Abstract
Key message
Seventeen PHS-QTLs and candidate genes were obtained, including eleven major loci, three under multiple environments and two with co-localization by the other map** methods; The functions of three candidate genes were validated using mutants; nine target proteins and five networks were filtered by joint analysis of GWAS and WGCNA.
Abstract
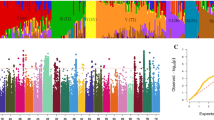
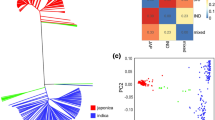
Seed dormancy (SD) and pre-harvest sprouting (PHS) affect yield, as well as grain and hybrid quality in seed production. Therefore, identification of genetic and regulatory pathways underlying PHS and SD is key to gene function analysis, allelic variation mining and genetic improvement. In this study, 78,360 SNPs by SLAF-seq of 230 maize chromosome segment introgression lines (ILs), PHS under five environments were used to conduct GWAS (genome wide association study) (a threshold of 1/n), and seventeen unreported PHS QTLs were obtained, including eleven QTLs with PVE > 10% and three QTLs under multiple environments. Two QTL loci were co-located between the other two genetic map** methods. Using differential gene expression analyses at two stages of grain development, gene functional analysis of Arabidopsis mutants, and gene functional analysis in the QTL region, seventeen PHS QTL-linked candidate genes were identified, and their five molecular regulatory networks constructed. Based on the Arabidopsis T-DNA mutations, three candidate genes were shown to regulate for SD and PHS. Meanwhile, using RNA-seq of grain development, the weighted correlation network analysis (WGCNA) was performed, deducing five regulatory pathways and target genes that regulate PHS and SD. Based on the conjoint analysis of GWAS and WGCNA, four pathways, nine target proteins and target genes were revealed, most of which regulate cell wall metabolism, cell proliferation and seed dehydration tolerance. This has important theoretical and practical significance for elucidating the genetic basis of maize PHS and SD, as well as mining of genetic resources and genetic improvement of traits.







Similar content being viewed by others
Data availability
All raw data generated of 36 samples used in this study were deposited in Bioproject in National Genomics Data Center (NGDC) database with the accession number PRJCA017883 subPRO026577.
References
Abbas N, Maurya JP, Senapati D, Gangappa SN, Chattopadhyay S (2014) Arabidopsis CAM7 and HY5 physically interact and directly bind to the HY5 promoter to regulate its expression and thereby promote photomorphogenesis. Plant Cell 26(3):1036–1052
Aderinola TA, Fagbemi TN, Enujiugha VN, Alashi AM, Aluko RE (2018) In vitro antihypertensive and antioxidative properties of trypsin-derived Moringa oleifera seed globulin hydrolyzate and its membrane fractions. Food Sci Nutr 7(1):132–138
Anderson AC, Stangherlin S, Pimentel KN, Weadge JT, Clarke AJ (2022) The SGNH hydrolase family: a template for carbohydrate diversity. Glycobiology 32(10):826–848
Arc E, Galland M, Godin B, Cueff G, Rajjou L (2013) Nitric oxide implication in the control of seed dormancy and germination. Front Plant Sci 4:346
Barkan A, Small I (2014) Pentatricopeptide repeat proteins in plants. Annu Rev Plant Biol 65:415–442
Bekturova A, Oshanova D, Tiwari P, Nurbekova Z, Kurmanbayeva A, Soltabayeva A, Yarmolinsky D, Srivastava S, Turecková V, Strnad M, Sagi M (2021) Adenosine 5’ phosphosulfate reductase and sulfite oxidase regulate sulfite-induced water loss in Arabidopsis. J Exp Bot 72(18):6447–6466
