Abstract
The escalating crisis of polyethylene terephthalate (PET) microplastic contamination in biological wastewater treatment systems is a pressing environmental concern. These microplastics inevitably accumulate in sewage sludge due to the absence of effective removal technologies. Addressing this urgent issue, this study introduces a novel approach using DuraPETase, a potent enzyme with enhanced PET hydrolytic activity at ambient temperatures. Remarkably, this enzyme was successfully secreted from Comamonas testosteroni CNB-1, a dominant species in the active sludge. The secreted DuraPETase showed significant hydrolytic activity toward p-NPB and PET nanoplastics. Furthermore, the CNB-1 derived whole-cell biocatalyst was able to depolymerize PET microplastics under ambient temperature, achieving a degradation efficiency of 9% within 7 days. The CNB-1-based whole biocatalysts were also capable of utilizing PET degradation intermediates, such as terephthalic acid (TPA) and ethylene glycol (EG), and bis(2-hydroxyethyl)-TPA (BHET), for growth. This indicates that it can completely mineralize PET, as opposed to merely breaking it down into smaller molecules. This research highlights the potential of activated sludge as a potent source for insitu microplastic removal.
Graphical Abstract

Similar content being viewed by others
Introduction
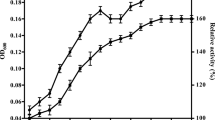
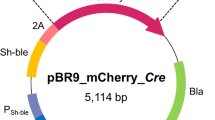
Plastics have revolutionized modern life, but the accumulation of synthetic polymers in landfills and the environment has created a global pollution crisis (Xu et al. 2023). Additionally, there has been increasing research interest in the potential adverse effects of microplastics (ranging from 1 μm to 5 mm) on ecosystems and human health (Wei et al. 2019). Microplastics are produced through the weathering and mechanical tearing (such as tires) of larger pieces of plastic, and are also directly manufactured and utilized in many personal care and cosmetic products (PCCPs) (Nizzetto et al. 2016; van der Laan et al. 2023). Furthermore, microplastics are transported from raw wastewater to wastewater treatment plants (WWTPs) (Browne et al. 2013; Sun et al. 2020), WWTPs can efficient remove over 90% of microplastics from raw wastewater (Carr et al. 2016). However, duo to this high removal efficiency, most microplastics end up being retained in the sewage sludge (Mahon et al. 2017; Zhang et al. C. testosteroni CNB-1(CGMCC 1.12282) was cultivated and maintained at 30 °C in LB broth or on LB plates with 1.5% (wt/vol) agar. For PET degradation, a mineral salt medium (MSM, pH 7.0) containing (NH4)2SO4 1 g/L, K2HPO4 6 g/L, KH2PO4 1 g/L, MgSO4·7H2O 0.1 g/L, NaCl 5 g/L was used by adding PET nanoplastics or microplastics. The pET-29a(+) vector served as the backbone for expressing enzymes (IsPETase, LCC, and DuraPETase) in E. coli BL21(DE3). For the overexpression of DuaPETase in CNB-1, the plasmid used was a derivative of pBBR1MCS-2 (pBBR1MCS2pfer), with its promoter replaced with a strong promoter from C. testosteroni CNB-1 (Huang et al. 2019). When necessary, antibiotics were added at the following final concentrations: 200 μg/ml kanamycin for C. testosteroni; 50 μg/ml kanamycin, and 100 μg/ml ampicillin for E. coli. DuraPETase (Genbank: GAP38373.1), which was mutated of 10 amino acid sites (S214H/I168R/W159H/S188Q/R280A/A180I/G165A/Q119Y/L117F/T140D) in the backbone of IsPETase. In this study, the ten mutated sites were regenerated in IsPETase and then commercially synthesized with codon optimization for expression in E. coli cells (GenScript, Nan**g, China). The genes were cloned into the pBBR1MCS2pfer vector, a pBBR1MCS-2 derivative plasmid whose promoter was replaced with a strong promoter from C. testosteroni, and contained a gfp gene (Huang et al. 1A), indicating that