Abstract
Purpose
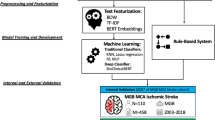
Data extraction from radiology free-text reports is time consuming when performed manually. Recently, more automated extraction methods using natural language processing (NLP) are proposed. A previously developed rule-based NLP algorithm showed promise in its ability to extract stroke-related data from radiology reports. We aimed to externally validate the accuracy of CHARTextract, a rule-based NLP algorithm, to extract stroke-related data from free-text radiology reports.
Methods
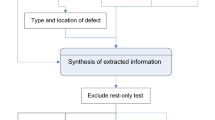
Free-text reports of CT angiography (CTA) and perfusion (CTP) studies of consecutive patients with acute ischemic stroke admitted to a regional stroke center for endovascular thrombectomy were analyzed from January 2015 to 2021. Stroke-related variables were manually extracted as reference standard from clinical reports, including proximal and distal anterior circulation occlusion, posterior circulation occlusion, presence of ischemia or hemorrhage, Alberta stroke program early CT score (ASPECTS), and collateral status. These variables were simultaneously extracted using a rule-based NLP algorithm. The NLP algorithm’s accuracy, specificity, sensitivity, positive predictive value (PPV), and negative predictive value (NPV) were assessed.
Results
The NLP algorithm’s accuracy was > 90% for identifying distal anterior occlusion, posterior circulation occlusion, hemorrhage, and ASPECTS. Accuracy was 85%, 74%, and 79% for proximal anterior circulation occlusion, presence of ischemia, and collateral status respectively.
The algorithm confirmed the absence of variables from radiology reports with an 87–100% accuracy.
Conclusions
Rule-based NLP has a moderate to good performance for stroke-related data extraction from free-text imaging reports. The algorithm’s accuracy was affected by inconsistent report styles and lexicon among reporting radiologists.
Similar content being viewed by others
Abbreviations
- ASPECTS:
-
Alberta stroke program early CT score
- CTA:
-
CT angiography
- CTP:
-
CT perfusion
- ML:
-
Machine learning
- NLP:
-
Natural language processing
References
Yu AY, Holodinsky JK, Zerna C, Svenson LW, Jetté N, Quan H, Hill MD (2016) Use and utility of administrative health data for stroke research and surveillance. Stroke 47(7):1946–1952. https://doi.org/10.1161/STROKEAHA.116.012390
Elkins JS, Friedman C, Boden-Albala B, Sacco RL, Hripcsak G (2000) Coding neuroradiology reports for the Northern Manhattan Stroke Study: a comparison of natural language processing and manual review. Comput Biomed Res 33(1):1–10. https://doi.org/10.1006/cbmr.1999.1535
Pons E, Braun LM, Hunink MG, Kors JA (2016) Natural language processing in radiology: a systematic review. Radiology 279(2):329–343. https://doi.org/10.1148/radiol.16142770
Garg R, Oh E, Naidech A, Kording K, Prabhakaran S (2019) Automating ischemic stroke subtype classification using machine learning and natural language processing. J Stroke Cerebrovasc Dis 28(7):2045–2051. https://doi.org/10.1016/j.jstrokecerebrovasdis.2019.02.004
Waqas M, Rai AT, Vakharia K, Chin F, Siddiqui AH (2020) Effect of definition and methods on estimates of prevalence of large vessel occlusion in acute ischemic stroke: a systematic review and meta-analysis. J Neurointerv Surg 12(3):260–265. https://doi.org/10.1136/neurintsurg-2019-015172
Casey A, Davidson E, Poon M, Dong H, Duma D, Grivas A, Grover C, Suárez-Paniagua V, Tobin R, Whiteley W et al (2021) A systematic review of natural language processing applied to radiology reports. BMC Med Inform Decis Mak 21(1):179. https://doi.org/10.1186/s12911-021-01533-7
Yu AYX, Liu ZA, Pou-Prom C, Lopes K, Kapral MK, Aviv RI, Mamdani M (2021) Automating Stroke Data extraction from free-text radiology reports using natural language processing: instrument validation study. JMIR Med Inform 9(5):e24381. https://doi.org/10.2196/24381
Ong CJ, Orfanoudaki A, Zhang R, Caprasse FPM, Hutch M, Ma L, Fard D, Balogun O, Miller MI, Minnig M et al (2020) Machine learning and natural language processing methods to identify ischemic stroke, acuity and location from radiology reports. PLoS One 15(6):e0234908. https://doi.org/10.1371/journal.pone.0234908
Li MD, Lang M, Deng F, Chang K, Buch K, Rincon S, Mehan WA, Leslie-Mazwi TM, Kalpathy-Cramer J (2021) Analysis of stroke detection during the COVID-19 pandemic using natural language processing of radiology reports. AJNR Am J Neuroradiol 42(3):429–434. https://doi.org/10.3174/ajnr.A6961
Mozayan A, Fabbri AR, Maneevese M, Tocino I, Chheang S (2021) Practical guide to natural language processing for radiology. Radiographics 41(5):1446–1453. https://doi.org/10.1148/rg.2021200113