Cai G, Kim SC, Li J, Zhou Y, Wang X (2020) Transcriptional regulation of lipid catabolism during seedling establishment. Mol Plant 13(7):984–1000
Cao Y, Dai Y, Cui SJ, Ma LG (2008) Histone H2B monoubiquitination in the chromatin of flowering locus c regulates flowering time in Arabidopsis. Plant Cell 20:2586–2602
Cascales-Miñana B, Muñoz-Bertomeu J, Flores-Tornero M, Anoman AD, Pertusa J, Alaiz M, Osorio S, Fernie AR, Segura J, Ros R (2013) The phosphorylated pathway of serine biosynthesis is essential both for male gametophyte and embryo development and for root growth in Arabidopsis. Plant Cell 25(6):2084–2101
Chen X, Wang H, Li X, Ma K, Zhan Y, Zeng F (2019) Molecular cloning and functional analysis of 4-Coumarate: CoA ligase 4(4CL-like 1) from Fraxinus mandshurica and its role in abiotic stress tolerance and cell wall synthesis. BMC Plant Biol 19(1):231
Collakova E, Goyer A, Naponelli V, Krassovskaya I, Gregory JF 3rd, Hanson AD, Shachar-Hill Y (2008) Arabidopsis 10-formyl tetrahydrofolate deformylases are essential for photorespiration. Plant Cell 20(7):1818–1832
Collin A, Daszkowska-Golec A, Szarejko I (2021) Updates on the role of abscisic acid insensitive 5 (ABI5) and abscisic acid-responsive element binding factors (ABFs) in ABA signaling in different developmental stages in plants. Cells 10(8):1996
Colucci G, Apone F, Alyeshmerni N, Chalmers D, Chrispeels MJ (2002) GCR1, the putative Arabidopsis G protein-coupled receptor gene is cell cycle-regulated, and its overexpression abolishes seed dormancy and shortens time to flowering. Proc Natl Acad Sci U S A 99(7):4736–4741
Do TH, Martinoia E, Lee Y (2018) Functions of ABC transporters in plant growth and development. Curr Opin Plant Biol 41:32–38
Dolui AK, Vijayaraj P (2020) Functional omics identifies serine hydrolases that mobilize storage lipids during rice seed germination. Plant Physiol 184(2):693–708
Dutilleul C, Garmier M, Noctor G, Mathieu C, Chétrit P, Foyer CH, de Paepe R (2003) Leaf mitochondria modulate whole cell redox homeostasis, set antioxidant capacity, and determine stress resistance through altered signaling and diurnal regulation. Plant Cell 15(5):1212–1226
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620
Finch-Savage WE, Leubner-Metzger G (2006) Seed dormancy and the control of germination. New Phytol 171(3):501–523
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415
Footitt S, Müller K, Kermode AR, Finch-Savage WE (2015) Seed dormancy cycling in Arabidopsis: chromatin remodelling and regulation of DOG1 in response to seasonal environmental signals. Plant J 81(3):413–425
Footitt S, Walley PG, Lynn JR, Hambidge AJ, Penfield S, Finch-Savage WE (2020) Trait analysis reveals DOG1 determines initial depth of seed dormancy, but not changes during dormancy cycling that result in seedling emergence timing. New Phytol 225(5):2035–2047
Fu J, Cheng Y, Linghu J, Yang X, Kang L, Zhang Z, Zhang J, He C, Du X, Peng Z, Wang B, Zhai L, Dai C, Xu J, Wang W, Li X, Zheng J, Chen L, Luo L, Liu J, Qian X, Yan J, Wang J, Wang G (2013) RNA sequencing reveals the complex regulatory network in the maize kernel. Nat Commun 4:2832
Gao X, Ren F, Lu YT (2006) The Arabidopsis mutant stg1 identifies a function for TBP-associated factor 10 in plant osmotic stress adaptation. Plant Cell Physiol 47(9):1285–1294
Gou M, Yang X, Zhao Y, Ran X, Song Y, Liu CJ (2019) Cytochrome b5 is an obligate electron shuttle protein for syringyl lignin biosynthesis in Arabidopsis. Plant Cell 31(6):1344–1366
Grennan AK (2011) Metallothioneins, a diverse protein family. Plant Physiol 155(4):1750–1751
