Abstract
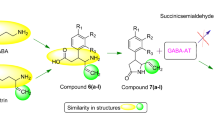
A new series, of γ-amino butyric acid analogs were designed and synthesized as novel potent GABA-AT inhibitors. A structure–activity relationship study was performed by correlating the effect of different substituents with GABA-AT inhibitory activity of the title compounds. The preliminary bioassays showed that acid hydrazones exhibited excellent inhibitory activities in micromolar (0.07–0.56 μM) range, while Schiff’s bases showed variable results. The most potent compound, 4-amino-N′-[(1Z)-1-(2-bromophenyl)ethylidene]butanehydrazide (AHG177) showed inhibitory potency (IC50) of 0.073 μM. Aminobutyrate transaminase is a pyridoxal-P enzyme which follows a bi–bi ** pong mechanism and in pyridoxamine form can readily transaminate only with succinic semialdehyde and 2-oxoglutarate. The results strongly suggest that only the pyridoxal form of the enzyme is capable of reacting with the ligands. Our findings open up the possibility to extend this protocol to different databases in order to find new potential inhibitor for promising targets based on a rational drug design process.








Similar content being viewed by others
References
Allison SA, Bacquet RJ, McCammon JA (1998) Simulation of the diffusion-controlled reaction between superoxide and superoxide dismutase II: detailed models. Biopolymers 27:251–269
Bansal SK, Sinha BN, Khosa RL, Olson AJ (2010) Novel GABA-AT inhibitors: QSAR and docking based virtual screening of phenyl-substituted β-phenyl ethylidene hydrazine analogs. Med Chem Res 19:S114
Bansal SK, Sinha BN, Khosa RL (2011a) QSAR and docking-based computational chemistry approach to novel GABA-AT inhibitors: kNN-MFA based 3DQSAR model for phenyl-substituted analogs of β-phenylethylidene hydrazine. Med Chem Res 20:549–553
Bansal SK, Sinha BN, Khosa RL (2011b) Docking based virtual screening of schiff’s bases of GABA- a prospective to novel GABA-AT inhibitors. Med Chem Res. doi:10.1007/s00044-011-9843-6
Bansal SK, Sinha BN, Khosa RL, Olson AJ (2011c) Novel GABA-AT inhibitors: QSAR and docking based virtual screening of phenyl-substituted β-phenyl ethylidene hydrazine analogs. Med Chem Res 20:1482–1489
Barbara M (2005) New anticonvulsant agents. Curr Top Med Chem 5:69–85
Bradford MM (1976) Rapid and sensitive method for quantitation of microgram quantities of protein utilizing the principle of dye binding. Anal Biochem 72:248–254
Gasteiger J, Marsili M (1980) Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron 36:3219–3228
Gerlach AC, Krajewski JL (2010) Antiepileptic drug discovery and development: what have we learned and where are we going? Pharmaceuticals 3:2884–2899
Goodford PJ (1985) A computational procedure for determining energetically favorable binding sites on biologically important macromolecules. J Med Chem 28:849–857
Goodsell DS, Olson AJ (1990) Automated docking of substrates to proteins by simulated annealing. Proteins Struct Funct Genet 8:195–202
Hardy LW, Abraham DJ, Sapo MK (2003) Structure based drug design. In: Abraham DJ (ed) Burger’s medicinal chemistry and drug discovery. Wiley, New Jersey
Jakoby WB (1962) Enzymes of γ-aminobutyrate metabolism. Methods Enzymol 5:771–774
Karlsson A, Fonnum F, Malthe-Sørenssen D, Storm-Mathisen J (1974) Effect of convulsive agent 3-mercaptopropionic acid on the levels of GABA other amino acids and glutamate decarboxylase in different regions of rat brain. Biochem Pharmacol 23:3053–3061
Koo BS, Park KS, Ha JH, Park JH, Lim JC, Lee DU (2003) Inhibitory effects of the fragrance inhalation of essential oil from Acorus gramineus on central nervous system. Biol Pharm Bull 26:978–982
