus. Front Microbiol 8:2058. https://doi.org/10.3389/fmicb.2017.02058
Korfanty GA, Dixon M, Jia H, Yoell H, Xu J (2021) Genetic diversity and dispersal of Aspergillus fumigatus in Arctic soils. Genes 13(1):19. https://doi.org/10.3390/genes13010019
Kraemer R, Deloséa N, Ballinari P, Gallati S, Crameri R (2006) Effect of allergic bronchopulmonary aspergillosis on lung function in children with cystic fibrosis. Am J Respir Crit Care Med 174(11):1211–1220. https://doi.org/10.1164/rccm.200603-423OC
Krappmann S, Bignell EM, Reichard U, Rogers T, Haynes K, Braus GH (2004) The Aspergillus fumigatus transcriptional activator CpcA contributes significantly to the virulence of this fungal pathogen. Mol Microbiol 52(3):785–799. https://doi.org/10.1111/j.1365-2958.2004.04015.x
Kumar V (2022) Toll-Like receptors in adaptive immunity. Handb Exp Pharmacol 276:95–131. https://doi.org/10.1007/164_2021_543
Lai CC, Yu WL (2021) COVID-19 associated with pulmonary aspergillosis: a literature review. J Microbiol Immunol Infect 54(1):46–53. https://doi.org/10.1016/j.jmii.2020.09.004
Lamarre C, Ibrahim-Granet O, Du C, Calderone R, Latgé JP (2007) Characterization of the SKN7 ortholog of Aspergillus fumigatus. Fungal Genet Biol 44(7):682–690. https://doi.org/10.1016/j.fgb.2007.01.009
Lambou K, Lamarre C, Beau R, Dufour N, Latge JP (2010) Functional analysis of the superoxide dismutase family in Aspergillus fumigatus. Mol Microbiol 75(4):910–923. https://doi.org/10.1111/j.1365-2958.2009.07024.x
Lanternier F, Lortholary O (2008) Liposomal amphotericin B: what is its role in 2008? Clin Microbiol Infect 14(Suppl 4):71–83. https://doi.org/10.1111/j.1469-0691.2008.01984.x
Latgé JP, Chamilos G (2019) Aspergillus fumigatus and aspergillosis in 2019. Clin Microbiol Rev 33(1):e00140-e218. https://doi.org/10.1128/cmr.00140-18
Lee T, Bae YJ, Park SK, Park HJ, Kim SH, Cho YS, Moon HB, Lee SO, Kim TB (2010) Severe pneumonia caused by combined infection with Pneumocystis jiroveci, parainfluenza virus type 3, cytomegalovirus, and Aspergillus fumigatus in a patient with Stevens-Johnson syndrome/toxic epidermal necrolysis. Acta Derm Venereol 90(6):625–629. https://doi.org/10.2340/00015555-0977
Lin CJ, Hou YH, Chen YL (2020) The histone acetyltransferase GcnE regulates conidiation and biofilm formation in Aspergillus fumigatus. Med Mycol 58(2):248–259. https://doi.org/10.1093/mmy/myz043
Liu H, Lee MJ, Solis NV, Phan QT, Swidergall M, Ralph B, Ibrahim AS, Sheppard DC, Filler SG (2016) Aspergillus fumigatus CalA binds to integrin α5β1 and mediates host cell invasion. Nat Microbiol 2:16211. https://doi.org/10.1038/nmicrobiol.2016.211
Liu J, Ran Z, Wang F, **n C, **ong B, Song Z (2021) Role of pulmonary microorganisms in the development of chronic obstructive pulmonary disease. Crit Rev Microbiol 47(1):1–12. https://doi.org/10.1080/1040841x.2020.1830748
Long N, Orasch T (2018) The Zn2Cys6-type transcription factor LeuB cross-links regulation of leucine biosynthesis and iron acquisition in Aspergillus fumigatus. PLoS Genet 14(10):e1007762. https://doi.org/10.1371/journal.pgen.1007762
Ma D, Li R (2013) Current understanding of HOG-MAPK pathway in Aspergillus fumigatus. Mycopathologia 175(1–2):13–23. https://doi.org/10.1007/s11046-012-9600-5
Macios M, Caddick MX, Weglenski P, Scazzocchio C, Dzikowska A (2012) The GATA factors AREA and AREB together with the co-repressor NMRA, negatively regulate arginine catabolism in Aspergillus nidulans in response to nitrogen and carbon source. Fungal Genet Biol 49(3):189–198. https://doi.org/10.1016/j.fgb.2012.01.004
Margalit A, Kavanagh K (2015) The innate immune response to Aspergillus fumigatus at the alveolar surface. FEMS Microbiol Rev 39(5):670–687. https://doi.org/10.1093/femsre/fuv018