Gu TY, Qi ZA, Chen SY, Yan J, Fang ZJ, Wang JM, Gong JM (2023) Dual-function DEFENSIN 8 mediates phloem cadmium unloading and accumulation in rice grains. Plant Physiol 191(1):515–527
Guan B, Jiang YT, Lin DL, Lin WH, Xue HW (2022) Phosphatidic acid suppresses autophagy through competitive inhibition by binding GAPC (glyceraldehyde-3-phosphate dehydrogenase) and PGK (phosphoglycerate kinase) proteins. Autophagy 18(11):2656–2670
Guo L, Yu Y, Law JA, Zhang X (2010) Set domain group2 is the major histone H3 lysine [corrected] 4 trimethyltransferase in Arabidopsis. Proc Natl Acad Sci U S A 107(43):18557–18562
Heras B, Drøbak BK (2002) PARF-1: an Arabidopsis thaliana FYVE-domain protein displaying a novel eukaryotic domain structure and phosphoinositide affinity. J Exp Bot 53(368):565–567
Hirano M, Hirano T (2002) Hinge-mediated dimerization of SMC protein is essential for its dynamic interaction with DNA. EMBO J 21(21):5733–5744
Huang AHC (2018) Plant lipid droplets and their associated proteins: potential for rapid advances. Plant Physiol 176(3):1894–1918
Huang X, Feng Q, Qian Q, Zhao Q, Wang L, Wang A, Guan J, Fan D, Weng Q, Huang T, Dong G, Sang T, Han B, Tao Y (2009) High-throughput genoty** by whole-genome resequencing. Genome Res 19(6):1068–1076
Huang T, Bo QuB, Li HP, Zuo DY, Zhao ZX, Liao YC (2012) A maize viviparous 1 gene increases seed dormancy and preharvest sprouting tolerance in transgenic wheat. J Cereal Sci 55(2):166–173
Jabrin S, Ravanel S, Gambonnet B, Douce R, Rébeillé F (2003) One-carbon metabolism in plants. Regulation of tetrahydrofolate synthesis during germination and seedling development. Plant Physiol 131(3):1431–1439
Jackson JP, Lindroth AM, Cao X, Jacobsen SE (2002) Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature 416:556–560
Jiang H, Fang Y, Yan D, Liu ST, Wei J, Guo FL, Wu XT, Cao H, Yin CB, Lu F, Gao LF, Liu YX (2022) Genome-wide association study reveals a NAC transcription factor TaNAC074 linked to pre-harvest sprouting tolerance in wheat. Theor Appl Genet 135(9):3265–3276
Kaewthai N, Gendre D, Eklöf JM, Ibatullin FM, Ezcurra I, Bhalerao RP, Brumer H (2013) Group III-A XTH genes of Arabidopsis encode predominant xyloglucan endohydrolases that are dispensable for normal growth. Plant Physiol 161(1):440–454
Kalachova T, Škrabálková E, Pateyron S, Soubigou-Taconnat L, Djafi N, Collin S, Sekereš J, Burketová L, Potocký M, Pejchar P, Ruelland E (2022) Diacylglycerol kinase 5 participates in flagellin-induced signaling in Arabidopsis. Plant Physiol 190(3):1978–1996
Kam**a HH, Craig EA (2010) The HSP70 chaperone machinery: J proteins as drivers of functional specificity. Nat Rev Mol Cell Biol 11(8):579–592
Kang J, Yim S, Choi H, Kim A, Lee KP, Lopez-Molina L, Martinoia E, Lee Y (2015) Abscisic acid transporters cooperate to control seed germination. Nat Commun 6:8113
Kelly AA, Quettier AL, Shaw E, Eastmond PJ (2011) Seed storage oil mobilization is important but not essential for germination or seedling establishment in Arabidopsis. Plant Physiol 157(2):866–875
Kent WJ (2002) BLAT-the BLAST-like alignment tool. Genome Res 12(4):656–664
Kim JS, Jung HJ, Lee HJ, Kim KA, Goh C-H, Woo Y, Oh SH, Han YS, Kang H (2008) Glycine-rich RNA-binding protein7 affects abiotic stress responses by regulating stomata opening and closing in Arabidopsis thaliana. Plant J 55(3):455–466
Kong MS, Qiao Q, Ma XL, Tao YS, Maharjan RP, Zhen WC (2020) Isolation and functional analysis of the ZmARM4 locus in a novel maize (Zea mays) grain-filling mutant. Plant Breeding 139(3):217–226