DuraPETase exhibited higher catalytic activity at 30 °C. To further confirm this, we measured the depolymerization efficacy of the three enzymes using PET films (Goodfellow) as a substrate. After treating the films with an equal amount of enzymes, we monitored the weight loss and the total products released from the PET films. The total released products (TPA, MHET, and BHET), determined by HPLC indicated that the highest degradation product release was observed for DuraPETase (Fig. 1B), which is about 1.5 folds than that of LCC, and over 10 times higher than that of IsPETase (Fig. 1B). These data also supported the weight loss of the PET films (without pretreatment) after enzymatic degradation. After 72 hours of enzymatic treatment, DuraPETase resulted in a weight loss of 2.2 mg (Fig. 1C), whereas IsPETase and LCC only showed weight losses of 0.4 mg and 1.4 mg, respectively (Fig. 1C). Overall, these results indicated that DuraPETase exhibited the highest catalytic activity at 30 °C, and is a suitable enzyme for subsequent heterologous expression in C. testosteroni CNB-1. Comparison of hydrolytic activity was conducted among IsPETase, DuraPETase, and LCC at a temperature of 30 °C. A Transparent zones were formed through the incubation of purified enzymes with Impranil® DLN as the substrate. B The release of degrading products from PET films after 72 h of incubation with IsPETase, LCC, and DuraPETase at 30 °C was analyzed. The total product release was quantified by summing the detected released compounds (TPA, MHET, and BHET). C The weight loss of PET films after incubation with purified enzymes for 72 h, with 100 mg PET films being utilized for degradation To efficiently degrade PET, the secretion of DuraPETase is crucial, as PET is a high molecular weight polymer that cannot enter cells. To achieve this, we selected a native signal peptide from CNB-1, named OmpC (gene locus: CtCNB1_1477), to assist the efficient secretion of DuraPETase in CNB-1. OmpC is a frequently used signal peptide for efficient secretion of heterologous proteins (Zhou et al. 2018). In addition to extracellular secretion, a high level of DuraPETase transcription is also essential for PET degradation. To accomplish this, a strong constitutive promoter from C. testosteroni CNB-1 to replace the promoter in the pBBR1MCS-2 plasmid, resulting in a new plasmid called pBBR1MCS2pfer was used (Huang et al. 2019). The signal peptide and the DuraPETase gene were inserted into the pBBR1MCS2pfer. Then this recombinant plasmid was transformed into C. testosteroni, generating the CNB-1B strain. Similarly, we also constructed a recombinant strain containing the DuraPETase gene with GFP, a strain without the signal peptide, and a strain with the original empty plasmid (Fig. 2A). Heterogeneous expression of DuraPETase was observed in C. testosteroni CNB-1. A DuraPETase was fused with different signal peptide and promoters to facilitate extracellular secretion in CNB-1. Different recombinant plasmids were constructed and introduced into CNB-1 to generate corresponding strains, which were designated as CNB-1A to CNB-1D, respectively. B The secretion efficiency of DuraPETase with or without OmpC signal peptide, was determined by measuring the intensity of extracellular GFP, which was fused with DuraPETase. C The hydrolytic activity of the recombinant strains was assessed on Impranil® DLN plates We chose CNB-1C (with SPOmpC) and CNB-1D (without SPOmpC) to test the secretion efficiency of the recombinant strains. Both of these strains are tagged with GFP, and the only difference between them is the signal peptide. As shown in Fig. 2B, OmpC signal peptides (SPOmpC) successfully direct the secretion of DuraPETase (Fig. 2B), resulting in a two-fold increase in GFP intensity in the supernatants than the recombinant strain CNB-1D (Fig. 