Marr KA, Platt A, Tornheim JA, Zhang SX, Datta K, Cardozo C, Garcia-Vidal C (2021) Aspergillosis complicating severe coronavirus disease. Emerg Infect Dis 27(1):18–25. https://doi.org/10.3201/eid2701.202896
Martin-Vicente A, Capilla J, Guarro J (2016) In vivo synergy of amphotericin B plus posaconazole in murine aspergillosis. Antimicrob Agents Chemother 60(1):296–300. https://doi.org/10.1128/aac.01462-15
Matthaiou EI, Sass G, Stevens DA, Hsu JL (2018) Iron: an essential nutrient for Aspergillus fumigatus and a fulcrum for pathogenesis. Curr Opin Infect Dis 31(6):506–511. https://doi.org/10.1097/qco.0000000000000487
Mattos EC, Palmisano G, Goldman GH (2020) Phosphoproteomics of Aspergillus fumigatus exposed to the antifungal drug caspofungin. mSphere 5(3):e00365–20. https://doi.org/10.1128/mSphere.00365-20
McCormick A, Heesemann L, Wagener J, Marcos V, Hartl D, Loeffler J, Heesemann J, Ebel F (2010) NETs formed by human neutrophils inhibit growth of the pathogenic mold Aspergillus fumigatus. Microbes Infect 12(12–13):928–936. https://doi.org/10.1016/j.micinf.2010.06.009
McCormick A, Jacobsen ID, Broniszewska M, Beck J, Heesemann J, Ebel F (2012) The two-component sensor kinase TcsC and its role in stress resistance of the human-pathogenic mold Aspergillus fumigatus. PLoS ONE 7(6):e38262. https://doi.org/10.1371/journal.pone.0038262
Meneau I, Coste AT, Sanglard D (2016) Identification of Aspergillus fumigatus multidrug transporter genes and their potential involvement in antifungal resistance. Med Mycol 54(6):616–627. https://doi.org/10.1093/mmy/myw005
Mezger M, Kneitz S, Wozniok I, Kurzai O, Einsele H, Loeffler J (2008) Proinflammatory response of immature human dendritic cells is mediated by dectin-1 after exposure to Aspergillus fumigatus germ tubes. J Infect Dis 197(6):924–931. https://doi.org/10.1086/528694
Monin L, Mehta S, Elsegeiny W (2018) Aspergillus fumigatus preexposure worsens pathology and improves control of Mycobacterium abscessus pulmonary infection in mice. Infect Immun 86(3):e00859-e917. https://doi.org/10.1128/iai.00859-17
Monk BC, Goffeau A (2008) Outwitting multidrug resistance to antifungals. Science 321(5887):367–369. https://doi.org/10.1126/science.1159746
Moodley L, Pillay J, Dheda K (2014) Aspergilloma and the surgeon. J Thorac Dis 6(3):202–209. https://doi.org/10.3978/j.issn.2072-1439.2013.12.40
Moser D, Biere K, Han B, Hoerl M, Schelling G, Choukér A, Woehrle T (2021) COVID-19 impairs immune response to Candida albicans. Front Immunol 12:640644. https://doi.org/10.3389/fimmu.2021.640644
Mousavi SAA, Robson GD (2004) Oxidative and amphotericin B-mediated cell death in the opportunistic pathogen Aspergillus fumigatus is associated with an apoptotic-like phenotype. Microbiology 150(Pt 6):1937–1945. https://doi.org/10.1099/mic.0.26830-0
Nasri E, Fakhim H (2019) Airway colonisation by Candida and Aspergillus species in Iranian cystic fibrosis patients. Mycoses 62(5):434–440. https://doi.org/10.1111/myc.12898
Netea MG, Warris A, Van der Meer JW, Fenton MJ, Verver-Janssen TJ, Jacobs LE, Andresen T, Verweij PE, Kullberg BJ (2003) Aspergillus fumigatus evades immune recognition during germination through loss of toll-like receptor-4-mediated signal transduction. J Infect Dis 188(2):320–326. https://doi.org/10.1086/376456
Neupane AS, Willson M, Chojnacki AK, Vargas ESCF, Morehouse C, Carestia A, Keller AE, Peiseler M, DiGiandomenico A, Kelly MM, Amrein M, Jenne C, Thanabalasuriar A, Kubes P (2020) Patrolling alveolar macrophages conceal bacteria from the immune system to maintain homeostasis. Cell 183(1):110-125.e11. https://doi.org/10.1016/j.cell.2020.08.020