Kong Y, Pei S, Wang Y, Xu Y, Wang X, Zhou G, Hu R (2021) Homeodomain glabrous2 regulates cellulose biosynthesis in seed coat mucilage by activating cellulose synthase5. Plant Physiol 185(1):77–93
Koo HJ, Park SM, Kim KP, Suh MC, Lee MO, Lee SK, **nli X, Hong CB (2015) Small heat shock proteins can release light dependence of tobacco seed during germination. Plant Physiol 167(3):1030–1038
Kumar MN, Bau YC, Longkumer T, Verslues PE (2019) Low water potential and At14a-Like1 (AFL1) effects on endocytosis and actin filament organization. Plant Physiol 179(4):1594–1607
Lai CP, Huang LM, Chen LO, Chan MT, Shaw JF (2017) Genome-wide analysis of GDSL-type esterases/lipases in Arabidopsis. Plant Mol Biol 95(1–2):181–197
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinform 9:559
Lee S, Cheng H, King KE, Wang W, He Y, Hussain A, Lo J, Harberd NP, Peng J (2002) Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Genes Dev 16(5):646–658
Li Y, Sun N, Lu Z, Sun S, Huang J, Chen Z, He J (2017) Prognostic alternative mRNA splicing signature in non-small cell lung cancer. Cancer Lett 393:40–51
Li X, Sun M, Liu S, Teng Q, Li S, Jiang Y (2021) Functions of PPR proteins in plant growth and development. Int J Mol Sci 22(20):11274
Lin KF, Lee TR, Tsai PH, Hsu MP, Chen CS, Lyu PC (2007) Structure-based protein engineering for alpha-amylase inhibitory activity of plant defensin. Proteins 68(2):530–540
Liu J, Zhu JK (1998) A calcium sensor homolog required for plant salt tolerance. Science 280(5371):1943–1945
Liu X, Yue Y, Li B, Nie Y, Li W, Wu WH, Ma L (2007a) A G protein-coupled receptor is a plasma membrane receptor for the plant hormone abscisic acid. Science 315(5819):1712–1716
Liu Y, Koornneef M, Soppe WJJ (2007b) The absence of histone H2B monoubiquitination in the Arabidopsis hub1 (rdo4) mutant reveals a role for chromatin remodeling in seed dormancy. Plant Cell 19:433–444
Liu WC, Song RF, Zheng SQ, Li TT, Zhang BL, Gao X, Lu YT (2022) Coordination of plant growth and abiotic stress responses by tryptophan synthase β subunit 1 through modulation of tryptophan and ABA homeostasis in Arabidopsis. Mol Plant 15(6):973–990
Lubkowitz M (2011) The oligopeptide transporters: a small gene family with a diverse group of substrates and functions? Mol Plant 4(3):407–415
Ma J, Cao YY, Wang LF, Li JJ, Wang H, Fan YP, Li HY (2020) Identification of gene co-expression modules of maize plant height and ear height by WGCNA. Acta Agron Sin 46(3):385–394
Martinez SA, Godoy J, Huang M, Zhang Z, Carter AH, Garland Campbell KA, Steber CM (2018) Genome-wide association map** for tolerance to preharvest sprouting and low falling numbers in wheat. Front Plant Sci 9:141
Maruyama K, Urano K, Yoshiwara K, Morishita Y, Sakurai N, Suzuki H, Kojima M, Sakakibara H, Shibata D, Saito K, Shinozaki K, Yamaguchi-Shinozaki K (2014) Integrated analysis of the effects of cold and dehydration on rice metabolites, phytohormones, and gene transcripts. Plant Physiol 164(4):1759–1771
Mei S, Zhang M, Ye J, Du J, Jiang Y, Hu Y (2023) Auxin contributes to jasmonate-mediated regulation of abscisic acid signaling during seed germination in Arabidopsis. Plant Cell 35(3):1110–1133
Merilo E, Jalakas P, Laanemets K, Mohammadi O, Hõrak H, Kollist H, Brosché M (2015) Abscisic acid transport and homeostasis in the context of stomatal regulation. Mol Plant 8(9):1321–1333
Misra VA, Wafula EK, Wang Y, dePamphilis CW, Timko MP (2019) Genome-wide identification of MST, SUT and SWEET family sugar transporters in root parasitic angiosperms and analysis of their expression during host parasitism. BMC Plant Biol 19(1):196