2B). Interestingly, even when no signal peptide was fused, the strain CNB-1D was still able to export extracellular DuraPETase-GFP, although the GFP intensity was much lower than in CNB-1C (Fig. 2B). This could be due to the phospholipid hydrolase activity of DuraPETase. Phospholipid is an important component of the cell membrane, and hydrolysis of phospholipids can enhance membrane permeability and release of DuraPETase into the extracellular milieu (Su et al. 2013). Similar phenomena have been observed in many PET-degrading cutinases expressed in E. coli in previous studies (Su et al. 2013). Also, CNB-1C generates a higher total GFP than CNB-1D, indicating that the signal peptide may contribute to the expression of DuraPETase (Fig. 2B). This is reasonable considering that high concentrations of DuraPETase can be toxic to microbial cells, and a secretion system may reduce cytotoxicity and facilitate higher-level expression of DuraPETase. Furthermore, when using Impranil® DLN as a substrate, all the recombinant strains showed similar transparent zones except CNB-1A, which only contains an empty vector (Fig. 2C). This is consistent with the extracellular GFP intensity data, as DuraPETase can be successfully secreted when no signal peptide was fused. Taken together, these data suggested that DuraPETase was functionally expressed in C. testosteroni CNB-1, and the signal peptide SPOmpC contributed to the secretion. To investigate whether the DuraPETase secreted by C. testosteroni CNB-1 is active against PET, we conducted enzymatic analysis using model substrates such as p-NPB. Although the structure of p-NPB is different from PET, it has been reported that the conversion rates of small-molecule para-nitrophenyl esters are comparable to PET oligomers (Wei et al. 2022). Therefore, we first used p-NPB as a substrate to assess the hydrolytic activity of DuraPETase when expressed in CNB-1. As expected, CNB-1B and CNB-1C exhibited significant kinetic hydrolytic activity against p-NPB compared to the blank control CNB-1A, and CNB-1D (Fig. 3A), another recombinant strain lacking a secretion signal peptide. Among them, CNB-1C showed a little higher hydrolytic activity than CNB-1B, but the difference was not significant (Fig. 3A). Moreover, the SEM images further confirmed that CNB-1B can induce surface erosion to PET films after a two-week incubation period (Fig. 3C), whereas the PET films incubated with the control strain (CNB-1A) maintained a smooth surface (Fig. 3B). Consequently, we selected the recombinant strain CNB-1B for further experiments. Enzymatic activity analysis was conducted on recombinant strains derived from CNB-1. A The catalyst activity of the secreted DuraPETase was determined using p-NPB as a substrate. The strains were incubated with MSM medium at 30 °C for 3 days, and 1 mL of the supernatant was collected for incubation with p-NPB at 30 °C for 30 min, and the absorbance at 410 nm was measured. SEM images revealed surface erosion of PET films after 14 days of degradation by CNB-1A (B) and CNB-1B (C), at 30 °C We then investigated the hydrolytic activity of CNB-1B on PET nanoplastics, which were prepared from PET films. These nanoplastics possess a high surface-to-volume ratio, making them significantly more susceptible to hydrolysis in comparison to the films (Barth et al. 2015; Walter et al. 1995). After incubating CNB-1B with PET nanoplastics as the sole carbon source for 10 days, we use thermogravimetric analysis to study the nanoplastics degradation. The thermogram in Fig. 4A illustrates the degradation of nanoplastics by CNB-1 without the heterologous expression of DuraPETase, which shows two stages of weight loss (Fig. 4A). Previous publications have suggested that the first stage of weight loss (approximately 5%) between 25 and 100 °C is caused by the presence of water