Abstract
Two major porcine circovirus type 2 (PCV2) genotypes, PCV2a and PCV2b, are recognized. PCV2a was predominant in the global pig population until 2000 while PCV2b became predominant from 2003 onward. The aim of this study was to analyze the immune protection conferred by two PCV2a and two PCV2b capsid proteins (Caps) in pigs challenged with a mutant PCV2b/YJ (mPCV2b/YJ) strain. Pigs vaccinated with PCV2a/LG-Cap and PCV2a/CL-Cap elicited significantly higher levels of PCV2-specific antibodies and neutralizing antibodies compared with PCV2b/JF-Cap and mPCV2b/YJ-Cap. Following a mPCV2b/YJ challenge, no viremia was detected in the PCV2a/LG-Cap and PCV2a/CL-Cap groups, while viremias were found in 20 and 40 % of the pigs in the PCV2b/JF-Cap and mPCV2b/YJ-Cap groups, respectively. Viral loads in the inguinal lymph nodes of pigs from the PCV2b/JF-Cap and mPCV2b/YJ-Cap groups were significantly higher than those in the PCV2a/LG-Cap and PCV2a/CL-Cap groups, but significantly lower than those of the challenge control group. Furthermore, PCV2 antigens were not detected in the inguinal lymph nodes of pigs from commercial vaccine groups, as well as the PCV2a/LG-Cap and PCV2a/CL-Cap groups, but were found in the challenge control (100 %, 5/5), PCV2b/JF-Cap (20 %, 1/5), and mPCV2b/YJ-Cap (20 %, 1/5) groups. These findings suggest that mPCV2b/YJ-Cap and PCV2b/JF-Cap were less immunogenic than PCV2a/LG-Cap and PCV2a/CL-Cap. We speculate that a genotypic shift from PCV2a to PCV2b might be the result of the majority of PCV2a strains being more immunogenic than the majority of PCV2b strains. These results provide a possible explanation for why PCV2b strains are more likely to cause epidemics than PCV2a strains. It tells us that PCV2 pathogenesis may be associated with its immunogenicity to some extent.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Porcine circovirus type 2 (PCV2) was isolated from pigs suffering from post-weaning multisystemic wasting disease (PMWS) in 1991 in Canada (Allan et al. 1998; Ellis et al. 1998), as well as from pigs suffering from other conditions, such as dermatitis and nephropathy syndrome, and some enteric, respiratory, and reproductive disorders. Collectively, these conditions have been grouped under the name of porcine circovirus diseases (PCVDs) or porcine circovirus-associated diseases (PCVADs) (Opriessnig et al. 2007; Segalés 2012). PCV2 is a non-enveloped virus containing a circular, single-stranded DNA molecule of approximately 1768 nucleotides (nt) (Allan et al. 1998; Huang et al. 2011a). The sole structural capsid protein (Cap) is the primary immunogenic protein and has been the target for vaccine development (Lekcharoensuk et al. 2004; Blanchard et al. 2003; Nawagitgul et al. 2000; Mahé et al. 2000).
Several studies have demonstrated that four genotypes of PCV2 exist, including PCV2a, PCV2b, PCV2c, and the recently approved PCV2d (Guo et al. 2010; **ao et al. 2015). PCV2a was the most widely circulating genotype in the field from the early 1990s until 2000, whereas PCV2b became more predominant from 2003 to 2004 onward (Gagnon et al. 2007; Dupont et al. 2008; Wang et al. 2009). While PCV2a and PCV2b are widespread, PCV2c has been identified in archived Danish tissues from the late 1990s and in a fetal pig from Brazil recently (Dupont et al. 2008; Franzo et al. 2015), and PCV2d has been identified in China and in the USA (Guo et al. 2010; **ao et al. 2015). Until now, it has not been clear why the PCV2b genotype has replaced the PCV2a genotype. Our hypothesis is that the immunogenicity of the majority of PCV2b strains is worse than that of the majority of PCV2a strains.
Commercially available PCV2 vaccines are based mostly on the PCV2a genotype and have generally been very effective in pigs, as they provide effective protection against PCV2b strains and some PCV2 mutants (Fort et al. 2008; Opriessnig et al. 2008; Guo et al. 2015). A PCV2 vaccine based on genotype 2b was more effective than a 2a-based vaccine in protecting pigs against PCV2b or a combination of PCV2a and 2b strains (Opriessnig et al. 2013). It is unknown whether a homologous PCV2 Cap could provide better immune protection than a heterologous Cap. PCV2b mutants were reported to be more virulent in vivo (Guo et al. 2012), which raises the possibility that a PCV2b mutant-based vaccine could also provide effective protection in pigs.
In this study, PCV2a/LG-Cap, PCV2a/CL-Cap, PCV2b/JF-Cap, and mutant PCV2b/YJ (mPCV2b/YJ)-Cap were expressed respectively in a baculovirus system, and immune protection was investigated in pigs challenged with the mPCV2b/YJ strain.