Nogueira MF, Pereira L, Jenull S, Kuchler K, Lion T (2019) Klebsiella pneumoniae prevents spore germination and hyphal development of Aspergillus species. Sci Rep 9(1):218. https://doi.org/10.1038/s41598-018-36524-8
Oliveira BR, Marques AP, Ressurreição M, Moreira CJS, Pereira CS, Crespo MTB, Pereir VJ (2021) Inactivation of Aspergillus species in real water matrices using medium pressure mercury lamps. J Photochem Photobiol B 221:112242. https://doi.org/10.1016/j.jphotobiol.2021.112242
Orasch T, Dietl AM, Shadkchan Y, Binder U, Bauer I (2019) The leucine biosynthetic pathway is crucial for adaptation to iron starvation and virulence in Aspergillus fumigatus. Virulence 10(1):925–934. https://doi.org/10.1080/21505594.2019.1682760
Ortiz SC, Pennington K, Thomson DD, Bertuzzi M (2022) Novel insights into Aspergillus fumigatus pathogenesis and host response from state-of-the-art imaging of host-pathogen interactions during infection. J Fungi 8(3):264. https://doi.org/10.3390/jof8030264
Ostapska H, Le Mauff F, Gravelat FN, Snarr BD (2022) Co-operative biofilm interactions between Aspergillus fumigatus and Pseudomonas aeruginosa through secreted galactosaminogalactan exopolysaccharide. J Fungi 8(4):336. https://doi.org/10.3390/jof8040336
Panepinto JC, Oliver BG, Fortwendel JR, Smith DL, Askew DS, Rhodes JC (2003) Deletion of the Aspergillus fumigatus gene encoding the Ras-related protein RhbA reduces virulence in a model of invasive pulmonary aspergillosis. Infect Immun 71(5):2819–2826. https://doi.org/10.1128/iai.71.5.2819-2826.2003
Paris S, Wysong D, Debeaupuis JP, Shibuya K, Philippe B, Diamond RD, Latgé JP (2003) Catalases of Aspergillus fumigatus. Infect Immun 71(6):3551–3562. https://doi.org/10.1128/iai.71.6.3551-3562.2003
Park H, Youk J, Shin DY, Hong J, Kim I, Kim NJ, Lee JO, Bang SM, Yoon SS, Park WB, Koh Y (2019) Micafungin prophylaxis for acute leukemia patients undergoing induction chemotherapy. BMC Cancer 19(1):358. https://doi.org/10.1186/s12885-019-5557-9
Pasqualotto AC, Powell G, Niven R, Denning DW (2009) The effects of antifungal therapy on severe asthma with fungal sensitization and allergic bronchopulmonary aspergillosis. Respirology 14(8):1121–1127. https://doi.org/10.1111/j.1440-1843.2009.01640.x
Peccini L, Pennoni S, Mencarini V, Saponara M, Palladino N, Principi N, Pennoni G, Esposito S (2019) A peculiar case of pneumonia due to Mycoplasma pneumoniae in a child with cystic fibrosis and sensibilization to Aspergillus fumigatus. Pathogens 9(1):15. https://doi.org/10.3390/pathogens9010015
Percopo CM, Dyer KD, Ochkur SI, Luo JL, Fischer ER, Lee JJ, Lee NA, Domachowske JB, Rosenberg HF (2014) Activated mouse eosinophils protect against lethal respiratory virus infection. Blood 123(5):743–752. https://doi.org/10.1182/blood-2013-05-502443
Pérez-Cantero A, López-Fernández L, Guarro J, Capilla J (2020) Azole resistance mechanisms in Aspergillus: update and recent advances. Int J Antimicrob Agents 55(1):105807. https://doi.org/10.1016/j.ijantimicag.2019.09.011
Perez-Cuesta U, Guruceaga X (2021) Nitrogen, iron and zinc acquisition: key nutrients to Aspergillus fumigatus virulence. J Fungi 7(7):518. https://doi.org/10.3390/jof7070518
Pfaller MA (2004) Anidulafungin: an echinocandin antifungal. Expert Opin Investig Drugs 13(9):1183–1197. https://doi.org/10.1517/13543784.13.9.1183
Pongpom M, Liu H, Xu W, Snarr BD, Sheppard DC, Mitchell AP, Filler SG (2015) Divergent targets of Aspergillus fumigatus AcuK and AcuM transcription factors during growth in vitro versus invasive disease. Infect Immun 83(3):923–933. https://doi.org/10.1128/iai.02685-14