Nagel M, Alqudah AM, Bailly M, Rajjou L, Pistrick S, Matzig G, Börner A, Kranner I (2019) Novel loci and a role for nitric oxide for seed dormancy and preharvest sprouting in barley. Plant Cell Environ 42(4):1318–1327
Nakabayashi K, Bartsch M, **ang Y, Miatton E, Pellengahr S, Yano R, Seo M, Soppe WJJ (2012) The time required for dormancy release in Arabidopsis is determined by delay of germination1 protein levels in freshly harvested seeds. Plant Cell 24(7):2826–2838
Niu L, Du C, Wang W, Zhang M, Wang W, Liu H, Zhang J, Wu X (2022) Transcriptome and co-expression network analyses of key genes and pathways associated with differential abscisic acid accumulation during maize seed maturation. BMC Plant Biol 22(1):359
Pandey GK, Cheong YH, Kim KN, Grant JJ, Li L, Hung W, D’Angelo C, Weinl S, Kudla J, Luan S (2004) The calcium sensor calcineurin B-like 9 modulates abscisic acid sensitivity and biosynthesis in Arabidopsis. Plant Cell 16(7):1912–1924
Park C, Lim CW, Baek W, Lee SC (2015) RING Type E3 Ligase CaAIR1 in pepper acts in the regulation of ABA signaling and drought stress response. Plant Cell Physiol 56(9):1808–1819
Pauly M, Albersheim P, Darvill A, York WS (1999) Molecular domains of the cellulose/xyloglucan network in the cell walls of higher plants. Plant J 20(6):629–639
Quettier AL, Eastmond PJ (2009) Storage oil hydrolysis during early seedling growth. Plant Physiol Biochem 47:485–490
Rao V, Petla BP, Verma P, Salvi P, Kamble NU, Ghosh S, Kaur H, Saxena SC, Majee M (2018) Arabidopsis SKP1-like protein13 (ASK13) positively regulates seed germination and seedling growth under abiotic stress. J Exp Bot 69(16):3899–3915
Rothbart SB, Strahl BD (2014) Interpreting the language of histone and DNA modifications. Biochim Biophys Acta 1839(8):627–643
Sahi C, Kominek J, Ziegelhoffer T, Yu HY, Baranowski M, Marszalek J, Craig EA (2013) Sequential duplications of an ancient member of the DnaJ-family expanded the functional chaperone network in the eukaryotic cytosol. Mol Biol Evol 30(5):985–998
Sahtoe DD, van Dijk WJ, El Oualid F, Ekkebus R, Ovaa H, Sixma TK (2015) Mechanism of UCH-L5 activation and inhibition by DEUBAD domains in RPN13 and INO80G. Mol Cell 57(5):887–900
Samtani H, Sharma A, Khurana P (2022) Overexpression of HVA1 enhances drought and heat stress tolerance in triticum aestivum doubled haploid plants. Cells 11(5):912
Sato K, Yamane M, Yamaji N, Kanamori H, Tagiri A, Schwerdt JG, Fincher GB, Matsumoto T, Takeda K, Komatsuda T (2016) Alanine aminotransferase controls seed dormancy in barley. Nat Commun 7:11625
Semrad K, Green R, Schroeder R (2004) RNA chaperone activity of large ribosomal subunit proteins from Escherichia coli. RNA 10(12):1855–1860
Shahpiri A, Svensson B, Finnie C (2008) The NADPH-dependent thioredoxin reductase/thioredoxin system in germinating barley seeds: gene expression, protein profiles, and interactions between isoforms of thioredoxin h and thioredoxin reductase. Plant Physiol 146(2):789–799
Shao Q, Liu X, Su T, Ma C, Wang P (2019) New insights into the role of seed oil body proteins in metabolism and plant development. Front Plant Sci 10:1568
Shen H, Luong P, Huq E (2007) The F-box protein MAX2 functions as a positive regulator of photomorphogenesis in Arabidopsis. Plant Physiol 145(4):1471–1483
Shi J, Shi J, Liang W, Zhang D (2021) Integrating GWAS and transcriptomics to identify genes involved in seed dormancy in rice. Theor Appl Genet 134(11):3553–3562
Shim JS, Park SH, Lee DK, Kim YS, Park SC, Redillas MC, Seo JS, Kim JK (2021) The rice Glycine-rich protein 3 confers drought tolerance by regulating mRNA stability of ROS scavenging-related genes. Rice (n y) 14(1):31