trapped in nanoplastics and bacterial cells. The second stage of weight loss, occurring between 100 and 500 °C, is attributed to the removal of organic matter from bacterial cells and most of the PET nanoplastics (Fig. 4A). It is important to note that PET decomposes above 250 °C, as indicated by previous reports (Sorolla-Rosario et al. 2022). In comparison, when the nanoplastics are treated with DuraPETase, we observed three distinct stages of weight loss (Fig. 4B). The first two stages are consistent with the findings mentioned above, but the third stage, occurring from 400 to 700 °C (Fig. 4B), is primarily due to the repolymerized of BHET during the heating process in the thermogravimetric study. This observation aligns with a similar thermogram of BHET reported in another study (Scé et al. 2019). These results indicate that PET nanoplastics are degraded by DuraPETase, resulting in the formation of the intermediate compound BHET. This finding suggests that the secreted DuraPETase enzyme is capable of hydrolyzing PET. Thermogravimetric analysis (TGA) was conducted to examine the degradation of PET nanoplastics. The differential scanning calorimetry (DSC) and TGA curves of glycolysis products by CNB-1A (A) and CNB-1B (B) are displayed, with the red line representing the DSC curve and the black line representing the TGA curve. When PET nanoplastics were degraded by the secreted DuraPETase from CNB-1B, the product BHET was released from PET, and the TGA curve showed a distinct BHET polymerization stage, which can be considered as evidence of PET nanoplastics degradation by CNB-1 derived whole-cell biocatalyst Wastewater treatment plants play a significant role in the entry of microplastics into the environment (Murphy et al. 2016; Sun et al. 2019). It is crucial to develop technology that specifically targets and treats microplastics. Microplastics are primarily found in sludge, which makes separating them difficult and inefficient (Talvitie et al. 2017). Therefore, we have proposed an in situ degradation framework in which engineered C. testosteroni CNB-1 is introduced to secrete DuraPETase to hydrolyze microplastics (PET films with an initial crystallinity of approximately 4%). To this end, we first examined the degradation of PET microplastics by the CNB-1 derived PET-degrading whole-cell biocatalyst. After incubating the biocatalysts in MSM medium with 5% (v/v) LB for 7 days, we found that the whole-cell biocatalysts degraded 9.2 mg of the microplastics, which corresponds to a 9% degradation of the total microplastics added (Fig. 5B). This result suggests that when DuraPETase is secreted, the CNB-1 derived whole-cell biocatalyst is capable of degrading PET microplastics. The performance of the CNB-1 derived whole-cell biocatalyst in PET microplastic degradation. A The growth curve of CNB-1B using TPA, EG, and BHET as the sole carbon sources. The cell density was measured for TPA and EG, as these compounds are easily utilized by CNB-1B. For BHET, the colony-forming units (CFU) was counted. B The degradation of PET microplastics by CNB-1B, 100 mg of PET microplastics was incubated with CNB-1B in MSM + 5% LB (v/v) as a carbon source. For blank control, no carbon source was added. The incubation took place at 30 °C for 7 days More importantly, we also found that the CNB-1 derived PET-degrading whole-cell biocatalyst can utilize main intermediates of enzymatic PET degradation, including TPA, EG, and BHET, as sole carbon sources for growth (Fig. 5A). While Comamonas is commonly recognized as capable of degrading TPA (Dierkes Robert et al. 2022; Hosaka et al. 2013), the utilization of EG and BHET has not been characterized before. This means that CNB-1-based whole-cell biocatalyst can completely mineralize the PET microplastics, rather than merely degrading it into smaller molecules. In