Materials and methods
Viruses, cells, and plasmids
The viral strain mPCV2b/YJ (GenBank accession: HM038032), the natural deletion at position 1039 of genome which contributed to the elongation of two amino acids (N and E) at the C terminus of Cap protein, was grown in PK-15 cells and utilized for the challenge experiment (Guo et al. 2015). Two high-titer seed stocks of a recombinant baculovirus expressing recombinant Cap, rBac-PCV2a/LG-Cap and rBac-PCV2a/CL-Cap, were previously produced (Wang et al. 2013; Liu et al. 2004). Sf21 cells were propagated and maintained at 27 °C in Grace’s medium (Invitrogen, Carlsbad, CA, USA) supplemented with 10 % fetal bovine serum (FBS), 100 μg/mL streptomycin, and 100 IU/mL penicillin. PCV1-free PK-15 cells that were grown in minimal essential medium (Invitrogen) containing 10 % heat-inactivated FBS were used for virus propagation. Two recombinant plasmids, pMD18-PCV2b/JF and pMD18-PCV2b/YJ, containing PCV2b/JF and PCV2b genomes, respectively, were previously constructed (Guo et al. 2010).
Analysis of ORF2 from different PCV2 strains
Multiple alignments of amino acid sequences in the capsid protein of four strains of PCV2 (PCV2a/LG, PCV2a/CL, PCV2b/JF, mPCV2b/YJ) were performed using Clustal W within the DNASTAR software (version 7.0).
Construction of recombinant baculoviruses
Two recombinant baculoviruses expressing recombinant Cap from PCV2b/JF and PCV2b/YJ strains, respectively, were constructed as described previously (Wang et al. 2013; Liu et al. 2004). Briefly, the two Cap genes were cloned into the pFastBac Dual™ transfer vector, and the resulting transfer vectors were confirmed by sequencing and called pFBD-PCV2b/JF-Cap and pFBD-PCV2b/YJ-Cap, respectively. Two recombinant baculoviruses, rBac-PCV2b/JF-Cap and rBac-PCV2b/YJ-Cap, were subsequently produced using the Bac-to-Bac Baculovirus Expression System (Invitrogen).
Expression and identification of recombinant proteins
Sf21 cells were separately infected with rBac-PCV2a/CL-Cap, rBac-PCV2a/LG-Cap, rBac-PCV2b/JF-Cap, and rBac-PCV2b/YJ-Cap; harvested 72 h post-infection; and washed twice with phosphate-buffered saline. Baculovirus-containing protein samples were separated using 12 % sodium dodecyl sulfate–polyacrylamide gel electrophoresis and then transferred onto nitrocellulose membranes as described before (Wang et al. 2013; Liu et al. 2004). Western blotting was performed with an anti-PCV2-Cap monoclonal antibody (mAb) 5F2 supernatant [28] (1:10 dilution, primary antibody) and HRP-labeled goat anti-mouse IgG (1:3000 dilution, secondary antibody). The target protein bands were visualized using 3,3ʹ-diaminobenzidine substrate (Zsgb-bio, Bei**g, China).
Preparation of immunogens
The four recombinant baculoviruses described above were harvested from Sf21 cells 72 h post-infection and purified under denaturing conditions (8 M urea) with HisPur™ Ni-NTA Superflow agarose (Pierce Biotechnology, Rockford, IL, USA) according to the manufacturer’s instructions. The eluted fractions containing purified Cap were pooled and dialyzed overnight at 4 °C against 10 mM Tris–HCl buffer (pH 8.0, 0.1 % Triton-X100) to remove the urea and allow the refolding of the protein. After dialysis, the samples were aliquoted and stored at −70 °C until use. The concentrations of the purified proteins were measured as described previously (Wang et al. 2013).
The PCV2a/LG-Cap, PCV2a/CL-Cap, PCV2b/JF-Cap, and PCV2b/YJ-Cap vaccines were prepared by mixing the target protein (1 mg/mL) with the oil-in-water adjuvant ISA 15A VG (Seppic, Castres, France) at a ratio of 85:15 (v/v).
