Abstract
Purpose
The relation between functional imaging and intrapatient genetic heterogeneity remains poorly understood. The aim of our study was to investigate spatial sampling and functional imaging by FDG-PET/MRI to describe intrapatient tumour heterogeneity.
Methods
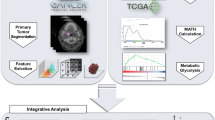
Six patients with oropharyngeal cancer were included in this pilot study. Two tumour samples per patient were taken and sequenced by next-generation sequencing covering 327 genes relevant in head and neck cancer. Corresponding regions were delineated on pretherapeutic FDG-PET/MRI images to extract apparent diffusion coefficients and standardized uptake values.
Results
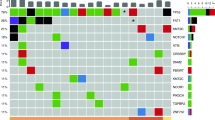
Samples were collected within the primary tumour (n = 3), within the primary tumour and the involved lymph node (n = 2) as well as within two independent primary tumours (n = 1). Genetic heterogeneity of the primary tumours was limited and most driver gene mutations were found ubiquitously. Slightly increasing heterogeneity was found between primary tumours and lymph node metastases. One private predicted driver mutation within a primary tumour and one in a lymph node were found. However, the two independent primary tumours did not show any shared mutations in spite of a clinically suspected field cancerosis. No conclusive correlation between genetic heterogeneity and heterogeneity of PET/MRI-derived parameters was observed.
Conclusion
Our limited data suggest that single sampling might be sufficient in some patients with oropharyngeal cancer. However, few driver mutations might be missed and, if feasible, spatial sampling should be considered. In two independent primary tumours, both lesions should be sequenced. Our data with a limited number of patients do not support the concept that multiparametric PET/MRI features are useful to guide biopsies for genetic tumour characterization.





Similar content being viewed by others
References
Bentzen SM, Gregoire V (2011) Molecular imaging-based dose painting: a novel paradigm for radiation therapy prescription. Semin Radiat Oncol 21(2):101–110. https://doi.org/10.1016/j.semradonc.2010.10.001
Grönlund E, Johansson S, Montelius A, Ahnesjö A (2017) Dose painting by numbers based on retrospectively determined recurrence probabilities. Radiother Oncol 122(2):236–241. https://doi.org/10.1016/j.radonc.2016.09.007
Cancer Genome Atlas Network (2015) Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517(7536):576–582. https://doi.org/10.1038/nature14129
Leemans CR, Snijders PJF, Brakenhoff RH (2018) The molecular landscape of head and neck cancer. Nat Rev Cancer 18(5):269–282. https://doi.org/10.1038/nrc.2018.11
Zwirner K, Hilke FJ, Demidov G, Socarras Fernandez J, Ossowski S, Gani C, Thorwarth D, Riess O, Zips D, Schroeder C, Welz S (2019) Radiogenomics in head and neck cancer: correlation of radiomic heterogeneity and somatic mutations in TP53, FAT1 and KMT2D. Strahlenther Onkol 195(9):771–779. https://doi.org/10.1007/s00066-019-01478-x
Greaves M (2015) Evolutionary determinants of cancer. Cancer Discov 5(8):806–820. https://doi.org/10.1158/2159-8290.CD-15-0439
McGranahan N, Swanton C (2017) Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168(4):613–628. https://doi.org/10.1016/j.cell.2017.01.018
Morris LG, Riaz N, Desrichard A, Şenbabaoğlu Y, Hakimi AA, Makarov V, Reis-Filho JS, Chan TA (2016) Pan-cancer analysis of intratumor heterogeneity as a prognostic determinant of survival. Oncotarget 7(9):10051–10063. https://doi.org/10.18632/oncotarget.7067