another publication, by functionally immobilizing PETase on the self-assembled E. coli curli nanofibers (Zhu et al. 2022), this system could depolymerize highly crystalline post-consumer PET waste materials under ambient conditions with a degradation efficiency of 9.1% in 7 days (Zhu et al. 2022), which is similar to the degradation efficiency observed in this study. Moreover, the whole-cell biocatalyst may have better performance in the real-world sewage sludge system due to the presence of unlimited nutrients (Zhu et al. 2022). This would result in the consumption of organic matter to express depolymerase to hydrolyze microplastics. Our results suggested the potential of CNB-1 derived PET-degrading whole-cell biocatalyst for the removal of PET microplastics in advanced wastewater treatment applications. With the sludge-indigenous strain C. testosteroni CNB-1 as the host, we have proposed an insitu framework for the treatment of sludge microplastics. We have designed a novel whole-cell biocatalyst that facilitates the degradation of PET microplastics by incorporating DuraPETase as the key enzyme, thereby allowing for the efficient depolymerization of PET into TPA and EG. These released intermediates can be further utilized by CNB-1 as the sole carbon source. Thus, the PET-degrading whole-cell biocatalyst derived from CNB-1 opens up a promising approach that could potentially lead to the complete mineralization of PET microplastics present in the sewage sludge. It is worth noting that the microplastic degradation carried out in this study was conducted under small-scale lab conditions. To facilitate the practical application of the PET-degrading whole-cell biocatalyst derived from CNB-1, further research is necessary to evaluate its performance in the degradation of PET microplastics in larger-scale systems such as wastewater treatment bioreactors. In this study, we developed a novel whole-cell biocatalyst for the potential removal of microplastics in situ, using sludge-derived C. testosteroni CNB-1. We compared the catalytic activity of the reported PET hydrolyses (IsPETase, LCC, and DuraPETase) at 30 °C and chose DuaPETase for heterologous expression in C. testosteroni CNB-1. After optimizing the promoter and signal peptide, the DuraPETase was successfully secreted into the extracellular milieu from C. testosteroni CNB-1. The secretion of DuraPETase led to an efficient hydrolytic activity towards PET nanoplastics and microplastics.Material and methods
Bacterial strains, plasmids, and culture conditions
Expression of DuraPETase in C. testosteroni
Functional expression of DuraPETase enabled PET degradation by C. testosteroni CNB-1
Hydrolytic activity of secreted DuraPETase on PET
PET microplastic degradation by CNB-1 derived whole-cell biocatalyst
Conclusion
Availability of data and materials
Data may be made available on request.
References
Aksu D, Diallo MM, Şahar U, Uyaniker TA, Ozdemir G (2021) High expression of ring-hydroxylating dioxygenase genes ensure efficient degradation of p-toluate, phthalate, and terephthalate by Comamonas testosteroni strain 3a2. Arch Microbiol 203(7):4101–4112. https://doi.org/10.1007/s00203-021-02395-3
Ateşlier ZBB, Metin K (2006) Production and partial characterization of a novel thermostable esterase from a thermophilic Bacillus sp. Enzyme Microb Technol 38(5):628–635. https://doi.org/10.1016/j.enzmictec.2005.07.015
Barth M, Oeser T, Wei R, Then J, Schmidt J, Zimmermann W (2015) Effect of hydrolysis products on the enzymatic degradation of polyethylene terephthalate nanoparticles by a polyester hydrolase from Thermobifida fusca. Biochem Eng J 93:222–228. https://doi.org/10.1016/j.bej.2014.10.012
Brott S, Pfaff L, Schuricht J, Schwarz J-N, Böttcher D, Badenhorst CPS, Wei R, Bornscheuer UT (2022) Engineering and evaluation of thermostable IsPETase variants for PET degradation. Eng Life Sci 22(3–4):192–203. https://doi.org/10.1002/elsc.202100105