Vaccination and challenge experiments
Forty healthy, 5-week-old, commercial Large White–Dutch Landrace crossbred piglets were obtained and confirmed to be PCV2-, porcine reproductive and respiratory syndrome virus-, classical swine fever virus-, porcine parvovirus type 1-, and pseudorabies virus-negative as described previously (Guo et al. 2015). These piglets were randomly divided into eight groups with five piglets per group, and they were raised separately in different isolation rooms with individual ventilation. Groups A, B, C, and D were intramuscularly immunized with the PCV2a/LG-Cap, PCV2a/CL-Cap, PCV2b/JF-Cap, and mPCV2b/YJ-Cap vaccines, respectively, at a dose of 1 mL/piglet. Groups E and F were intramuscularly inoculated with 1 mL/piglet of the commercial PCV2-LG strain inactivated vaccine (lot number: 2013009, Weike, Harbin, China) and the subunit vaccine (Batch number: 309-835A, Boehringer Ingelheim Vetmedica, St. Joseph, MO, USA), respectively, whereas group G was intramuscularly injected with 1 mL/piglet of the mock inoculum and served as the challenge control group. Group H was the mock group that was not vaccinated and challenged. An identical booster immunization was conducted 3 weeks after the primary immunization. At 0, 7, 14, 21, 28, and 35 days post-primary immunization (dpi), sera from all piglets were collected to detect PCV2 antibodies. All of the piglets (except those in group H) were challenged with mPCV2/YJ strain (104.5 median tissue culture infective doses (TCID50)/mL), l mL intramuscularly and 1 mL intranasally, at 35 dpi. The pigs were weighed at 0 and 28 days post-challenge (dpc), and relative weight gain rates (RWGRs) (%) were calculated according to the following formula: [(the average daily weight gain of vaccination group − the average daily weight gain of challenge group) / the average daily weight gain of challenge group] × 100 %. Sera were collected for viral nucleic acid and antibody analysis at 0, 7, 14, 21, and 28 dpc. All pigs were euthanized by intravenous pentobarbital sodium overdose at 28 dpc. Inguinal lymph nodes were collected for pathology and viral nucleic acid and antigen detection. For microscopic lesion, the size of inguinal lymph nodes was estimated (score range from 0 [normal] to 3 [three times the normal size]) and recorded for each pig.
Serology
All sera were analyzed for the presence of PCV2 antibodies by the immunoperoxidase monolayer assay (IPMA), as previously described (Timmusk et al. 2008). Additionally, PCV2-neutralizing antibodies present in these sera were also identified using a blocking enzyme-linked immunosorbent assay (ELISA), as previously described (Huang et al. 2011b). PCV2a/LG strain was used as antigen in IPMA and ELISA.
Detection of viremia
To investigate the development of viremia after challenge, viral DNA was extracted from sera collected at 0, 7, 14, 21, and 28 dpc using a DNA extraction kit (Tiangen, Bei**g, China), and polymerase chain reactions (PCRs) were performed as previously described (Guo et al. 2015).
Quantification of PCV2 nucleic acid
To test the PCV2 loads of inguinal lymph nodes, viral DNA was extracted from inguinal lymph nodes using a DNA extraction kit (Tiangen). All DNA extracts were tested for the presence of PCV2 DNA by a quantitative PCR (qPCR) targeting a conserved region, as described previously (Wang et al. 2013). The results were calculated as the mean of the logarithmic viral DNA copy number per gram of inguinal lymph node (Log10 copies/gram).
Microscopic lesions and antigen detection
Inguinal lymph nodes were evaluated for lymphoid depletion and histiocytic replacement of follicles, with scores ranging from 0 (none) to 3 (severe) (Opriessnig et al. 2004). The PCV2 antigen in all of the collected inguinal lymph nodes of PCV2-inoculated pigs was detected by indirect immunofluorescence staining described previously with minor modifications (Sanchez et al. 2001). mAb 3A5 (1:800 dilution), which was generated against PCV2a/LG via a procedure described previously (Huang et al. 2011b), was used as the primary antibody. DyLight 488-labeled goat anti-mouse IgG (H+L) (Thermo Fisher Scientific, Waltham, MA, USA) (1:400 dilution) was used as the secondary antibody. Finally, the stained sections were viewed and representative digital images of the stained tissue sections were made using the EVOS® FL Imaging System (Thermo Fisher Scientific).
Statistical analysis
For data analysis, SAS software version 8.1 (SAS Institute, Cary, NC, USA) was used. All data from the IPMA, blocking ELISA, and the qPCR were analyzed using a one-way repeated measurement analysis of variance (ANOVA), followed by a least significance difference (LSD) test. A p value less than 0.05 was considered to be significant.
Results
Expression of recombinant Caps
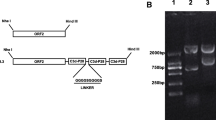
The main amino acid difference between Caps of PCV2a and PCV2b in this study was located at positions 51, 57, 59, 63, 80, 86, 88, 89, 91, 121, 151, 190, 191, 200, 206, 210, and 232 (Fig. 1a). The immunoreactivity of the four Caps was confirmed via an immunoblot analysis. The expression of the PCV2-Cap protein was verified using the PCV2 mAb 5F2, and it exhibited an approximate molecular weight of 28 kDa (Fig. 1b). In contrast, no protein was detected in the negative controls.