Posch W, Blatzer M, Wilflingseder D, Lass-Flörl C (2018) Aspergillus terreus: novel lessons learned on amphotericin B resistance. Med Mycol 56(suppl_1):73–82. https://doi.org/10.1093/mmy/myx119
Qadri H, Qureshi MF, Mir MA, Shah AH (2021) Glucose - The X factor for the survival of human fungal pathogens and disease progression in the host. Microbiol Res 247:126725. https://doi.org/10.1016/j.micres.2021.126725
Ramírez Granillo A, Canales MG, Espíndola ME, Martínez Rivera MA, de Lucio VM, Tovar AV (2015) Antibiosis interaction of Staphylococccus aureus on Aspergillus fumigatus assessed in vitro by mixed biofilm formation. BMC Microbiol 15:33. https://doi.org/10.1186/s12866-015-0363-2
Reese S, Chelius C, Riekhof W, Marten MR, Harris SD (2021) Micafungin-induced cell wall damage stimulates morphological changes consistent with microcycle conidiation in Aspergillus nidulans. J Fungi 7(7):525. https://doi.org/10.3390/jof7070525
Reeves EP, Reiber K, Neville C, Scheibner O, Kavanagh K, Doyle S (2006) A nonribosomal peptide synthetase (Pes1) confers protection against oxidative stress in Aspergillus fumigatus. FEBS J 273(13):3038–3053. https://doi.org/10.1111/j.1742-4658.2006.05315.x
Revie NM, Iyer KR, Robbins N, Cowen LE (2018) Antifungal drug resistance: evolution, mechanisms and impact. Curr Opin Microbiol 45:70–76. https://doi.org/10.1016/j.mib.2018.02.005
Richard N, Marti L, Varrot A, Guillot L (2018) Human bronchial epithelial cells inhibit Aspergillus fumigatus germination of extracellular conidia via FleA recognition. Sci Rep 8(1):15699. https://doi.org/10.1038/s41598-018-33902-0
Ries LNA, Beattie S, Cramer RA, Goldman GH (2018) Overview of carbon and nitrogen catabolite metabolism in the virulence of human pathogenic fungi. Mol Microbiol 107(3):277–297. https://doi.org/10.1111/mmi.13887
Ries LNA, Alves de Castro P, Pereira Silva L, Valero C, Dos Reis TF, Saborano R, Duarte IF, Persinoti GF, Steenwyk JL, Rokas A, Almeida F, Costa JH, Fill T, Sze Wah Wong S, Aimanianda V, Rodrigues FJS, Gonçales RA, Duarte-Oliveira C, Carvalho A, Goldman GH (2021) Aspergillus fumigatus acetate utilization impacts virulence traits and pathogenicity. mBio 12(4):e0168221. https://doi.org/10.1128/mBio.01682-21
Rivero-Menendez O, Alastruey-Izquierdo A (2016) Triazole resistance in Aspergillus spp.: a worldwide problem? J Fungi 2(3):21 https://doi.org/10.3390/jof2030021
Robinson JR, Isikhuemhen OS, Anike FN (2021) Fungal-metal interactions: a review of toxicity and homeostasis. J Fungi 7(3):225. https://doi.org/10.3390/jof7030225
Rocha MC, Minari K (2021) Aspergillus fumigatus Hsp90 interacts with the main components of the cell wall integrity pathway and cooperates in heat shock and cell wall stress adaptation. Cell Microbiol 23(2):e13273. https://doi.org/10.1111/cmi.13273
Rocha MC, Godoy KF, de Castro PA, Hori JI, Bom VL, Brown NA, Cunha AF, Goldman GH, Malavazi I (2015) The Aspergillus fumigatus pkcAG579R mutant is defective in the activation of the cell wall integrity pathway but is dispensable for virulence in a neutropenic mouse infection model. PLoS ONE 10(8):e0135195. https://doi.org/10.1371/journal.pone.0135195
Rocha MC, Fabri JH, Franco de Godoy K, Alves de Castro P, Hori JI, Ferreira da Cunha A, Arentshorst M, Ram AF, van den Hondel CA, Goldman GH, Malavazi I (2016) Aspergillus fumigatus MADS-Box transcription factor rlmA is required for regulation of the cell wall integrity and virulence. G3 (Bethesda) 6(9):2983–3002. https://doi.org/10.1534/g3.116.031112
Samalova M, Flamant P, Beau R, Bromley M, Moya-Nilges M, Fontaine T, Latgé JP, Mouyna I (2023) The new GPI-anchored protein, SwgA, is involved in nitrogen metabolism in the pathogenic filamentous fungus Aspergillus fumigatus. J Fungi 9(2):256. https://doi.org/10.3390/jof9020256