Krogsgaard-Larsen P (1981) Gamma-aminobutyric acid agonists antagonists and uptake inhibitors Design and therapeutic aspects. J Med Chem 24:1377–1383
Kwon OS, Park J, Churchich JE (1992) Brain 4-aminobutyrate aminotransferase: isolation and sequence of a cDNA encoding the enzyme. J Biol Chem 267:7215–7216
Morris GM, Goodsell DS, Huey R, Olson AJ (1996) Distributed automated docking of flexible ligands to proteins: parallel applications of Autodock 2.4. J Comput Aided Mol Des 10:293–304
Nogardy T, Weaver DF (2005) Medicinal chemistry: a molecular and biochemical approach. Oxford University Press, New York
Novikov FN, Chilov GG (2009) Molecular docking: theoretical background, practical applications and perspectives. Mendeleev Commun 19:237–242
Osolodkin DI, Chupakhin VI, Palyulin VA, Zefirov NS (2009) Molecular modeling of ligand-receptor interaction in GABAC receptor. J Mol Graph Model 27:813–821
Ricci L, Frosini M, Gaggelli N, Valensin G, Machetti F, Sgaragli G, Valoti M (2006) Inhibition of rabbit brain 4-aminobutyrate transaminase by some taurine analogues: a kinetic analysis. Biochem Pharm 71:1510–1519
Scott EM, Jakoby WB (1959) Soluble γ-aminobutyric-glutamic transaminase from Pseudomonas fluorescens. J Biol Chem 234:932–936
Sharp K, Fine R, Honig B (1987) Computer simulations of the diffusion of a substrate to an active site of an enzyme. Science 236:1460–1463
Silverman RB, Clift MD (2008) Synthesis and evaluation of novel aromatic substrates and competitive inhibitors of GABA aminotransferase. Bioorg Med Chem Lett 18:3122–3125
Silverman RB, Durkee SC, Invergo BJ (1986) 4-Amino-2-(substituted methyl)-2-butenoic acids: substrates and potent inhibitors of gamma-aminobutyric acid aminotransferase. J Med Chem 29:764–770
Smith AJ, Simpson PB (2003) Methodological approaches for the study of GABAA receptor pharmacology and functional responses. Anal Bioanal Chem 377:843–851
Solis FJ, Wets RJ-B (1981) Minimization by random search techniques. Math Oper Res 6:19–30
Sowa B, Rauw G, Davood A, Fassihi A, Knaus EE, Baker GB (2005) Design and biological evaluation of phenyl-substituted analogs of β-phenyl ethylidene hydrazine. Bioorg Med Chem 13:4389–4395
Storici P, Capitani G, Baise DD, Moser M, John RA, Jansonius JN, Schirmer T (1999) Crystal structure of GABA aminotransferase a target for antiepileptic drug therapy. Biochemistry 38:8628–8634
Storici P, Baise DD, Bossa F, Bruno S, Mozzarelli A, Peneff C, Silverman RB, Schirmer T (2005) Structure of γ-amino butyric acid (GABA) aminotransferase a pyridoxal 5′-phosphate and [2Fe-2S] cluster-containing enzyme complexed with γ-ethynyl-GABA and with the antiepilepsy drug vigabatrin. J Biol Chem 279:363–373
Toney MD, Pascarella S, Baise DD (1995) Active site model for γ-amino butyrate aminotransferase explains substrate specificity and inhibitor reactivities. Protein Sci 4:2366–2374
Weiner SJ, Kollman PA, Case DA, Singh UC, Ghio C, Alagona G, Profeta S, Weiner P (1984) A new force field for molecular mechanical simulation of nucleic acid and proteins. J Am Chem Soc 106:765–784
Acknowledgments
Author wishes to thank Vice Chancellor, Birla Institute of Technology, Mesra, Ranchi and Chairman, Bharat Institute of Technology, Meerut to provide necessary infrastructural and other facilities.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1
3D crystal structure of GABA-AT
Rights and permissions
About this article
Cite this article
Bansal, S.K., Sinha, B.N. & Khosa, R.L. γ-Amino butyric acid analogs as novel potent GABA-AT inhibitors: molecular docking, synthesis, and biological evaluation. Med Chem Res 22, 134–146 (2013). https://doi.org/10.1007/s00044-012-0023-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-012-0023-0