Gerlinger M, Horswell S, Larkin J, Rowan AJ, Salm MP, Varela I, Fisher R, McGranahan N, Matthews N, Santos CR, Martinez P, Phillimore B, Begum S, Rabinowitz A, Spencer-Dene B, Gulati S, Bates PA, Stamp G, Pickering L, Gore M, Nicol DL, Hazell S, Futreal PA, Stewart A, Swanton C (2014) Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat Genet 46(3):225–233. https://doi.org/10.1038/ng.2891
Gerlinger M, Rowan AJ, Horswell S, Math M, Larkin J, Endesfelder D, Gronroos E, Martinez P, Matthews N, Stewart A, Tarpey P, Varela I, Phillimore B, Begum S, McDonald NQ, Butler A, Jones D, Raine K, Latimer C, Santos CR, Nohadani M, Eklund AC, Spencer-Dene B, Clark G, Pickering L, Stamp G, Gore M, Szallasi Z, Downward J, Futreal PA, Swanton C (2012) Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 366(10):883–892. https://doi.org/10.1056/NEJMoa1113205
Zhang J, Fujimoto J, Zhang J, Wedge DC, Song X, Zhang J, Seth S, Chow CW, Cao Y, Gumbs C, Gold KA, Kalhor N, Little L, Mahadeshwar H, Moran C, Protopopov A, Sun H, Tang J, Wu X, Ye Y, William WN, Lee JJ, Heymach JV, Hong WK, Swisher S, Wistuba II, Futreal PA (2014) Intratumor heterogeneity in localized lung adenocarcinomas delineated by multiregion sequencing. Science 346(6206):256–259. https://doi.org/10.1126/science.1256930
O’Connor JP, Rose CJ, Waterton JC, Carano RA, Parker GJ, Jackson A (2015) Imaging intratumor heterogeneity: role in therapy response, resistance, and clinical outcome. Clin Cancer Res 21(2):249–257. https://doi.org/10.1158/1078-0432.CCR-14-0990
Aerts HJ, Velazquez ER, Leijenaar RT, Parmar C, Grossmann P, Carvalho S, Bussink J, Monshouwer R, Haibe-Kains B, Rietveld D, Hoebers F, Rietbergen MM, Leemans CR, Dekker A, Quackenbush J, Gillies RJ, Lambin P (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun 5:4006. https://doi.org/10.1038/ncomms5006
Jansen RW, van Amstel P, Martens RM, Kooi IE, Wesseling P, de Langen AJ, Menke-Van der Houven van Oordt CW, Jansen BHE, Moll AC, Dorsman JC, Castelijns JA, de Graaf P, de Jong MC (2018) Non-invasive tumor genoty** using radiogenomic biomarkers, a systematic review and oncology-wide pathway analysis. Oncotarget 9(28):20134–20155. https://doi.org/10.18632/oncotarget.24893
Lin G, Keshari KR, Park JM (2017) Cancer metabolism and tumor heterogeneity: imaging perspectives using MR imaging and spectroscopy. Contrast Media Mol Imaging 2017:6053879. https://doi.org/10.1155/2017/6053879
Leibfarth S, Winter RM, Lyng H, Zips D, Thorwarth D (2018) Potentials and challenges of diffusion-weighted magnetic resonance imaging in radiotherapy. Clin Transl Radiat Oncol 13:29–37. https://doi.org/10.1016/j.ctro.2018.09.002
Becker M, Zaidi H (2014) Imaging in head and neck squamous cell carcinoma: the potential role of PET/MRI. Br J Radiol 87(1036):20130677. https://doi.org/10.1259/bjr.20130677
Mehanna H, Wong WL, McConkey CC, Rahman JK, Robinson M, Hartley AG, Nutting C, Powell N, Al-Booz H, Robinson M, Junor E, Rizwanullah M, von Zeidler SV, Wieshmann H, Hulme C, Smith AF, Hall P, Dunn J (2016) PET-CT surveillance versus neck dissection in advanced head and neck cancer. N Engl J Med 374(15):1444–1454. https://doi.org/10.1056/NEJMoa1514493
Ledgerwood LG, Kumar D, Eterovic AK, Wick J, Chen K, Zhao H, Tazi L, Manna P, Kerley S, Joshi R, Wang L, Chiosea SI, Garnett JD, Tsue TT, Chien J, Mills GB, Grandis JR, Thomas SM (2016) The degree of intratumor mutational heterogeneity varies by primary tumor sub-site. Oncotarget 7(19):27185–27198. https://doi.org/10.18632/oncotarget.8448
Zhang XC, Xu C, Mitchell RM, Zhang B, Zhao D, Li Y, Huang X, Fan W, Wang H, Lerma LA, Upton MP, Hay A, Méndez E, Zhao LP (2013) Tumor evolution and intratumor heterogeneity of an oropharyngeal squamous cell carcinoma revealed by whole-genome sequencing. Neoplasia 15(12):1371–1378