Browne Mark A, Niven Stewart J, Galloway Tamara S, Rowland Steve J, Thompson Richard C (2013) Microplastic moves pollutants and additives to worms, reducing functions linked to health and biodiversity. Curr Biol 23(23):2388–2392. https://doi.org/10.1016/j.cub.2013.10.012
Carr SA, Liu J, Tesoro AG (2016) Transport and fate of microplastic particles in wastewater treatment plants. Water Res 91:174–182. https://doi.org/10.1016/j.watres.2016.01.002
Chen K, Hu Y, Dong X, Sun Y (2021) Molecular insights into the enhanced performance of EKylated PETase toward PET degradation. ACS Catal 11(12):7358–7370. https://doi.org/10.1021/acscatal.1c01062
Chen J, Wu J, Sherrell PC, Chen J, Wang H, Zhang W-x, Yang J (2022) How to build a microplastics-free environment: strategies for microplastics degradation and plastics recycling. Adv Sci 9(6):2103764. https://doi.org/10.1002/advs.202103764
Cui Y, Chen Y, Liu X, Dong S, Ye T, Qiao Y, Mitra R, Han J, Li C, Han X, Liu W, Chen Q, Wei W, Wang X, Du W, Tang S, **ang H, Liu H, Liang Y, Houk KN, Wu B (2021) Computational redesign of a PETase for plastic biodegradation under ambient condition by the GRAPE strategy. ACS Catal 11(3):1340–1350. https://doi.org/10.1021/acscatal.0c05126
Dierkes Robert F, Wypych A, Pérez-García P, Danso D, Chow J, Streit Wolfgang R (2022) An ultra-sensitive comamonas thiooxidans biosensor for the rapid detection of enzymatic polyethylene terephthalate (PET) degradation. Appl Environ Microbiol 89(1):e01603-e1622. https://doi.org/10.1128/aem.01603-22
Dong Q, Yuan S, Wu L, Su L, Zhao Q, Wu J, Huang W, Zhou J (2020) Structure-guided engineering of a Thermobifida fusca cutinase for enhanced hydrolysis on natural polyester substrate. Bioresources Bioprocess 7(1):37. https://doi.org/10.1186/s40643-020-00324-8
Hosaka M, Kamimura N, Toribami S, Mori K, Kasai D, Fukuda M, Masai E (2013) Novel tripartite aromatic acid transporter essential for terephthalate uptake in Comamonas sp. strain E6. Appl Environ Microbiol 79(19):6148–6155. https://doi.org/10.1128/AEM.01600-13
Huang Z, Wang Y-H, Zhu H-Z, Andrianova Ekaterina P, Jiang C-Y, Li D, Ma L, Feng J, Liu Z-P, **ang H, Zhulin Igor B, Liu S-J (2019) Cross talk between chemosensory pathways that modulate chemotaxis and biofilm formation. Mbio. https://doi.org/10.1128/mbio.02876-18
Knott BC, Erickson E, Allen MD, Gado JE, Graham R, Kearns FL, Pardo I, Topuzlu E, Anderson JJ, Austin HP, Dominick G, Johnson CW, Rorrer NA, Szostkiewicz CJ, Copié V, Payne CM, Woodcock HL, Donohoe BS, Beckham GT, McGeehan JE (2020) Characterization and engineering of a two-enzyme system for plastics depolymerization. Proc Natl Acad Sci 117(41):25476–25485. https://doi.org/10.1073/pnas.2006753117
Li X, Chen L, Mei Q, Dong B, Dai X, Ding G, Zeng EY (2018) Microplastics in sewage sludge from the wastewater treatment plants in China. Water Res 142:75–85. https://doi.org/10.1016/j.watres.2018.05.034
Li X-y, Liu H-t, Wang L-x, Guo H-n, Zhang J, Gao D (2022) Effects of typical sludge treatment on microplastics in China—characteristics, abundance and micro-morphological evidence. Sci Total Environ 826:154206. https://doi.org/10.1016/j.scitotenv.2022.154206
Liu L, Jiang C-Y, Liu X-Y, Wu J-F, Han J-G, Liu S-J (2007) Plant–microbe association for rhizoremediation of chloronitroaromatic pollutants with Comamonas sp. strain CNB-1. Environ Microbiol 9(2):465–473. https://doi.org/10.1111/j.1462-2920.2006.01163.x
Ma Y-F, Zhang Y, Zhang J-Y, Chen D-W, Zhu Y, Zheng H, Wang S-Y, Jiang C-Y, Zhao G-P, Liu S-J (2009) The complete genome of Comamonas testosteroni reveals its genetic adaptations to changing environments. Appl Environ Microbiol 75(21):6812–6819. https://doi.org/10.1128/AEM.00933-09
Mahon AM, O’Connell B, Healy MG, O’Connor I, