Abstract
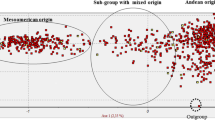
This study focuses on the expansion of Phaseolus vulgaris in Europe. The pathways of distribution of beans into and across Europe were very complex, with several introductions from the New World that were combined with direct exchanges between European and other Mediterranean countries. We have analyzed here six chloroplast microsatellite (cpSSR) loci and two unlinked nuclear loci (for phaseolin types and Pv-shatterproof1). We have assessed the genetic structure and level of diversity of a large collection of European landraces of P. vulgaris (307) in comparison to 94 genotypes from the Americas that are representative of the Andean and Mesoamerican gene pools. First, we show that most of the European common bean landraces (67%) are of Andean origin, and that there are no strong differences across European regions for the proportions of the Andean and Mesoamerican gene pools. Moreover, cytoplasmic diversity is evenly distributed across European regions. Secondly, the cytoplasmic bottleneck that was due to the introduction of P. vulgaris into the Old World was very weak or nearly absent. This is in contrast to evidence from nuclear analyses that have suggested a bottleneck of greater intensity. Finally, we estimate that a relatively high proportion of the European bean germplasm (about 44%) was derived from hybridization between the Andean and Mesoamerican gene pools. Moreover, although hybrids are present everywhere in Europe, they show an uneven distribution, with high frequencies in central Europe, and low frequencies in Spain and Italy. On the basis of these data, we suggest that the entire European continent and not only some of the countries therein can be regarded as a secondary diversification center for P. vulgaris. Finally, we outline the relevance of these inter-gene pool hybrids for plant breeding.




Similar content being viewed by others
References
Acosta-Gallegos JA, Kelly JD, Gepts P (2007) Prebreeding in common bean and use of genetic diversity from wild germplasm. Crop Sci 47(S3):S44–S59
Angioi SA, Rau D, Rodriguez M, Logozzo G, Desiderio F, Papa R, Attene G (2009a) Nuclear and chloroplast microsatellite diversity in Phaseolus vulgaris L. from Sardinia (Italy). Mol Breed 23:413–429
Angioi SA, Desiderio F, Rau D, Bitocchi E, Attene G, Papa R (2009b) Development and use of chloroplast microsatellites in Phaseolus spp. and other legumes. Plant Biol 11:598–612
Arroyo-Garcia R, Lefort F, de Andrés F, Ibanez J, Borrego J, Jouve N, Cabello F, Martìnez-Zapater JM (2002) Chloroplast microsatellite polymorphisms in Vitis species. Genome 45:1142–1149
Bassam BJ, Caetano-Anolles G, Gresshoff PM (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196:80–83
Beebe S, Skroch PW, Tohme J, Duque MC, Pedraza F, Nienhuis J (2000) Structure of genetic diversity between common bean landraces of Middle American origin based on correspondence analysis of RAPD. Crop Sci 40:264–273
Beebe S, Rengifo J, Gaitan E, Duque MC, Tohme J (2001) Diversity and origin of Andean landraces of common bean. Crop Sci 41:854–862
Berglund-Brücher B, Brücher H (1976) The South American wild bean (Phaseolus aborigenus Burk.) as an ancestor of the common bean. Econ Bot 30:257–272
Blair MW, Iriarte G, Beebe S (2006) QTL analysis of yield traits in an advanced backcross population derived from a cultivated Andean × wild common bean (Phaseolus vulgaris L.) cross. Theor Appl Genet 112:1149–1163
Chung S, Staub JE (2003) The development and evaluation of consensus chloroplast primer pairs that possess highly variable sequence regions in a diverse array of plant taxa. Theor Appl Genet 107:757–767
Cuenca A, Escalante E, Pinero D (2003) Long-distance colonization, isolation by distance, and historical demography in a relictual Mexican pinyon pine (Pinus nelsonii Shaw) as revealed by paternally inherited genetic markers (cpSSRs). Mol Ecol 12:2087–2097