Santiago V, Rezvani K, Sekine T, Stebbing J, Kelleher P, Armstrong-James D (2018) Human NK cells develop an exhaustion phenotype during polar degranulation at the Aspergillus fumigatus hyphal synapse. Front Immunol 9:2344. https://doi.org/10.3389/fimmu.2018.02344
Sass G, Nazik H, Penner J, Shah H, Ansari SR, Clemons KV, Groleau MC, Dietl AM, Visca P, Haas H, Déziel E, Stevens DA (2019) Aspergillus-Pseudomonas interaction, relevant to competition in airways. Med Mycol 57(Supplement_2):S228–S232. https://doi.org/10.1093/mmy/myy087
Sasse C, Bignell EM, Hasenberg M, Haynes K, Gunzer M, Braus GH, Krappmann S (2008) Basal expression of the Aspergillus fumigatus transcriptional activator CpcA is sufficient to support pulmonary aspergillosis. Fungal Genet Biol 45(5):693–704. https://doi.org/10.1016/j.fgb.2007.12.008
Satish S, Jiménez-Ortigosa C, Zhao Y, Lee MH, Dolgov E, Krüger T, Park S, Denning DW, Kniemeyer O (2019) Stress-induced changes in the lipid microenvironment of β-(1,3)-d-glucan synthase cause clinically important echinocandin resistance in Aspergillus fumigatus. mBio 10(3):e00779–19. https://doi.org/10.1128/mBio.00779-19
Schafferer L, Beckmann N, Binder U, Brosch G, Haas H (2015) AmcA-a putative mitochondrial ornithine transporter supporting fungal siderophore biosynthesis. Front Microbiol 6:252. https://doi.org/10.3389/fmicb.2015.00252
Schiefermeier-Mach N, Haller T (2020) Migrating lung monocytes internalize and inhibit growth of Aspergillus fumigatus Conidia. Pathogens 9(12):983. https://doi.org/10.3390/pathogens9120983
Schlam D, Canton J, Carreño M, Kopinski H, Freeman SA, Grinstein S, Fairn GD (2016) Gliotoxin suppresses macrophage immune function by subverting phosphatidylinositol 3,4,5-trisphosphate homeostasis. mBio 7(2):e02242. https://doi.org/10.1128/mBio.02242-15
Schmidt S, Tramsen L, Hanisch M, Latgé JP, Huenecke S, Koehl U, Lehrnbecher T (2011) Human natural killer cells exhibit direct activity against Aspergillus fumigatus hyphae, but not against resting conidia. J Infect Dis 203(3):430–435. https://doi.org/10.1093/infdis/jiq062
Schrettl M, Bignell E, Kragl C, Sabiha Y, Loss O, Eisendle M, Wallner A, Arst HN Jr, Haynes K, Haas H (2007) Distinct roles for intra- and extracellular siderophores during Aspergillus fumigatus infection. PLoS Pathog 3(9):1195–1207. https://doi.org/10.1371/journal.ppat.0030128
Schrettl M, Kim HS, Eisendle M, Kragl C, Nierman WC, Heinekamp T, Werner ER, Jacobsen I, Illmer P, Yi H, Brakhage AA, Haas H (2008) SreA-mediated iron regulation in Aspergillus fumigatus. Mol Microbiol 70(1):27–43. https://doi.org/10.1111/j.1365-2958.2008.06376.x
Schrettl M, Beckmann N, Varga J, Heinekamp T, Jacobsen ID, Jöchl C, Moussa TA, Wang S, Gsaller F, Blatzer M, Werner ER, Niermann WC, Brakhage AA, Haas H (2010) HapX-mediated adaption to iron starvation is crucial for virulence of Aspergillus fumigatus. PLoS Pathog 6(9):e1001124. https://doi.org/10.1371/journal.ppat.1001124
Schweer KE, Bangard C, Hekmat K, Cornely OA (2014) Chronic pulmonary aspergillosis. Mycoses 57(5):257–270. https://doi.org/10.1111/myc.12152
Seidel C, Moreno-Velásquez SD, Ben-Ghazzi N, Gago S, Read ND, Bowyer P (2020) Phagolysosomal survival enables non-lytic hyphal escape and ramification through lung epithelium during Aspergillus fumigatus infection. Front Microbiol 11:1955. https://doi.org/10.3389/fmicb.2020.01955
Seo K, Akiyoshi H, Ohnishi Y (1999) Alteration of cell wall composition leads to amphotericin B resistance in Aspergillus flavus. Microbiol Immunol 43(11):1017–1025. https://doi.org/10.1111/j.1348-0421.1999.tb01231.x