a Amino acid alignment of the capsid protein of PCV2 strains used in this study. Boxes show residues that differ from the consensus. The consensus was used as the majority sequence for this alignment. b Identification of expression of PCV2b/JF-Cap, PCV2b/YJ-Cap, PCV2a/CL-Cap, and PCV2a/LG-Cap by Western blotting. PCV2 mAb 5F2 (as the primary antibody) and horseradish peroxidase (HRP)-labeled goat anti-mouse IgG (H + L) (1:4000, as the secondary antibody, Invitrogen, Carlsbad, CA, USA) were used to detect the expression of PCV2 Caps. Lanes 2, 3, 4, and 5 show cell lysate from Sf21 cells infected with rBac-PCV2b/JF-Cap, rBac-mPCV2b/YJ-Cap, rBac-PCV2a/CL-Cap, and rBac-PCV2a/LG-Cap; lane 1 shows the cell lysate from normal Sf21 cells as negative control; lane M is a protein molecular weight marker
Relative weight gain rates and macroscopic lesions
The RWGRs of different groups were calculated and are shown in Fig. 2. Significantly lower RWGRs were observed in the PCV2b/JF-Cap and PCV2b/YJ-Cap vaccine groups than in the other two PCV2a-Cap vaccine groups and the commercial vaccine groups (p < 0.05), while no significant difference in the RWGR was observed among the two PCV2a-Cap vaccine groups and the commercial vaccine groups (Fig. 2). There were mild pathological lesions in the inguinal lymph nodes from the PCV2b/JF-Cap and PCV2b/YJ-Cap vaccine groups and severe pathological lesions in the inguinal lymph nodes from the challenge group (Table 1). No significant macroscopic lesions were observed in the other groups.
Relative weight gain rates (RWGRs) in different groups. RWGRs (%) were calculated according to the following formula: [(the average daily weight gain of the vaccination group − the average daily weight gain of the challenge group) / the average daily weight gain of the challenge group] × 100 %. The RWGR ± SD was calculated for each treatment group throughout the experiment. Different letters (A and B) indicate a statistically significant difference between the different experimental groups (p < 0.05). Error bars represent SDs
Serology
IPMA was used to test pigs for the presence of PCV2 total antibodies, and the IPMA antibody titers in the different groups are shown in Fig. 3a. Pigs vaccinated with PCV2a/LG-Cap and PCV2a/CL-Cap had a stronger and earlier PCV2 antibody response compared with pigs vaccinated with PCV2b/JF-Cap and PCV2b/YJ-Cap. Fourteen days post-vaccination, 80 % (4/5) and 60 % (3/5) of the pigs vaccinated with PCV2a/LG-Cap and PCV2a/CL-Cap, respectively, were seropositive (IPMA titer ≥100), whereas 40 % (2/5) and 20 % (1/5) of the pigs vaccinated with PCV2b/JF-Cap and PCV2b/YJ-Cap, respectively, were seropositive (Fig. 3a). Furthermore, PCV2b/JF-Cap- and PCV2b/YJ-Cap-vaccinated pigs had significantly (p < 0.05) lower mean group IPMA titers compared with PCV2a/LG-Cap- and PCV2a/CL-Cap-vaccinated pigs at 35 dpv (Fig. 3a). All pigs in the challenge group were seronegative before challenge, while 100 % of pigs in the challenge group were seropositive at 28 dpc. The mock group pigs had no detectable anti-PCV2 antibodies throughout the trial period.
PCV2 antibody tests of the different groups. a Detection of PCV2 antibody titers in sera at different times using IPMA. Different letters (a and b) indicate a statistically significant difference in the antibody titers between the different experimental groups (p < 0.05). b Detection of PCV2-neutralizing antibodies in sera at different times using the blocking enzyme-linked immunosorbent assay
PCV2-neutralizing antibodies present in the above sera were also tested using a blocking ELISA (Huang et al. 2011b), and the results are shown in Fig. 3b. PCV2-seropositive pigs were found in PCV2a/LG-Cap and PCV2a/CL-Cap groups at 21 dpv, and 100 % of the pigs from these two groups seroconverted to PCV2 at 28 dpv. At 28 dpv, 60 % of the pigs vaccinated with PCV2b/JF-Cap were seropositive, while 40 % of the pigs vaccinated with PCV2b /YJ-Cap seroconverted to PCV2. All of the pigs in the PCV2b/JF-Cap group and 80 % of the pigs in the PCV2b/YJ-Cap group were PCV2 seropositive at 35 dpv. All of the pigs in the mock group were seronegative throughout the trial period, while all of the pigs in the challenge group were PCV2 seronegative from 0 to 56 dpv/21 dpc and were seropositive at 28 dpc.
Viremia
No viremia was detected in the piglets of the mock, commercial vaccine, PCV2a/LG-Cap, and PCV2a/CL-Cap groups. In contrast, all of the piglets in the challenge group showed viremias from around 14 to 28 dpc (Table 1). For the PCV2b/JF-Cap and PCV2b/YJ-Cap groups, viremias were tested from 21 to 28 dpc, and the rates of viremia were 20 % (1/5) and 40 % (2/5), respectively, at 28 dpc (Table 1).