Zech J, Pain M, Titano J, Badgeley M, Schefflein J, Su A, Costa A, Bederson J, Lehar J, Oermann EK (2018) Natural language-based machine learning models for the annotation of clinical radiology reports. Radiology 287(2):570–580. https://doi.org/10.1148/radiol.2018171093
Kim C, Zhu V, Obeid J, Lenert L (2019) Natural language processing and machine learning algorithm to identify brain MRI reports with acute ischemic stroke. PLoS One 14(2):e0212778. https://doi.org/10.1371/journal.pone.0212778
Peng Y, Wang X, Lu L, Bagheri M, Summers R, Lu Z (2018) NegBio: a high-performance tool for negation and uncertainty detection in radiology reports. AMIA Jt Summits Transl Sci Proc 2017:188–196
Sykes D, Grivas A, Grover C, Tobin R, Sudlow C, Whiteley W, Mcintosh A, Whalley H, Alex B (2021) Comparison of rule-based and neural network models for negation detection in radiology reports. Nat Lang Eng 27(2):203–224. https://doi.org/10.1017/S1351324920000509
Gorinski PJ, Wu H, Grover C, Tobin R, Talbot C, Whalley H, Whiteley W, Alex B (2019) Named entity recognition for electronic health records: a comparison of rule-based and machine learning approaches. CoRR [Internet]. https://arxiv.org/abs/1903.03985. Accessed 20 Feb 2022
Davidson EM, Poon MTC, Casey A, Grivas A, Duma D, Dong H, Suárez-Paniagua V, Grover C, Tobin R, Whalley H et al (2021) The reporting quality of natural language processing studies: systematic review of studies of radiology reports. BMC Med Imaging 21(1):142. https://doi.org/10.1186/s12880-021-00671-8
Houssein EH, Mohamed RE, Ali AA (2021) Machine learning techniques for biomedical natural language processing: a comprehensive review. IEEE Access 9:140628–140653. https://doi.org/10.1109/ACCESS.2021.3119621
CHARTextract – Li Ka Shing Centre for Healthcare Analytics Research & Training (LKS-CHART). 2019. https://lks-chart.github.io/CHARTextract-docs/. Accessed 20 Feb 2022
Wintermark M, Albers GW, Broderick JP, Demchuk AM, Fiebach JB, Fiehler J, Grotta JC, Houser G, Jovin TG, Lees KR et al (2013) Stroke Imaging Research (STIR) and Virtual International Stroke Trials Archive (VISTA)-Imaging Investigators. Acute Stroke Imaging Research Roadmap II. Stroke 44(9):2628–39. https://doi.org/10.1161/STROKEAHA.113.002015
Zaidat OO, Yoo AJ, Khatri P, Tomsick TA, von Kummer R, Saver JL, Marks MP, Prabhakaran S, Kallmes DF, Fitzsimmons BF et al (2013) Cerebral Angiographic Revascularization Grading (CARG) Collaborators; STIR Revascularization working group; STIR Thrombolysis in Cerebral Infarction (TICI) Task Force. Recommendations on angiographic revascularization grading standards for acute ischemic stroke: a consensus statement. Stroke 44(9):2650–63. https://doi.org/10.1161/STROKEAHA.113.001972
Shijie Wu, Mark Dredze. Beto, Bentz, Becas (2019) The surprising cross-lingual effectiveness of BERT. In Kentaro Inui, **g Jiang, Vincent Ng, **aojun Wan 0001, editors, Proceedings of the 2019 conference on empirical methods in natural language processing and the 9th International Joint Conference on Natural Language Processing, EMNLP-IJCNLP 2019, Association for Computational Linguistics, Hong Kong, China, pp. 833–844. https://doi.org/10.18653/v1/D19-1077
Peng Y, Yan K, Sandfort V, Summers RM, Lu Z (2019) A self-attention based deep learning method for lesion attribute detection from CT reports. Int Conf Healthcare Info (ICHI). 1–5. IEEE. https://doi.org/10.1109/ICHI.2019.8904668
Funding
Paulo Puac-Polanco was supported by a University of Ottawa Brain and Mind Research Institute Trainee Award. The remaining authors did not receive support from any organization for the submitted work. The authors have no relevant financial or non-financial interests to disclose.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest or non-financial interest in the subject matter or materials discussed in this manuscript.
Ethics approval
This retrospective chart review study involving human participants was in accordance with the ethical standards of the institutional and national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Study was Ethics approved by the Ottawa Health Science Network Research Ethics Board (OHSN-REB) Protocol #20200309-01H.
Informed consent
The requirement for informed consent was waived by the Ottawa Health Science Network Research Ethics Board (OHSN-REB).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gunter, D., Puac-Polanco, P., Miguel, O. et al. Rule-based natural language processing for automation of stroke data extraction: a validation study. Neuroradiology 64, 2357–2362 (2022). https://doi.org/10.1007/s00234-022-03029-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00234-022-03029-1