Shibuya K, Paris S, Ando T, Nakayama H, Hatori T, Latgé JP (2006) Catalases of Aspergillus fumigatus and inflammation in aspergillosis. Nippon Ishinkin Gakkai Zasshi 47(4):249–255. https://doi.org/10.3314/jjmm.47.249
Shlezinger N, Hohl TM (2021) Mitochondrial reactive oxygen species enhance alveolar macrophage activity against Aspergillus fumigatus but are dispensable for host protection. mSphere 6(3):e0026021. https://doi.org/10.1128/mSphere.00260-21
Silva LP, Frawley D (2020) Putative membrane receptors contribute to activation and efficient signaling of mitogen-activated protein kinase cascades during adaptation of Aspergillus fumigatus to different stressors and carbon sources. mSphere 5(5):e00818–20. https://doi.org/10.1128/mSphere.00818-20
Simitsopoulou M, Roilides E, Georgiadou E, Paliogianni F, Walsh TJ (2011) Differential transcriptional profiles induced by amphotericin B formulations on human monocytes during response to hyphae of Aspergillus fumigatus. Med Mycol 49(2):176–185. https://doi.org/10.3109/13693786.2010.510539
Singh RK (2021) Chronic pulmonary aspergillosis in a patient with AIDS. Cureus 13(4):e14588. https://doi.org/10.7759/cureus.14588
Singh N, Yu VL, Rihs JD (1991) Invasive aspergillosis in AIDS. South Med 84(7):822–827. https://doi.org/10.1097/00007611-199107000-00003
Singh A, Sharma B, Mahto KK, Meis JF, Chowdhary A (2020) High-frequency direct detection of triazole resistance in Aspergillus fumigatus from patients with chronic pulmonary fungal diseases in India. J Fungi 6(2):67. https://doi.org/10.3390/jof6020067
Singhal D, Baker L, Wormald PJ, Tan L (2011) Aspergillus fumigatus biofilm on primary human sinonasal epithelial culture. Am J Rhinol Allergy 25(4):219–225. https://doi.org/10.2500/ajra.2011.25.3622
Siopi M, Perlin DS, Arendrup MC (2021) Comparative pharmacodynamics of echinocandins against Aspergillus fumigatus using an in vitro pharmacokinetic/pharmacodynamic model that correlates with clinical response to caspofungin therapy: is there a place for dose optimization? Antimicrob Agents Chemother 65(4):e01618-e1620. https://doi.org/10.1128/aac.01618-20
Snelders E, van der Lee HAL, Kuijpers J, Rijs A, Varga J, Samson RA, Mellado E, Donders ART, Melchers WJG, Verweij PE (2008) Emergence of azole resistance in Aspergillus fumigatus and spread of a single resistance mechanism. Plos Med 5(11):1629–1637. https://doi.org/10.1371/journal.pmed.0050219
Steenwyk JL, Mead ME, de Castro PA, Valero C, Damasio A, Dos Santos RAC, Labella AL, Li Y, Knowles SL, Raja HA, Oberlies NH, Zhou X, Cornely OA, Fuchs F, Koehler P, Goldman GH, Rokas A (2021) Genomic and phenotypic analysis of COVID-19-associated pulmonary aspergillosis isolates of Aspergillus fumigatus. Microbiol Spectr 9(1):e0001021. https://doi.org/10.1128/Spectrum.00010-21
Stemler J, Többen C, Lass-Flörl C, Steinmann J, Ackermann K, Rath PM, Simon M, Cornely OA, Koehler P (2023) Diagnosis and treatment of invasive aspergillosis caused by non-fumigatus Aspergillus spp. J Fungi 9(4):500. https://doi.org/10.3390/jof9040500
Storisteanu DM, Pocock JM, Cowburn AS, Juss JK, Nadesalingam A, Nizet V, Chilvers ER (2017) Evasion of neutrophil extracellular traps by respiratory pathogens. Am J Respir Cell Mol Biol 56(4):423–431. https://doi.org/10.1165/rcmb.2016-0193PS
Sun Y, Tan L, Yao Z, Gao L (2022) In vitro and in vivo interactions of TOR inhibitor AZD8055 and azoles against pathogenic fungi. Microbiol Spectr 10(1):e0200721. https://doi.org/10.1128/spectrum.02007-21
Tavakoli M, Shokohi T, Lass Flörl C, Hedayati MT, Hoenigl M (2020) Immunological response to COVID-19 and its role as a predisposing factor in invasive aspergillosis. Curr Med Mycol 6(4):75–79. https://doi.org/10.18502/cmm.6.4.5442
Tobin JM, Nickolich KL (2020) Influenza suppresses neutrophil recruitment to the lung and exacerbates secondary invasive pulmonary aspergillosis. J Immunol 205(2):480–488. https://doi.org/10.4049/jimmunol.2000067
Toledo H, Sánchez CI, Marín L, Amich J (2022) Regulation of zinc homeostatic genes by environmental pH in the filamentous fungus Aspergillus fumigatus. Environ Microbiol 24(2):643–666. https://doi.org/10.1111/1462-2920.15452
Tsitsigiannis DI, Bok JW, Andes D, Nielsen KF, Frisvad JC, Keller NP (2005) Aspergillus cyclooxygenase-like enzymes are associated with prostaglandin production and virulence. Infect Immun 73(8):4548–4559. https://doi.org/10.1128/iai.73.8.4548-4559.2005
Twumasi-Boateng K, Yu Y, Chen D, Gravelat FN, Nierman WC, Sheppard DC (2009) Transcriptional profiling identifies a role for BrlA in the response to nitrogen depletion and for StuA in the regulation of secondary metabolite clusters in Aspergillus fumigatus. Eukaryot Cell 8(1):104–115. https://doi.org/10.1128/ec.00265-08
Valero C, Colabardini AC, Chiaratto J, Pardeshi L, de Castro PA, Ferreira Filho JA, Silva LP, Rocha MC, Malavazi I (2020) Aspergillus fumigatus transcription factors involved in the caspofungin paradoxical effect. mBio 11(3):e00816–20. https://doi.org/10.1128/mBio.00816-20
Verburg K, van Neer J, Duca M, de Cock H (2022) Novel treatment approach for aspergilloses by targeting germination. J Fungi 8(8):758. https://doi.org/10.3390/jof8080758
Vicentefranqueira R, Amich J (2018) The transcription factor ZafA regulates the homeostatic and adaptive response to zinc starvation in Aspergillus fumigatus. Genes 9(7):318. https://doi.org/10.3390/genes9070318
Vicentefranqueira R, Amich J, Laskaris P, Ibrahim-Granet O, Latgé JP, Toledo H, Leal F, Calera JA (2015) Targeting zinc homeostasis to combat Aspergillus fumigatus infections. Front Microbiol 6:160. https://doi.org/10.3389/fmicb.2015.00160
Vicentefranqueira R, Leal F, Marín L, Sánchez CI, Calera JA (2019) The interplay between zinc and iron homeostasis in Aspergillus fumigatus under zinc-replete conditions relies on the iron-mediated regulation of alternative transcription units of zafA and the basal amount of the ZafA zinc-responsiveness transcription factor. Environ Microbiol 21(8):2787–2808. https://doi.org/10.1111/1462-2920.14618
S J, Vipparti H (2014) Mixed fungal lung infection with Aspergillus fumigatus and Candida albicans in a immunocomprimised patient: case report. J Clin Diagn Res 8(4):Dd08– DD10. https://doi.org/10.7860/jcdr/2014/8048.4272
Wagener J, Echtenacher B, Rohde M, Kotz A, Krappmann S, Heesemann J, Ebel F (2008) The putative alpha-1,2-mannosyltransferase AfMnt1 of the opportunistic fungal pathogen Aspergillus fumigatus is required for cell wall stability and full virulence. Eukaryot Cell 7(10):1661–1673. https://doi.org/10.1128/ec.00221-08
Wang F, **n C, Liu J, Ran Z, Zhao C, Song Z (2020) Interactions between invasive fungi and symbiotic bacteria. World J Microbiol Biotechnol 36(9):137. https://doi.org/10.1007/s11274-020-02913-3
Wei T, Zheng N, Zheng H, Chen Y, Hong P, Liu W, Liu M (2022) Proteomic perspective of azole resistance in Aspergillus fumigatus biofilm extracellular matrix in response to itraconazole. Med Mycol 60(10):myac084. https://doi.org/10.1093/mmy/myac084
White PL, Dhillon R, Cordey A, Hughes H, Faggian F, Soni S, Pandey M, Whitaker H, May A, Morgan M, Wise MP, Healy B, Blyth I, Price JS, Vale L, Posso R, Kronda J, Blackwood A, Rafferty H, Moffitt A, Tsitsopoulou A, Gaur S, Holmes T, Backx M (2021) A national strategy to diagnose coronavirus disease 2019-associated invasive fungal disease in the intensive care unit. Clin Infect Dis 73(7):e1634–e1644. https://doi.org/10.1093/cid/ciaa1298