El Mousadik A, Petit RJ (1996) High level of genetic differentiation for allelic richness among populations of the argan tree [Argania spinosa (L.) Skeels] endemic to Morocco. Theor Appl Genet 92:832–839
Elbert D, Peakall R (2009) Chloroplast simple sequence repeats (cpSSRs): technical resources and recommendations for expanding cpSSR discovery and applications to a wide array of plant species. Mol Ecol Resour 9:673–690
Ennos RA, Sinclair WT, Hu XS, Langdon A (1999) Using organelle markers to elucidate the history, ecology and evolution of plant populations. In: Hollingsworth PM, Bateman RM, Gornall RJ (eds) Molecular systematics and plant evolution. Taylor & Francis, London, pp 1–19
Escribano MR, Santalla M, Casquero PA, De Ron AR (1998) Patterns of genetic diversity in landraces of common bean (Phaseolus vulgaris L.) from Galicia. Plant Breed 117:49–56
Gepts P (1998) Origin and evolution of common bean: past events and recent trends. Hort Sci 33:1121–1130
Gepts P (1999) Development of an integrated genetic linkage map in common bean (Phaseolus vulgaris L.) and its use. In: Singh S (ed) Bean breeding for the 21st century, 53–91. Kluwer, Dordrecht, pp 389–400
Gepts P, Bliss FA (1985) F1 hybrid weakness in the common bean: differential geographic origin suggests two gene pools in cultivated bean germplasm. J Hered 76(6):447–450
Gepts P, Bliss FA (1988) Dissemination pathways of common bean (Phaseolus vulgaris, Fabaceae) deduced from phaseolin electrophoretic variability. II Europe and Africa. Econ Bot 42:86–104
Gepts P, Debouck DG (1991) Origin, domestication and evolution of common bean, Phaseolus vulgaris. In: Schoonhoven A, Voysest O (eds) Common beans: research for crop improvement. CAB International, Wallingford, UK, pp 4–54
Gepts P, Osborne TC, Rashka K, Bliss FA (1986) Electrophoretic analysis of phaseolin protein variability in wild forms and landraces of the common bean Phaseolus vulgaris L.: evidence for two centers of domestications. Econ Bot 40:451–468
Gepts P, Kmieciz K, Pereira P, Bliss FA (1988) Dissemination pathways of common bean (Phaseolus vulgaris, Fabaceae) deduced from phaseolin electrophoretic variability. I The Americas. Econ Bot 42:86–104
Gepts P, Papa R, Gonzales Mejia J, Acosta J, Delgado Salinas A (1999) Human effects on Phaseolus adaptation during and after domestication. In: van Raamsdonk L, den Njis J (eds) Proceeding VIIth international IOPB symposium. Amsterdam, 10–15 August
González AM, Rodiño AP, Santalla M, De Ron AM (2009) Genetics of intra-gene pool and inter-gene pool hybridization for seed traits in common bean (Phaseolus vulgaris L.) germplasm from Europe. Field Crop Res 112:66–76
Goudet J (2002) FSTAT: a program to estimate and test gene diversities and fixation indices (version 2.9.3.2). http://www.unil.ch/izea/softwares/fstat.html
Goudet J, Raymond M, de-Meeus T, Rousset F (1996) Testing differentiation in diploid populations. Genetics 144:1933–1940
Hale ML, Borland AM, Gustafsson MH, Wolff K (2004) Causes of size homoplasy among chloroplast microsatellites in closely related Clusia species. J Mol Evol 58:182–190
Hurlbert SH (1971) The non-concept of species diversity: a critique and alternative parameters. Ecology 52:577–586
Ishii T, McCouch SR (2000) Microsatellites and microsynteny in the chloroplast genomes of Oryza and eight other Gramineae species. Theor Appl Genet 100:1257–1266
Johnson WC, Gepts P (1999) Segregation for performance in recombinant inbred populations resulting from inter-gene pool crosses of common bean (Phaseolus vulgaris L.). Euphytica 106:45–56
Johnson WC, Gepts P (2002) The role of epistasis in controlling seed yield and other agronomic traits in an Andean × Mesoamerican cross of common bean (Phaseolus vulgaris L.). Euphytica 125:69–79
Koenig R, Gepts P (1989) Allozyme diversity in wild Phaseolus vulgaris: further evidence for two major centers of genetic diversity. Theor Appl Genet 78:809–817
Kwak M, Gepts P (2009) Structure of genetic diversity in the two major gene pools of common bean (Phaseolus vulgaris L., Fabaceae). Theor Appl Genet 118:979–992