Tabatabaeifar S, Thomassen M, Larsen MJ, Larsen SR, Kruse TA, Sørensen JA (2017) The subclonal structure and genomic evolution of oral squamous cell carcinoma revealed by ultra-deep sequencing. Oncotarget 8(10):16571–16580. https://doi.org/10.18632/oncotarget.15014
Hedberg ML, Goh G, Chiosea SI, Bauman JE, Freilino ML, Zeng Y, Wang L, Diergaarde BB, Gooding WE, Lui VW, Herbst RS, Lifton RP, Grandis JR (2016) Genetic landscape of metastatic and recurrent head and neck squamous cell carcinoma. J Clin Invest 126(4):1606. https://doi.org/10.1172/JCI86862
de Roest RH, Mes SW, Poell JB, Brink A, van de Wiel MA, Bloemena E, Thai E, Poli T, Leemans CR, Brakenhoff RH (2019) Molecular characterization of locally relapsed head and neck cancer after concomitant chemoradiotherapy. Clin Cancer Res 25(23):7256–7265. https://doi.org/10.1158/1078-0432.CCR-19-0628
Winter RM, Leibfarth S, Schmidt H, Zwirner K, Monnich D, Welz S, Schwenzer NF, la Fougere C, Nikolaou K, Gatidis S, Zips D, Thorwarth D (2018) Assessment of image quality of a radiotherapy-specific hardware solution for PET/MRI in head and neck cancer patients. Radiother Oncol. https://doi.org/10.1016/j.radonc.2018.04.018
Winter RM, Schmidt H, Leibfarth S, Zwirner K, Welz S, Schwenzer NF, la Fougere C, Nikolaou K, Gatidis S, Zips D, Thorwarth D (2017) Distortion correction of diffusion-weighted magnetic resonance imaging of the head and neck in radiotherapy position. Acta Oncol 56(11):1659–1663. https://doi.org/10.1080/0284186X.2017.1377347
Zwirner K, Thorwarth D, Winter RM, Welz S, Weiss J, Schwenzer NF, Schmidt H, la Fougere C, Nikolaou K, Zips D, Gatidis S (2018) Voxel-wise correlation of functional imaging parameters in HNSCC patients receiving PET/MRI in an irradiation setup. Strahlenther Onkol 194(8):719–726. https://doi.org/10.1007/s00066-018-1292-4
Eder T, Hess AK, Konschak R, Stromberger C, Johrens K, Fleischer V, Hummel M, Balermpas P, von der Grun J, Linge A, Lohaus F, Krause M, Baumann M, Stuschke M, Zips D, Grosu AL, Abdollahi A, Debus J, Belka C, Pigorsch S, Combs SE, Budach V, Tinhofer I (2019) Interference of tumour mutational burden with outcome of patients with head and neck cancer treated with definitive chemoradiation: a multicentre retrospective study of the German Cancer Consortium Radiation Oncology Group. Eur J Cancer 116:67–76. https://doi.org/10.1016/j.ejca.2019.04.015
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Kim S, Scheffler K, Halpern AL, Bekritsky MA, Noh E, Kallberg M, Chen X, Kim Y, Beyter D, Krusche P, Saunders CT (2018) Strelka2: fast and accurate calling of germline and somatic variants. Nat Methods 15(8):591–594. https://doi.org/10.1038/s41592-018-0051-x
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso‑2; iso‑3. Fly (Austin) 6(2):80–92. https://doi.org/10.4161/fly.19695
Tamborero D, Rubio-Perez C, Deu-Pons J, Schroeder MP, Vivancos A, Rovira A, Tusquets I, Albanell J, Rodon J, Tabernero J, de Torres C, Dienstmann R, Gonzalez-Perez A, Lopez-Bigas N (2018) Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med 10(1):25. https://doi.org/10.1186/s13073-018-0531-8
Nowell PC (1976) The clonal evolution of tumor cell populations. Science 194(4260):23–28