PCV2 DNA, antigen, and microscopic lesions in inguinal lymph nodes
qPCRs were used to quantify PCV2 DNA in inguinal lymph nodes of all of the pigs at 28 dpc, and the results are shown in Fig. 4a. The PCV2 loads in inguinal lymph nodes from the commercial vaccine groups were significantly lower than those from the challenge group (p < 0.05), while PCV2 DNA was not detectable in the mock group (Fig. 4a). No apparent difference in the PCV2 loads was found between the PCV2a/LG-Cap group (or PCV2a/CL-Cap group) and the commercial vaccine groups (p > 0.05), while significant differences in the viral loads were observed between the PCV2a/LG-Cap group (or PCV2a/CL-Cap group) and the PCV2b/JF-Cap group (or PCV2b/YJ-Cap group) (p < 0.05, Fig. 4a).
Detection of PCV2 genome and antigen in the inguinal lymph nodes. a Quantification of viral DNA loads from different groups at 28 dpc. Quantification of viral DNA loads in lymphoid tissues was performed using a previously described quantitative PCR assay (Wang et al. 2013). The group mean logarithm of the number of viral genomic copies/gram of tissue (±SD) was calculated as the corresponding value on the x axis for each treatment group. Different letters (a, b, and c) indicate a statistically significant difference in the viral DNA loads between the different experimental groups (p < 0.05). b PCV2 antigen detection and microscopic lesions of inguinal lymph nodes observed in pigs from different groups at 28 dpc. PCV2-specific antigens were tested with the PCV2 mAb 5F2 using an indirect immunofluorescence assay. a the PCV2a/LG-Cap group, b the PCV2a/CL-Cap group, c the PCV2b/JF-Cap group, d the PCV2b/YJ-Cap group, e the commercial PCV2-LG strain inactivated vaccine group, f the commercial subunit vaccine group, g the mPCV2b/YJ challenge group without vaccination, and h the mock control without vaccination and challenge
Specific signals of PCV2 Caps were found in inguinal lymph nodes from challenge group pigs at 28 dpc, and the positive rate was 80 %, while no signal was found in those tissues from the mock, commercial vaccine, PCV2a/LG-Cap, and PCV2a/CL-Cap groups at 28 dpc (Fig. 4b). However, PCV2 signals were also observed in the PCV2b/JF-Cap and PCV2b/YJ-Cap groups at 28 dpc, and the positive rates of two groups were 20 and 40 %, respectively.
The microscopic lesion scores of inguinal lymph nodes for all of the pigs are summarized in Table 1. The majority of the pigs in the commercial vaccine groups, two PCV2a-Cap groups, and mock groups had normal lymphoid tissues or developed only mild lesions. Moderate lesions (scores ranging from 3 to 4) were observed in one fifth of PCV2b/JF-Cap-vaccinated pigs, two fifths 2/5 of PCV2b/YJ-Cap-vaccinated pigs, and three fifths of challenged pigs, while severe lesions (scores ranging from 5 to 6) were only observed in two fifths of challenged pigs at 28 dpc.
Overall, two pigs in the challenge group developed serve lymphoid lesions (Fig. 5g) associated with abundant PCV2 antigen (Fig. 4b) and high PCV2 DNA loads (Fig. 3) in their inguinal lymph nodes. Five pigs (one from the PCV2b/JF-Cap group, two from the PCV2b/YJ-Cap group, and two from the challenge group) developed moderate lymphoid lesions (Fig. 5) associated with a small amount of PCV2 antigen (Fig. 4b) and medium PCV2 DNA loads (Fig. 3) in their inguinal lymph nodes at 28 dpc.
Clinical lesions of inguinal lymph nodes in the different groups at 28 dpc. Sections were processed as described in the “Materials and methods” section and stained with hematoxylin and eosin; magnification scale (×200). a the PCV2a/LG-Cap group, b the PCV2a/CL-Cap group, c the PCV2b/JF-Cap group, d the PCV2b/YJ-Cap group, e the commercial PCV2-LG strain inactivated vaccine group, f the commercial subunit vaccine group, g the mPCV2b/YJ challenge group without vaccination, and h the mock control without vaccination and challenge
Discussion
PCV2a was the most widely circulating genotype in the field from the early 1990s until 2000, whereas PCV2b became predominant from 2003 to 2004 onward (Gagnon et al. 2007; Dupont et al. 2008; Wang et al. 2009). Since 2010, pigs in China have begun to be vaccinated with PCV2a vaccines, and the number of vaccinated pigs has increased with time. However, the PCV2-seropositive rate in pigs was high all over the world before vaccination (Segalés et al. 2005). Therefore, the emergence of PCV2b might be the result of immune pressure not only from vaccination, but also from the high seropositive rate in pigs.
The most common form of PCV2 manifestation is a subclinical infection (Segalés 2012). Two outcomes of subclinical infections are recovery and PCVAD. A decrease in viremia most often occurred simultaneously with an increase in the serum neutralizing antibody titer (Fort et al. 2007). Therefore, neutralizing antibodies play an important role in limiting PCV2 infections. In this study, the levels of PCV2-neutralizing antibodies (as determined by the blocking ELISA) of the two PCV2b-Cap groups were significantly lower than those of the two PCV2a-Cap groups and the commercial vaccine groups. Accordingly, the viremias of the two PCV2b-Cap groups were more serious than those of the two PCV2a-Cap groups. Therefore, we conclude that the immunogenicities of PCV2b/YJ Cap and PCV2b/JF Cap were worse than those of PCV2a/LG-Cap and PCV2a/CL Cap. We speculate that a genotypic shift from PCV2a to PCV2b might be the result of the majority of PCV2a strains being more immunogenic than the majority of PCV2b strains. It is not difficult to explain that PCV2b is most frequently isolated from naturally occurring cases of PMWS (Gagnon et al. 2007; Dupont et al. 2008; Cortey et al. 2011; Timmusk et al. 2008) if the aforementioned hypothesis is correct. In this study, only two PCV2a strains and two PCV2b strains were selected and further studies should be performed to reach a more definitive conclusion.
Under the study conditions, the commercial PCV2a-based vaccines (the PCV2a/LG inactivated vaccine and Ingelvac CircoFLEX) and the two PCV2a subunit vaccines (PCV2a/LG-Cap and PCV2a/CL-Cap) significantly reduced detectable PCV2 viremias (Table 1), decreased viral antigen and pathological lesions in the lymph nodes (Figs.4 and 5), and increased the percentages of weight gain (Fig. 2) after mPCV2b/YJ challenge. These results are in accordance with studies from other research groups (Fort et al. 2008; Beach et al. 2011; Beach and Meng 2012). However, vaccination does not completely prevent infection or spread of PCV2 (Beach and Meng 2012). In addition, more genetic variations and recombination associated with PCV2 have been reported worldwide under immune pressure (Li et al. 2010; Guo et al. 2011; Cai et al. 2004; Shang et al. 2009). Therefore, the amino acid difference might result in the difference of epitopes and immunogenicity of Caps. Furthermore, the numbers of predicted glycation sites of ε amino groups of lysines in PCV2b/JF-Cap (positions 41, 63, 100, 132, 164, 179, and 227) and mPCV2b/YJ-Cap (positions 41, 63, 100, 164, 179, and 227) were more than that of PCV2a/LG-Cap (positions 41, 164, and 179) and PCV2a/CL-Cap (positions 41, 102, 164, and 179) in this study with NetGlycate 1.0 Server from Technical University of Denmark (http://www.cbs.dtu.dk/services/NetGlycate/). The extra different glycation sites (positions 63 and 227) of two PCV2b-Caps were also located in and near to the highest protrusion (Khayat et al. 2011) which is demonstrated as one of the main immunoreactive regions (Saha et al. 2012; Huang et al. 2011a; Liu et al. 2013). It was reported that the loss of glycan residues in the ectodomain of GP5 enhances the immunogenicity of the nearby neutralization epitope (Ansari et al. 2006). Therefore, it is speculated that the glycations of lysines at positions 63 and 227 may result in less immunogenicity of PCV2b/JF-Cap and mPCV2b/YJ-Cap than other two PCV2a Caps.
In summary, the immunoprotection of two PCV2a-Caps and two PCV2b-Caps was analyzed in pigs, and the two PCV2a-Caps were more effective based on the RWGRs, PCV2 DNA loads, levels of PCV2 antigens, the presence of macroscopic and microscopic lesions in inguinal lymph nodes, viremias, and the presence of PCV2-specific antibodies. These results may explain why PCV2b strains are more likely to cause epidemics than PCV2a strains, and they enlighten us to consider the role of PCV2 immunogenicity in viral pathogenesis.
References
Allan GM, McNeilly F, Kennedy S, Daft B, Ellis JA, Haines DM, Meehan BM, Adair BM (1998) Isolation of porcine circovirus-like viruses from pigs with a wasting disease in the USA and Europe. J Vet Diagn Investig 10:3–10
Ansari IH, Kwon B, Osorio FA, Pattnaik AK (2006) Influence of N-linked glycosylation of porcine reproductive and respiratory syndrome virus GP5 on virus infectivity, antigenicity, and ability to induce neutralizing antibodies. J Virol 80:3994–4004
Beach NM, Meng XJ (2012) Efficacy and future prospects of commercially available and experimental vaccines against porcine circovirus type 2 (PCV2). Virus Res 164:33–42
Beach NM, Ramamoorthy S, Opriessnig T, Wu SQ, Meng XJ (2011) Novel chimeric porcine circovirus (PCV) with the capsid gene of the emerging PCV2b subtype cloned in the genomic backbone of the non-pathogenic PCV1 is attenuated in vivo and induces protective and cross-protective immunity against PCV2b and PCV2a subtypes in pigs. Vaccine 29:221–232
Blanchard P, Mahé D, Cariolet R, Keranflec’h A, Baudouard MA, Cordioli P, Albina E, Jestin A (2003) Protection of swine against post-weaning multisystemic wasting syndrome (PMWS) by porcine circovirus type 2 (PCV2) proteins. Vaccine 21:4565–4575
Cai L, Ni J, **a Y, Zi Z, Ning K, Qiu P, Li X, Wang B, Liu Q, Hu D, Yu X, Zhou Z, Zhai X, Han X, Tian K (2012) Identification of an emerging recombinant cluster in porcine circovirus type 2. Virus Res 165:95–102
Cortey M, Pileri E, Sibila M, Pujols J, Balasch M, Plana J (2011) Genotypic shift of porcine circovirus type 2 from PCV-2a to PCV-2b in Spain from 1985 to 2008. Vet J 187:363–368
Dupont K, Nielsen EO, Bækbo P, Larsen LE (2008) Genomic analysis of PCV2 isolates from Danish archives and a current PMWS case-control study supports a shift in genotypes with time. Vet Microbiol 128:56–64
Ellis J, Hassard L, Clark E, Harding J, Allan G, Willson P, Strokappe J, Martin K, McNeilly F, Meehan B, Todd D, Haines D (1998) Isolation of circovirus from lesions of pigs with postweaning multisystemic wasting syndrome. Can Vet J 39:44
Fort M, Olvera A, Sibila M, Segalés J, Mateu E (2007) Detection of neutralizing antibodies in postweaning multisystemic wasting syndrome (PMWS)-affected and non-PMWS-affected pigs. Vet Microbiol 125:244–255
Fort M, Sibila M, Allepuz A, Mateu E, Roerink F, Segalés J (2008) Porcine circovirus type 2 (PCV2) vaccination of conventional pigs prevents viremia against PCV2 isolates of different genotypes and geographic origins. Vaccine 26:1063–1071
Franzo G, Cortey M, de Castro AM, Piovezan U, Szabo MP, Drigo M, Segalés J, Richtzenhain LJ (2015) Genetic characterisation of Porcine circovirus type 2 (PCV2) strains from feral pigs in the Brazilian Pantanal: an opportunity to reconstruct the history of PCV2 evolution. Vet Microbiol 178(1–2):158–162
Gagnon CA, Tremblay D, Tijssen P, Venne MH, Houde A, Elahi SM (2007) The emergence of porcine circovirus 2b genotype (PCV-2b) in swine in Canada. Can Vet J 48:811–819
Guo LJ, Lu YH, Wei YW, Huang LP, Liu CM (2010) Porcine circovirus type 2 (PCV2):genetic variation and newly emerging genotypes in China. Virol J 7:273
Guo L, Lu Y, Wei Y, Huang L, Wu H, Liu C (2011) Porcine circovirus genotype 2a (PCV2a)and genotype 2b (PCV2b) recombinant mutants showed significantly enhanced viral replication and altered antigenicity in vitro. Virology 419:57–63
Guo L, Fu Y, Wang Y, Lu Y, Wei Y, Tang Q, Fan P, Liu J, Zhang L, Zhang F, Huang L, Liu D, Li S, Wu H, Liu C (2012) A porcine circovirus type 2(PCV2) mutant with 234 amino acids in capsid protein showed more virulencein vivo, compared with classical PCV2a/b strain. PLoS One 7:e41463
Guo LJ, Fu YJ, Huang LP, Wang YP, Wei YW, Wu HL, Liu CM (2015) A commercial PCV2a-based vaccine is effective in protection from experimental challenge of PCV2 mutant with two amino acids elongation in capsid protein. Vaccine 33:3752–3757
Huang LP, Lu YH, Wei YW, Guo LJ, Liu CM (2011a) Identification of one critical amino acid that determines a conformational neutralizing epitope in the capsid protein of porcine circovirus type 2. BMC Microbiol 11:188
Huang LP, Lu YH, Wei YW, Guo LJ, Liu CM (2011b) Development of a blocking ELISA for detection of serum neutralizing antibodies against porcine circovirus type 2. J Virol Methods 171:26–33
Khayat R, Brunn N, Speir JA, Hardham JM, Ankenbauer RG, Schneemann A, Johnson JE (2011) The 2.3-angstrom structure of porcine circovirus 2. J Virol 85:7856–7862
Lekcharoensuk P, Morozov I, Paul PS, Thangthumniyom N, Wajjawalku W, Meng XJ (2004) Epitope map** of the major capsid protein of type 2 porcine circovirus (PCV2) by using chimeric PCV1 and PCV2. J Virol 78:8135–8145
Li W, Wang X, Ma T, Feng Z, Li Y, Jiang P (2010) Genetic analysis of porcine circovirus type 2 (PCV2) strains isolated between 2001 and 2009: genotype PCV2b predominate in postweaning multisystemic wasting syndrome occurrences in eastern China. Virus Genes 40:244–251
Liu C, Ihara T, Nunoya T, Ueda S (2004) Development of an ELISA based on the baculovirus-expressed capsid protein of porcine circovirus type 2 as antigen. J Vet Med Sci 66:237–242
Liu J, Huang L, Wei Y, Tang Q, Liu D, Wang Y, Li S, Guo L, Wu H, Liu C (2013) Amino acid mutations in the capsid protein produce novel porcine circovirus type 2 neutralizing epitopes. Vet Microbiol 165:260–267
Mahé D, Blanchard P, Truong C, Arnauld C, Le Cann P, Cariolet R, Madec F, Albina E, Jestin A (2000) Differential recognition of ORF2 protein from type 1 and type 2 porcine circoviruses and identification of immunorelevant epitopes. J Gen Virol 81:1815–1824
Nawagitgul P, Morozov I, Bolin SR, Harms PA, Sorden SD, Paul PS (2000) Open reading frame 2 of porcine circovirus type 2 encodes a major capsid protein. J Gen Virol 81:2281–2287
Opriessnig T, Thacker EL, Yu S, Fenaux M, Meng XJ, Halbur PG (2004) Experimental reproduction of postweaning multisystemic wasting syndrome in pigs by dual infection with Mycoplasma hyopneumoniae and porcine circovirus type 2. Vet Pathol 41:624–640
Opriessnig T, Meng XJ, Halbur PG (2007) Porcine circovirus type 2 associated disease: update on current terminology, clinical manifestations, pathogenesis, diagnosis, and intervention strategies. J Vet Diagn Investig 19:591–615
Opriessnig T, Ramamoorthy S, Madson DM, Patterson AR, Pal N, Carman S, Meng XJ, Halbur PG (2008) Differences in virulence among porcine circovirus type 2 isolates are unrelated to cluster type 2a or 2b and prior infection provides heterologous protection. J Gen Virol 89:2482–2491
Opriessnig T, O’Neill K, Gerber PF, de Castro AM, Gimenéz-Lirola LG, Beach NM, Zhou L, Meng XJ, Wang C, Halbur PG (2013) A PCV2 vaccine based on genotype 2b is more effective than a 2a-based vaccine to protect against PCV2b or combined PCV2a/2b viremia in pigs with concurrent PCV2, PRRSV and PPV infection. Vaccine 31:487–494
Saha D, Huang L, Bussalleu E, Lefebvre D, Fort M, Doorsselaere JV, Nauwynck HJ (2012) Antigenic subty** and epitopes’ competition analysis of porcine circovirus type 2 using monoclonal antibodies. Vet Microbiol 157:13–22
Sanchez RE, Nauwynck HJ, McNeilly F, Allan G, Pensaert MB (2001) Porcine circovirus 2 infection in swine foetuses inoculated at different ages of gestation. Vet Microbiol 83:169–176
Segalés J (2012) Porcine circovirus type 2 (PCV2) infections: clinical signs, pathology and laboratory diagnosis. Virus Res 164:10–19
Segalés J, Allan GM, Domingo M (2005) Porcine circovirus diseases. Anim Health Res Rev 6:119–142
Shang SB, ** YL, Jiang XT, Zhou JY, Zhang X, ** of antigenic epitopes on capsid proteins of porcine circovirus, and antigenic phenotype of porcine circovirus type 2. Mol Immunol 46:327–334
Timmusk S, Wallgren P, Brunborg IM, Wikström FH, Allan G, Meehan B, McMenamy M, McNeilly F, Fuxler L, Belák K, Põdersoo D, Saar T, Berg M, Fossum C (2008) Phylogenetic analysis of porcine circovirus type 2 (PCV2) pre- and post-epizootic postweaning multisystemic wasting syndrome (PMWS). Virus Genes 36:509–520
Wang F, Guo X, Ge X, Wang Z, Chen Y, Cha Z (2009) Genetic variation analysis of Chinese strains of porcine circovirus type 2. Virus Res 145:151–156
Wang YP, Liu D, Guo LJ, Tang QH, Wei YW, Wu HL, Liu JB, Li SB, Huang LP, Liu CM (2013) Enhanced protective immune response to PCV2 subunit vaccine by co-administration of recombinant porcine IFN-γ in mice. Vaccine 31:833–838
**ao CT, Halbur PG, Opriessnig T (2015) Global molecular genetic analysis of porcine circovirus type 2 (PCV2) sequences confirms the presence of four main PCV2 genotypes and reveals a rapid increase of PCV2d. J Gen Virol 96:1830–1841
Acknowledgments
This study was supported by the National Natural Science Foundation of China (Grant No. 31302110) and the Public Welfare Special Funds for Agricultural Scientific Research (Grant No. 201203039).