Shu K, Liu XD, **e Q, He ZH (2016) Two faces of one seed: hormonal regulation of dormancy and germination. Mol Plant 9(1):34–45
Sidorov V, Gilbertson L, Addae P, Duncan D (2006) Agrobacterium-mediated transformation of seedling-derived maize callus. Plant Cell Rep 25(4):320–328
Simsek S, Ohm JB, Lu H, Rugg M, Berzonsky W, Alamri MS, Mergoum M (2014) Effect of pre-harvest sprouting on physicochemical changes of proteins in wheat. J Sci Food Agric 94(2):205–212
Stroud DA, Surgenor EE, Formosa LE, Reljic B, Frazier AE, Dibley MG, Osellame LD, Stait T, Beilharz TH, Thorburn DR, Salim A, Ryan MT (2016) Accessory subunits are integral for assembly and function of human mitochondrial complex I. Nature 538(7623):123–126
Sun Y, Wu Z, Wang Y, Yang J, Wei G, Chou M (2019) Identification of phytocyanin gene family in legume plants and their involvement in Nodulation of Medicago truncatula. Plant Cell Physiol 60(4):900–915
Suzuki M, Kao CY, Cocciolone S, McCarty DR (2001) Maize VP1 complements Arabidopsis ABI3 and confers a novel ABA/auxin interaction in roots. Plant J 28(4):409–418
Suzuki M, Settles AM, Tseung CW, Li QB, Latshaw S, Wu S, Porch TG, Schmelz EA, James MG, McCarty DR (2006) The maize viviparous15 locus encodes the molybdopterin synthase small subunit. Plant J 45(2):264–274
Swain SM, Tseng TS, Olszewski NE (2001) Altered expression of SPINDLY affects gibberellin response and plant development. Plant Physiol 126(3):1174–1185
Tai L, Wang HJ, Xu XJ, Sun WH, Ju L, Liu WT, Li WQ, Sun JQ, Chen KM (2021) Pre-harvest sprouting in cereals: genetic and biochemical mechanisms. J Exp Bot 72(8):2857–2876
Vishnu Varthini L, Selvaraju K, Srinivasan M, Nachiappan V (2015) ROG1 encodes a monoacylglycerol lipase in Saccharomyces cerevisiae. FEBS Lett 589(1):23–30
Wang Z, Wang L, Liu Z, Li Y, Liu Q, Liu B (2016) Phylogeny, seed trait, and ecological correlates of seed germination at the community level in a degraded sandy grassland. Front Plant Sci 7:1532
Wang JJ, Lei JJ, Liang CL, Zhang ZK, Chang PP, Zhang ZY, He HJ (2019) Progress on the study of translationally controlled tumor protein in plants. J Nuc Agricul Sci 33(4):696–704
Watanabe M, Mochida K, Kato T, Tabata S, Yoshimoto N, Noji M, Saito K (2008) Comparative genomics and reverse genetics analysis reveal indispensable functions of the serine acetyltransferase gene family in Arabidopsis. Plant Cell 20(9):2484–2496
Waters MT, Scaffidi A, Flematti GR, Smith SM (2013) The origins and mechanisms of karrikin signalling. Curr Opin Plant Biol 16(5):667–673
Wolven AK, Belmont LD, Mahoney NM, Almo SC, Drubin DG (2000) In vivo importance of actin nucleotide exchange catalyzed by profilin. J Cell Biol 150(4):895–904
Wu CT, Bradford KJ (2003) Class I chitinase and beta-1,3-glucanase are differentially regulated by wounding, methyl jasmonate, ethylene, and gibberellin in tomato seeds and leaves. Plant Physiol 133(1):263–273
**ang Y, Nakabayashi K, Ding J, He F, Bentsink L, Soppe WJ (2014) Reduced dormancy5 encodes a protein phosphatase 2C that is required for seed dormancy in Arabidopsis. Plant Cell 26(11):4362–4375
Xu R (2003) Measuring explained variation in linear mixed effects models. Stat Med 22(22):3527–3541
Yang XY, Chen ZW, Xu T, Qu Z, Pan XD, Qin XH, Ren DT, Liu GQ (2011) Arabidopsis kinesin KP1 specifically interacts with VDAC3, a mitochondrial protein, and regulates respiration during seed germination at low temperature. Plant Cell 23(3):1093–1106