de Bruin EC, McGranahan N, Mitter R, Salm M, Wedge DC, Yates L, Jamal-Hanjani M, Shafi S, Murugaesu N, Rowan AJ, Gronroos E, Muhammad MA, Horswell S, Gerlinger M, Varela I, Jones D, Marshall J, Voet T, Van Loo P, Rassl DM, Rintoul RC, Janes SM, Lee SM, Forster M, Ahmad T, Lawrence D, Falzon M, Capitanio A, Harkins TT, Lee CC, Tom W, Teefe E, Chen SC, Begum S, Rabinowitz A, Phillimore B, Spencer-Dene B, Stamp G, Szallasi Z, Matthews N, Stewart A, Campbell P, Swanton C (2014) Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346(6206):251–256. https://doi.org/10.1126/science.1253462
Reiter JG, Makohon-Moore AP, Gerold JM, Heyde A, Attiyeh MA, Kohutek ZA, Tokheim CJ, Brown A, DeBlasio RM, Niyazov J, Zucker A, Karchin R, Kinzler KW, Iacobuzio-Donahue CA, Vogelstein B, Nowak MA (2018) Minimal functional driver gene heterogeneity among untreated metastases. Science 361(6406):1033–1037. https://doi.org/10.1126/science.aat7171
Moon SH, Kim J, Joung JG, Cha H, Park WY, Ahn JS, Ahn MJ, Park K, Choi JY, Lee KH, Kim BT, Lee SH (2018) Correlations between metabolic texture features, genetic heterogeneity, and mutation burden in patients with lung cancer. Eur J Nucl Med Mol Imaging. https://doi.org/10.1007/s00259-018-4138-5
Hu LS, Ning S, Eschbacher JM, Baxter LC, Gaw N, Ranjbar S, Plasencia J, Dueck AC, Peng S, Smith KA, Nakaji P, Karis JP, Quarles CC, Wu T, Loftus JC, Jenkins RB, Sicotte H, Kollmeyer TM, O’Neill BP, Elmquist W, Hoxworth JM, Frakes D, Sarkaria J, Swanson KR, Tran NL, Li J, Mitchell JR (2017) Radiogenomics to characterize regional genetic heterogeneity in glioblastoma. Neuro-oncology 19(1):128–137. https://doi.org/10.1093/neuonc/now135
O’Connor JPB (2017) Cancer heterogeneity and imaging. Semin Cell Dev Biol 64:48–57. https://doi.org/10.1016/j.semcdb.2016.10.001
Acknowledgements
We acknowledge the support of Prof. B. Sipos, Institute of Pathology and Neuropathology; Department of General and Molecular Pathology and Pathological Anatomy, Tübingen, Germany and Prof. I. Tinhofer, Department of Radiation Oncology and Radiotherapy, Charité Berlin, Germany. Furthermore, we would like to thank C. Goltermann for the language editing.
Funding
This project was funded by the Center for Personalised Medicine (ZPM) Tübingen/Germany (grant number 025/2015BO1). Kerstin Clasen was supported by the Fortüne/PATE Program of the Medical Faculty, Eberhard Karls University Tübingen (grant number: 2447-0-0). O. Riess and S. Ossowski receive funding by the German Research Foundation (DFG) as an NGS Competence Center (INST 37/1049-1). Parts of the research leading to these results have received funding from the European Research Council under the European Union’s Seventh Framework Programme (FP/2007-2013), ERC Grant Agreement no. 335367. None of our funding sources was involved in the study design, data analysis or data interpretation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
C. la Fougère has research collaborations with Siemens Healthineers. D. Zips and D. Thorwarth have research collaborations with Elekta, Philips and Siemens. K. Clasen, S. Leibfarth, F.J. Hilke, J. Admard, R.M. Winter, S. Welz, S. Gatidis, D. Nann, S. Ossowski, T. Breuer, K. Nikolaou, O. Riess and C. Schroeder declare that they have no competing interests.
Ethical standards
The study was approved by the local ethics committee (reference number 025/2015) and all patients declared their written informed consent. The study was performed in accordance with the ethical standards as laid down in the 1964 Declaration of Helsinki and its later amendments.
Rights and permissions
About this article
Cite this article
Clasen, K., Leibfarth, S., Hilke, F.J. et al. PET/MRI and genetic intrapatient heterogeneity in head and neck cancers. Strahlenther Onkol 196, 542–551 (2020). https://doi.org/10.1007/s00066-020-01606-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00066-020-01606-y