Limongelli G, Laghetti G, Perrino P, Piergiovanni AR (1996) Variation of seed storage protein in landraces of common bean (Phaseolus vulgaris L.) from Basilicata, southern Italy. Plant Breed 119:513–516
Lioi L (1989) Geographical variation of phaseolin patterns in an old world collection of Phaseolus vulgaris. Seed Sci Technol 17:317–324
Logozzo G, Donnoli R, Macaluso L, Papa R, Knupffer H, Spagnoletti Zeuli PL (2007) Analysis of the contribution of Mesoamerican and Andean gene pools to European common bean (Phaseolus vulgaris L.) germplasm and strategies to establish a core collection. Genet Resour Crop Evol 54:1763–1779
Luikart G, Sherwin WB, Steele BM, Allendorf FW (1998a) Usefulness of molecular markers for detecting population bottlenecks via monitoring genetic change. Mol Ecol 7:963–974
Luikart G, Allendorf FW, Cornuet JM, Sherwin WB (1998b) Distortion of allele frequency distributions provide a test for recent population bottlenecks. J Hered 89:238–247
McClean PE, Lee RK, Otto C, Gepts P, Bassett MJ (2002) Molecular and phenotypic map** of genes controlling seed coat pattern and color in common bean (Phaseolus vulgaris L.). J Hered 93:148–152
McClean PE, Lee RK, Miklas PN (2004) Sequence diversity analysis of dihydroflavonol 4-reductase intron 1 in common bean. Genome 47:266–280
Miklas PN, Kelly JD, Beebe SE, Blair MW (2006) Common bean breeding for resistance against biotic and abiotic stresses: from classical to MAS breeding. Euphytica 147:106–131
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Nodari RO, Tsai SM, Gilbertson RL, Gepts P (1993) Towards an integrated linkage map of common bean. II. Development of an RFLP-based linkage map. Theor Appl Genet 85:513–520
Ocampo CH, Martin JP, Ortiz JM, Sanchez-Yelamo MD, Toro O, Debouck D (2002) Possible origins of common bean (Phaseolus vulgaris L.) cultivated in Spain in relation to the wild genetic pools of Americas. Annu Rep Bean Improv Coop 45:236–237
Ortwin-Sauer C (1966) The early Spanish man. University of California Press, Berkeley, pp 51–298
Papa R, Gepts P (2003) Asymmetry of gene flow and differential geographical structure of molecular diversity in wild and domesticated common bean (Phaseolus vulgaris L.) from Mesoamerica. Theor Appl Genet 106:239–250
Papa R, Nanni L, Sicard D, Rau D, Attene G (2006) The evolution of genetic diversity in Phaseolus vulgaris L. In: Motley TJ, Zerega N, Cross H (eds) New approaches to the origins, evolution and conservation of crops. Darwin’s harvest. Columbia University Press, USA
Paredes OM, Gepts P (1995) Extensive introgression of Middle American germplasm into Chilean common bean cultivars. Genet Resour Crop Evol 42:29–41
Park S, Michaels T, Myers J, Hunt D, Stewart-Williams K (1996) Outcrossing rates of common beans grow in Ontario and Idaho. Annu Rep Bean Improv Coop 39:90–91
Petit R, El Mousadik A, Pons O (1998) Identifying populations for conservation on the basis of genetic markers. Conserv Biol 12:844–855
Petit RJ, Dumilin J, Fineschi S, Hampe A, Salvini D, Vendramin GG (2005) Comparative organization of chloroplast, mitochondrial and nuclear diversity in plant populations. Mol Ecol 14:689–701
Piergiovanni AR, Cerbino D, Brandi M (2000a) The common bean populations from Basilicata (southern Italy). An evaluation of their variation. Genet Resour Crop Evol 47:489–495
Piergiovanni AR, Taranto G, Pignone D (2000b) Diversity among common bean populations from the Abruzzo region (central Italy): a preliminary inquiry. Genet Resour Crop Evol 47:467–470
Powell W, Morgante M, McDevitt R, Vendramin G, Rafalski J (1995) Polymorphic simple-sequence repeat regions in chloroplast genomes: applications to the population genetics of pines. Proc Natl Acad Sci USA 92:7759–7763
Provan J, Russell JR, Booth A, Powell W (1999a) Polymorphic chloroplast simple-sequence repeat primers for systematic and population studies in the genus Hordeum. Mol Ecol 8:505–511