Wiemann P, Lechner BE, Baccile JA, Velk TA, Yin WB, Bok JW, Pakala S, Losada L, Nierman WC, Schroeder FC, Haas H, Keller NP (2014) Perturbations in small molecule synthesis uncovers an iron-responsive secondary metabolite network in Aspergillus fumigatus. Front Microbiol 5:530. https://doi.org/10.3389/fmicb.2014.00530
Wuyts J, Van Dijck P, Holtappels M (2018) Fungal persister cells: the basis for recalcitrant infections? PLoS Pathog 14(10):e1007301. https://doi.org/10.1371/journal.ppat.1007301
Xue T, Nguyen CK, Romans A, May GS (2004) A mitogen-activated protein kinase that senses nitrogen regulates conidial germination and growth in Aspergillus fumigatus. Eukaryot Cell 3(2):557–560. https://doi.org/10.1128/ec.3.2.557-560.2004
Yan H, Li Z, **a H, Li Q, Bai H (2022) A case report on mixed pulmonary infection of Nocardia nova, Mycobacterium tuberculosis, and Aspergillus fumigatus based on metagenomic next-generation sequencing. Front Public Health 10:927338. https://doi.org/10.3389/fpubh.2022.927338
Zhai P, Shi L, Zhong G, Jiang J, Zhou J, Chen X, Dong G, Zhang L, Li R, Song J (2021) The OxrA protein of Aspergillus fumigatus is required for the oxidative stress response and fungal pathogenesis. Appl Environ Microbiol 87(22):e0112021. https://doi.org/10.1128/aem.01120-21
Zhang X, Jia X, Tian S, Zhang C, Lu Z, Chen Y, Chen F, Li Z, Su X, Han X, Sun Y, Han L (2018) Role of the small GTPase Rho1 in cell wall integrity, stress response, and pathogenesis of Aspergillus fumigatus. Fungal Genet Biol 120:30–41. https://doi.org/10.1016/j.fgb.2018.09.003
Zhang X, He D, Gao S, Wei Y, Wang L (2019) Aspergillus fumigatus enhances human NK cell activity by regulating M1 macrophage polarization. Mol Med Rep 20(2):1241–1249. https://doi.org/10.3892/mmr.2019.10365
Zhang C, Gao L, Ren Y, Gu H, Zhang Y, Lu L (2021a) The CCAAT-binding complex mediates azole susceptibility of Aspergillus fumigatus by suppressing SrbA expression and cleavage. Microbiologyopen 10(6):e1249. https://doi.org/10.1002/mbo3.1249
Zhang Q, Liu F, Zeng M, Mao Y, Song Z (2021b) Drug repurposing strategies in the development of potential antifungal agents. Appl Microbiol Biotechnol 105(13):5259–5279. https://doi.org/10.1007/s00253-021-11407-7
Zhou J, Cai Y, Liu Y, An H, Deng K, Ashraf MA, Zou L, Wang J (2022) Breaking down the cell wall: Still an attractive antibacterial strategy. Front Microbiol 13:952633. https://doi.org/10.3389/fmicb.2022.952633
Zhou X, Zeng M, Huang F, Qin G, Song Z, Liu F (2023) The potential role of plant secondary metabolites on antifungal and immunomodulatory effect Appl Microbiol Biotechnol:1–22. https://doi.org/10.1007/s00253-023-12601-5
Funding
This study was supported by the Sichuan Science and Technology Program (2022NSFSC1539, 2023NSFSC0529, 2023NSFSC1698, and 2022YFS0629), Technology Strategic Cooperation Project of Luzhou Municipal People’s Government–Southwest Medical University (2018LZNYD-ZK26), and the Foundation of Southwest Medical University (2022QN042, 2022QN085, 2022QN102, and 2022QN118).
Author information
Authors and Affiliations
Contributions
ZYS had the idea for the article drafted, FYL wrote the manuscript, MZ, XZ, and FJH performed the literature search and data analysis, and FYL and ZYS critically revised the work. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, F., Zeng, M., Zhou, X. et al. Aspergillus fumigatus escape mechanisms from its harsh survival environments. Appl Microbiol Biotechnol 108, 53 (2024). https://doi.org/10.1007/s00253-023-12952-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-023-12952-z