Abstract
Key message
QTL for 15 agronomic traits under two levels of salt stress in dry salinity field were mapped in a new constructed RIL population utilizing a Wheat55K SNP array. Furthermore, eight QTL were validated in a collected natural population.
Abstract
Soil salinity is one of the major abiotic stresses causing serious impact on crop growth, development and yield. As one of the three most important crops in the world, bread wheat (Triticum aestivum L.) is severely affected by salinity, too. In this study, an F7 recombinant inbred line (RIL) population derived from a cross between high-yield wheat cultivar Zhongmai 175 and salt-tolerant cultivar ** showed that 90 stable QTL for 15 traits were detected, and they were distributed on all wheat chromosomes except 4D, 6B and 7D. These QTL individually explained 2.34–32.43% of the phenotypic variation with LOD values ranging from 2.68 to 47.15. It was found that four QTL clusters were located on chromosomes 2D, 3D, 4B and 6A, respectively. Notably, eight QTL from the QTL clusters were validated in a collected natural population. Among them, QPh-4B was deduced to be an allele of Rht-B1. In addition, three kompetitive allele-specific PCR (KASP) markers derived from SNPs were successfully designed for three QTL clusters. This study provides an important base for salt-tolerant QTL (gene) cloning in wheat, and the markers, especially the KASP markers, will be useful for marker-assisted selection in salt-tolerant wheat breeding.



Similar content being viewed by others
References
Asif MA, Schilling RK, Tilbrook J, Brien C, Dowling K, Rabie H, Short L, Trittermann C, Garcia A, Barrett-Lennard EG, Berger B, Mather DE, Gilliham M, Fleury D, Tester M, Roy SJ, Pearson AS (2018) Map** of novel salt tolerance QTL in an Excalibur × Kukri doubled haploid wheat population. Theor Appl Genet 131:2179–2196
Azadi A, Mardi M, Hervan EM, Mohammadi SA, Moradi F, Tabatabaee MT, Pirseyedi SM, Ebrahimi M, Fayaz F, Kazemi M, Ashkani S, Nakhoda B, Mohammadi-Nejad G (2015) QTL map** of yield and yield components under normal and salt-stress conditions in bread wheat (Triticum aestivum L.). Plant Mol Biol Rep 33:102–120
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, Forrest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D, Bai G, Pumphrey M, Tomar L, Wong D, Kong S, Reynolds M, da Silva ML, Bockelman H, Talbert L, Anderson JA, Dreisigacker S, Baenziger S, Carter A, Korzun V, Morrell PL, Dubcovsky J, Morell MK, Sorrells ME, Hayden MJ, Akhunov E (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Natl Acad Sci 110:8057–8062
Chai L, Chen Z, Bian R, Zhai H, Cheng X, Peng H, Yao Y, Hu Z, **n M, Guo W, Sun Q, Zhao A, Ni Z (2018) Dissection of two quantitative trait loci with pleiotropic effects on plant height and spike length linked in coupling phase on the short arm of chromosome 2D of common wheat (Triticum aestivum L.). Theor Appl Genet 131:2621–2637
Cui F, Zhao C, Ding A, Li J, Wang L, Li X, Bao Y, Li J, Wang H (2014) Construction of an integrative linkage map and QTL map** of grain yield-related traits using three related wheat RIL populations. Theor Appl Genet 127:659–675
Cui F, Fan X, Chen M, Zhang N, Zhao C, Zhang W, Han J, Ji J, Zhao X, Yang L, Zhao Z, Tong Y, Wang T, Li J (2016) QTL detection for wheat kernel size and quality and the responses of these traits to low nitrogen stress. Theor Appl Genet 129:469–484
Cui F, Zhang N, Fan XL, Zhang W, Zhao CH, Yang LJ, Pan RQ, Chen M, Han J, Zhao XQ, Ji J, Tong YP, Zhang HX, Jia JZ, Zhao GY, Li JM (2017) Utilization of a Wheat660K SNP array-derived high-density genetic map for high-resolution map** of a major QTL for kernel number. Sci Rep 7:3788
Devi R, Ram S, Rana V, Malik VK, Pande V, Singh GP (2019) QTL map** for salt tolerance associated traits in wheat (Triticum aestivum L.). Euphytica 215:210
Ellis MH, Spielmeyer W, Gale KR, Rebetzke GJ, Richards RA (2002) "Perfect" markers for the Rht-B1b and Rht-D1b dwarfing genes in wheat. Theor Appl Genet 105:1038–1042