Provan J, Powell W, Dewar H, Bryan G, Machray GC, Waugh R (1999b) An extreme cytoplasmic bottleneck in the modern European cultivated potato (Solanum tuberosum) is not reflected in decreased levels of nuclear diversity. Proc Biol Sci 266:633–639
Rebourg C, Chastanet M, Gouesnard B, Welcker C, Dubreuil P, Charcosset A (2003) Maize introduction into Europe: the history reviewed in the light of molecular data. Theor Appl Genet 106:895–903
Rodiño AP, Santalla M, Montero I, Casquero PA, De Ron AM (2001) Diversity in common bean (Phaseolus vulgaris L.) germplasm from Portugal. Genet Resour Crop Evol 48:409–417
Rodiño AP, Santalla M, De Ron AM, Singh SP (2003) A core collection of common bean from the Iberian peninsula. Euphytica 131:165–175
Rodiño AP, Santalla M, Gonzàlez AM, De Ron AM, Singh SP (2006) Novel genetic variation in common bean from the Iberian peninsula. Crop Sci 46:2540–2546
Rohlf F (2000) NTSYS pc. Numerical taxonomy system exeter publishing, Setauket
Romero J, Sun SM, McLeester RC, Bliss FA, Hall TC (1975) Heritable variation in a polypeptide subunit of the major storage protein of the bean, Phaseolus vulgaris L. Plant Physiol 56:776–779
Rossi M, Bitocchi E, Bellucci E, Nanni L, Rau D, Attene G, Papa R (2009) Linkage disequilibrium and population structure in wild and domesticated populations of Phaseolus vulgaris L. Evolutionary applications. Published Online July 2009. doi:10.1111/j.1752-4571.2009.00082.x
Santalla M, Rodiño AP, De Ron AM (2002) Allozyme evidence supporting southwester Europe as a secondary center of genetic diversity for common bean. Theor Appl Genet 104:934–944
SAS Institute Inc. (2007) JMP Software ver. 7
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Sicard D, Nanni L, Porfiri O, Bulfon D, Papa R (2005) Genetic diversity of Phaseolus vulgaris L and P. coccineus L. landraces in central Italy. Plant Breed 124:464–472
Singh SP (2001) Broadening the genetic base of common bean cultivars: a review. Crop Sci 41:1659–1675
Singh SP, Nodari R, Gepts P (1991) Genetic diversity in cultivated common bean. I. allozymes. Crop Sci 31:19–23
Sukhotu T, Kamijima O, Hosaka K (2006) Chloroplast DNA variation in the most primitive cultivated diploid potato species Solanum stenotomum Juz. et Buk. and its putative wild ancestral species using high-resolution markers. Genet Resour Crop Evol 53:53–63
Sun SM, Hall TC (1975) Solubility characteristics of globulins of Phaseolus seeds in regard to their isolation and characterisation. J Agric Food Chem 23:184–189
Vendramin GG, Lelli L, Rossi P, Morgante M (1996) A set of primers for the amplification of 20 chloroplast microsatellites in Pinaceae. Mol Ecol 5:595–598
Vigouroux Y, Mitchell S, Matsuoka Y, Hamblin M, Kresovich S, Smith JSC, Jaqueth J, Smith OS, Doebley J (2005) An analysis of genetic diversity across the maize genome using microsatellites genetics. Genetics 169:1617–1630
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Xu DH, Abe J, Gai JY, Shimamoto Y (2002) Diversity of chloroplast DNA SSRs in wild and cultivated soybeans: evidence for multiple origins of cultivated soybean. Theor Appl Genet 105:645–653
Zhang X, Blair MW, Wang S (2008) Genetic diversity of Chinese common bean Phaseolus vulgaris L.) landraces assessed with simple sequence repeats markers. Theor Appl Genet 117:629–640
Acknowledgments
S.A. Angioi and D. Rau made equal contributions to this study, and so should be considered joint first authors. We are grateful to A.H.D. Brown for his suggestions on the statistical analyses of this study and for critical reading of the manuscript. This study was supported by the Italian Government (MIUR) Grant no. # 2005071310, Project PRIN 2005 and constituted part of the PhD thesis of S.A. Angioi.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Bervillé.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Angioi, S.A., Rau, D., Attene, G. et al. Beans in Europe: origin and structure of the European landraces of Phaseolus vulgaris L.. Theor Appl Genet 121, 829–843 (2010). https://doi.org/10.1007/s00122-010-1353-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-010-1353-2