Yang N, Lu Y, Yang X, Huang J, Zhou Y, Ali F, Wen W, Liu J, Li J, Yan J (2014) Genome wide association studies using a new nonparametric model reveal the genetic architecture of 17 agronomic traits in an enlarged maize association panel. PLoS Genet 10(9):e1004573
Yang K, Zhu L, Wang H, Jiang M, **ao C, Hu X, Vanneste S, Dong J, Le J (2019) A conserved but plant-specific CDK-mediated regulation of DNA replication protein A2 in the precise control of stomatal terminal division. Proc Natl Acad Sci U S A 116(36):18126–18131
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association map** that accounts for multiple levels of relatedness. Nat Genet 38(2):203–208
Zhan JJ, Wang F, ** and candidate gene prediction of a major QTL for kernel number perear in maize. Mol Breeding 38:27
Zhang Y, Hu Y, Guan Z, Liu P, He Y, Zou C, Li P, Gao S, Peng H, Yang C, Pan G, Shen Y, Ma L (2020) Combined linkage map** and association analysis reveals genetic control of maize kernel moisture content. Physiol Plant 170(4):508–518
Zhang M, Cheng W, Yuan X, Wang J, Cheng T, Zhang Q (2022) Integrated transcriptome and small RNA sequencing in revealing miRNA-mediated regulatory network of floral bud break in Prunus mume. Front Plant Sci 13:931454
Zhao P, Liu RX, Li CP, ** for grain yield associated traits using ye478 introgression lines in maize. Sci Agric Sin 44(17):3508–3519
Zhao ML, Ni J, Chen MS, Xu ZF (2019) Ectopic expression of jatropha curcas Trehalose-6-Phosphate Phosphatase j causes late-flowering and heterostylous phenotypes in Arabidopsis but not in Jatropha. Int J Mol Sci 20(9):2165
Zhao H, Jan A, Ohama N, Kidokoro S, Soma F, Koizumi S, Mogami J, Todaka D, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2021) Cytosolic HSC70s repress heat stress tolerance and enhance seed germination under salt stress conditions. Plant Cell Environ 44(6):1788–1801
Zheng J, Chen F, Wang Z, Cao H, Li X, Deng X, Soppe WJJ, Li Y, Liu Y (2012) A novel role for histone methyltransferase KYP/SUVH4 in the control of Arabidopsis primary seed dormancy. New Phytol 193(3):605–616
Acknowledgements
This work was supported by State Key Laboratory of North China Crop Improvement and Regulation (NCCIR2020ZZ-6), Natural Science Foundation of Heibei Province of China (C2021204078), and Hebei Key R&D Plan “Special project on scientific and technological innovation of modern seed industry” (21326328D). We thank Prof. Jianbing Yan of Huazhong Agricultural University for providing seeds of the 368 natural maize populations.
Funding
This study was funded by the State Key Laboratory of North China Crop Improvement and Regulation (NCCIR2020ZZ-6), Natural Science Foundation of Heibei Province of China (C2021204078), and Hebei Key R&D Plan “Special project on scientific and technological innovation of modern seed industry” (21326328D).
Author information
Authors and Affiliations
Contributions
YT and HD designed the experiment. XM, AT, LF, YH, and DZ collected the phenotypes and performed the data analysis. YT and HD wrote the manuscript. XM, AT, LF, TZ, YH, and DZ edited the manuscript. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationship that could be construed as a potential conflict of interest.
Human and animal rights
This study does not include human or animal subjects.
Additional information
Communicated by Thomas Lubberstedt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ma, X., Feng, L., Tao, A. et al. Identification and validation of seed dormancy loci and candidate genes and construction of regulatory networks by WGCNA in maize introgression lines. Theor Appl Genet 136, 259 (2023). https://doi.org/10.1007/s00122-023-04495-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04495-8