Gao F, Wen W, Liu J, Rasheed A, Yin G, ** of QTL for yield components, plant height and yield-related physiological traits in the Chinese wheat cross Zhou 8425B/Chinese Spring. Front Plant Sci 6:1099
Gasperini D, Greenland A, Hedden P, Dreos R, Harwood W, Griffiths S (2012) Genetic and physiological analysis of Rht8 in bread wheat: an alternative source of semidwarfism with a reduced sensitivity to brassinosteroids. J Exp Bot 63:4419–4436
Genc Y, Oldach K, Verbyla AP, Lott G, Hassan M, Tester M, Wallwork H, McDonald GK (2010) Sodium exclusion QTL associated with improved seedling growth in bread wheat under salinity stress. Theor Appl Genet 121:877–894
Genc Y, Oldach K, Gogel B, Wallwork H, McDonald GK, Smith AB (2013) Quantitative trait loci for agronomic and physiological traits for a bread wheat population grown in environments with a range of salinity levels. Mol Breed 32:39–59
Ghaedrahmati M, Mardi M, Naghavi MR, Haravan EM, Nakhoda B, Azadi A, Kazemi M (2014) Map** QTLs associated with salt tolerance related traits in seedling stage of wheat (Triticum aestivum L.). J Agric Sci Technol 16:1413–1428
Guan P, Lu L, Jia L, Kabir MR, Zhang J, Lan T, Zhao Y, **n M, Hu Z, Yao Y, Ni Z, Sun Q, Peng H (2018) Global QTL analysis identifies genomic regions on chromosomes 4A and 4B harboring stable loci for yield-related traits across different environments in wheat (Triticum aestivum L.). Front Plant Sci 9:529
Hong Y, Chen L, Du L-p, Su Z, Wang J, Ye X, Qi L, Zhang Z (2014) Transcript suppression of TaGW2 increased grain width and weight in bread wheat. Funct Integr Genom 14:341–349
Huang XQ, Cloutier S, Lycar L, Radovanovic N, Humphreys DG, Noll JS, Somers DJ, Brown PD (2006) Molecular detection of QTLs for agronomic and quality traits in a doubled haploid population derived from two Canadian wheats (Triticum aestivum L.). Theor Appl Genet 113:753–766
Huang Y, Kong Z, Wu X, Cheng R, Yu D, Ma Z (2015) Characterization of three wheat grain weight QTLs that differentially affect kernel dimensions. Theor Appl Genet 128:2437–2445
Hussain B, Lucas SJ, Ozturk L, Budak H (2017) Map** QTLs conferring salt tolerance and micronutrient concentrations at seedling stage in wheat. Sci Rep 7:15662
International Wheat Genome Sequencing C (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
International Wheat Genome Sequencing C (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:661–674
Kosambi DD (1943) The estimation of map distances from recombination values. Ann Eugen 12:172–175
León JLDD, Escoppinichi R, Geraldo N, Castellanos T, Mujeeb-Kazi A, Röder MS (2011) Quantitative trait loci associated with salinity tolerance in field grown bread wheat. Euphytica 181:371–383
Li FJ, Wen WE, He ZH, Liu JD, ** H, Cao SH, Geng HW, Yan J, Zhang PZ, Wan YX, ** of yield-related traits in three Chinese bread wheat populations using high-density SNP markers. Theor Appl Genet 131:1903–1924
Liu J, Luo W, Qin N, Ding P, Zhang H, Yang C, Mu Y, Tang H, Liu Y, Li W, Jiang Q, Chen G, Wei Y, Zheng Y, Liu C, Lan X, Ma J (2018) A 55 K SNP array-based genetic map and its utilization in QTL map** for productive tiller number in common wheat. Theor Appl Genet 131:2439–2450
Luo Q, Teng W, Fang S, Li H, Li B, Chu J, Li Z, Zheng Q (2019) Transcriptome analysis of salt-stress response in three seedling tissues of common wheat. Crop J 7:378–392
Ma LQ, Zhou EF, Huo NX, Zhou RH, Wang GY, Jia JZ (2007) Genetic analysis of salt tolerance in a recombinant inbred population of wheat (Triticum aestivum L.). Euphytica 153:109–117
Masoudi B, Mardi M, Hervan EM, Bihamta MR, Naghavi MR, Nakhoda B, Amini A (2015) QTL map** of salt tolerance traits with different effects at the seedling stage of bread wheat. Plant Mol Biol Rep 33:1790–1803
McCartney CA, Somers DJ, Humphreys DG, Lukow O, Ames N, Noll J, Cloutier S, McCallum BD (2005) Map** quantitative trait loci controlling agronomic traits in the spring wheat cross RL4452 × 'AC Domain'. Genome 48:870–883
Mo Y, Vanzetti LS, Hale I, Spagnolo EJ, Guidobaldi F, Al-Oboudi J, Odle N, Pearce S, Helguera M, Dubcovsky J (2018) Identification and characterization of Rht25, a locus on chromosome arm 6AS affecting wheat plant height, heading time, and spike development. Theor Appl Genet 131:2021–2035
Munns R, James RA (2003) Screening methods for salinity tolerance: a case study with tetraploid wheat. Plant Soil 253:201–218
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, Byrt CS, Hare RA, Tyerman SD, Tester M (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30:360–364
Narjesi V, Mardi M, Hervan EM, Azadi A, Naghavi MR, Ebrahimi M, Zali AA (2015) Analysis of quantitative trait loci (QTL) for grain yield and agronomic traits in wheat (Triticum aestivum L.) under normal and salt-stress conditions. Plant Mol Biol Rep 33:2030–2040
Mahdi Nezhad N, Jalal Kamali MR, McIntyre CL, Fakheri BA, Omidi M, Masoudi B (2019) Map** QTLs with main and epistatic effect on Seri "M82xBabax" wheat population under salt stress. Euphytica 215:1
Ogbonnaya FC, Rasheed A, Okechukwu EC, Jighly A, Makdis F, Wuletaw T, Hagras A, Uguru MI, Agbo CU (2017) Genome-wide association study for agronomic and physiological traits in spring wheat evaluated in a range of heat prone environments. Theor Appl Genet 130:1819–1835
Oyiga BC, Sharma RC, Baum M, Ogbonnaya FC, Leon J, Ballvora A (2018) Allelic variations and differential expressions detected at quantitative trait loci for salt stress tolerance in wheat. Plant Cell Environ 41:919–935
Parida SK, Jaiswal V, Gahlaut V, Mathur S, Agarwal P, Khandelwal MK, Khurana JP, Tyagi AK, Balyan HS, Gupta PK (2015) Identification of novel SNP in promoter sequence of TaGW2-6A associated with grain weight and other agronomic traits in wheat (Triticum aestivum L.). PLoS ONE 10:e0129400
Qin L, Hao C, Hou J, Wang Y, Li T, Wang L, Ma Z, Zhang X (2014) Homologous haplotypes, expression, genetic effects and geographic distribution of the wheat yield gene TaGW2. BMC Plant Biol 14:1–19
Quarrie SA, Steed A, Calestani C, Semikhodskii A, Lebreton C, Chinoy C, Steele N, Pljevljakusic D, Waterman E, Weyen J, Schondelmaier J, Habash DZ, Farmer P, Saker L, Clarkson DT, Abugalieva A, Yessimbekova M, Turuspekov Y, Abugalieva S, Tuberosa R, Sanguineti MC, Hollington PA, Aragues R, Royo A, Dodig D (2005) A high-density genetic map of hexaploid wheat (Triticum aestivum L.) from the cross Chinese Spring × SQ1 and its use to compare QTLs for grain yield across a range of environments. Theor Appl Genet 110:865–880
Ren T, Hu Y, Tang Y, Li C, Yan B, Ren Z, Tan F, Tang Z, Fu S, Li Z (2018) Utilization of a Wheat55K SNP array for map** of major QTL for temporal expression of the tiller number. Front Plant Sci 9:333
Roy SJ, Negrao S, Tester M (2014) Salt resistant crop plants. Curr Opin Biotechnol 26:115–124
Shi W, Hao C, Zhang Y, Cheng J, Zhang Z, Liu J, Yi X, Cheng X, Sun D, Xu Y, Zhang X, Cheng S, Guo P, Guo J (2017) A combined association map** and linkage analysis of kernel number per spike in common wheat (Triticum aestivum L.). Front Plant Sci 8:1412
Su Z, Hao C, Wang L, Dong Y, Zhang X (2011) Identification and development of a functional marker of TaGW2 associated with grain weight in bread wheat (Triticum aestivum L.). Theor Appl Genet 122:211–223
Su QN, Zhang XL, Zhang W, Zhang N, Song LQ, Liu L, Xue X, Liu GT, Liu JJ, Meng DY, Zhi LY, Ji J, Zhao XQ, Yang CL, Tong YP, Liu ZY, Li JM (2018) QTL detection for kernel size and weight in bread wheat (Triticum aestivum L.) using a high-density SNP and SSR-based linkage map. Front Plant Sci 9:13
Sukumaran S, Dreisigacker S, Lopes M, Chavez P, Reynolds MP (2015) Genome-wide association study for grain yield and related traits in an elite spring wheat population grown in temperate irrigated environments. Theor Appl Genet 128:353–363
Sukumaran S, Lopes M, Dreisigacker S, Reynolds M (2018) Genetic analysis of multi-environmental spring wheat trials identifies genomic regions for locus-specific trade-offs for grain weight and grain number. Theor Appl Genet 131:985–998
Sun C, Zhang F, Yan X, Zhang X, Dong Z, Cui D, Chen F (2017) Genome-wide association study for 13 agronomic traits reveals distribution of superior alleles in bread wheat from the Yellow and Huai Valley of China. Plant Biotechnol J 15:953–969
Tian X, Wen W, ** of reduced plant height gene Rht24 in bread wheat. Front Plant Sci 8:1379
Wang M, **a G (2017) The landscape of molecular mechanisms for salt tolerance in wheat. Crop J 6:42–47
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Whan A, Stephen S, Barker G, Wieseke R, Plieske J, Lillemo M, Mather D, Appels R, Dolferus R, Brown-Guedira G, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo M-C, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden M, Akhunov E, Int Wheat Genome S (2014) Characterization of polyploid wheat genomic diversity using a high-density 90000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796
Winfield MO, Allen AM, Burridge AJ, Barker GL, Benbow HR, Wilkinson PA, Coghill J, Waterfall C, Davassi A, Scopes G, Pirani A, Webster T, Brew F, Bloor C, King J, West C, Griffiths S, King I, Bentley AR, Edwards KJ (2016) High-density SNP genoty** array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol J 14:1195–1206
Wu X, Cheng R, Xue S, Kong Z, Wan H, Li G, Huang Y, Jia H, Jia J, Zhang L, Ma Z (2014) Precise map** of a quantitative trait locus interval for spike length and grain weight in bread wheat (Triticum aestivum L.). Mol Breed 33:129–138
Wurschum T, Langer SM, Longin CFH, Tucker MR, Leiser WL (2017) A modern green revolution gene for reduced height in wheat. Plant J 92:892–903
Xu Y, An D, Liu D, Zhang A, Xu H, Li B (2012) Map** QTLs with epistatic effects and QTL × treatment interactions for salt tolerance at seedling stage of wheat. Euphytica 186:233–245
Xu Y, Li S, Li L, Zhang X, Xu H, An D (2013) Map** QTLs for salt tolerance with additive, epistatic and QTL × treatment interaction effects at seedling stage in wheat. Plant Breed 132:276–283
Xu Y, Wang R, Tong Y, Zhao H, ** QTLs for yield and nitrogen-related traits in wheat: influence of nitrogen and phosphorus fertilization on QTL expression. Theor Appl Genet 127:59–72
Xu D, Wen W, Fu L, Li F, Li J, **e L, **a X, Ni Z, He Z, Cao S (2019) Genetic dissection of a major QTL for kernel weight spanning the Rht-B1 locus in bread wheat. Theor Appl Genet 132:3191–3200
Yang S, Zhang X, He Z, **a X, Zhou Y (2006) Distribution of dwarfing genes Rht-B1b and Rht-D1b in Chinese bread wheats detected by STS marker. Scientica Agricultura Sinica 39:1680–1688
Yang Z, Bai Z, Li X, Wang P, Wu Q, Yang L, Li L, Li X (2012) SNP identification and allelic-specific PCR markers development for TaGW2, a gene linked to wheat kernel weight. Theor Appl Genet 125:1057–1068
Zhai H, Feng Z, Li J, Liu X, **ao S, Ni Z, Sun Q (2016) QTL Analysis of spike morphological traits and plant height in winter wheat (Triticum aestivum L.) using a high-density SNP and SSR-based linkage map. Front Plant Sci 7:1617
Zhang X, Yang S, Zhou Y, He Z, **a X (2006) Distribution of the Rht-B1b, Rht-D1b and Rht8 reduced height genes in autumn-sown Chinese wheats detected by molecular markers. Euphytica 152:109–116
Zhang N, Fan X, Cui F, Zhao C, Zhang W, Zhao X, Yang L, Pan R, Chen M, Han J, Ji J, Liu D, Zhao Z, Tong Y, Zhang A, Wang T, Li J (2017) Characterization of the temporal and spatial expression of wheat (Triticum aestivum L.) plant height at the QTL level and their influence on yield-related traits. Theor Appl Genet 130:1235–1252
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324
Zimin AV, Puiu D, Hall R, Kingan S, Clavijo BJ, Salzberg SL (2017) The first near-complete assembly of the hexaploid bread wheat genome, Triticum aestivum. Gigascience 6:1–7
Acknowledgements
We particularly thank Dr. Yunfeng Xu for his generous help in genetic linkage map construction. This study was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (No. XDA24030302 and XDA08030105).
Author information
Authors and Affiliations
Contributions
ZSL and QZ supervised the research. QLL and QZ designed the experiment. QLL, QZ and PH performed the phenotypes of the RIL population. QLL created the RIL population, performed data analyses and QTL map**, confirmed the QTL effects and wrote the manuscript. QZ also put forward many constructive suggestions and revised the manuscript. LQL, GTY, HWL and BL provided a lot of help in the materials preparation and experiments performance. All authors read and approved the final manuscript.
Corresponding author
Additional information
Communicated by Peter Langridge.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
The frequency distribution histogram for all 15 traits and the parents positions among RILs under HS and LS in three years. All data were derived from the averages of three years. The first three rows were under HS, and the last three rows were under LS. (PDF 640 kb)
Fig. S2
Comparison of the genetic and physical positions of the SNPs on the genetic map. The bars in green indicate the percentage of markers whose physical and genetic positions are coincident (identical chromosome); yellow indicates the percentage of markers whose physical and genetic chromosomes are homoeologous; red indicates the percentage of markers whose physical and genetic positions are inconsistent (in disorder); and purple indicates the percentage of markers whose physical positions are unknown. The physical positions are from the Chinese Spring contigs which were the best hits for the corresponding markers. (PDF 345 kb)
Fig. S3
Syntenic relationships between the genetic and physical maps of a total of 16,008 SNP markers used in this study. Gen-1A to Gen-7D represent the 21 wheat chromosomal genetics maps released in this paper; Phy-1A to Phy-7D represent the 21 wheat chromosomal physical maps (IWGSC RefSeq v1.0). (TIF 7451 kb)
Fig. S4
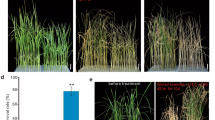
Confirmation of the QTL effect in the RIL population. Comparison of the QTL corresponding traits between two groups with different alleles derived from RIL parents ZM175 and XY60. SNP markers used in validation were shown as chart titles. Y-axis titles were the corresponding traits for the validated QTL. Error bars indicate the standard deviation. *, significant difference determined by the Student’s t test (P <0.05); **, significant difference determined by the Student’s t test (P <0.01). (PDF 548 kb)
Fig. S5
Genoty** of 24 RILs with the KASP markers developed in this study. The genotypes obtained using KASP markers were coincided with the genoty** results on the Wheat55K SNP array. (PDF 379 kb)
Fig. S6
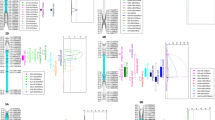
QSl-7A was detected in both RIL and natural populations. (a) QSl-7A involved SNP markers and the LOD curves in the RIL population. SNPs tightly linked to QSl-7A in purple were near to the significantly associated SNPs by GWAS. (b) Quantile–quantile (Q-Q) plot of SL in the natural population. (C) Manhattan plot for SL on chromosome 7A under LS with mixed linear model (MLM) by TASSEL 5.0 (Bradbury et al. 2007). Two vertical black lines showed the position of the SNP markers which were significantly associated with QSl-7A in the natural population. (PDF 361 kb)
Rights and permissions
About this article
Cite this article
Luo, Q., Zheng, Q., Hu, P. et al. Map** QTL for agronomic traits under two levels of salt stress in a new constructed RIL wheat population. Theor Appl Genet 134, 171–189 (2021). https://doi.org/10.1007/s00122-020-03689-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03689-8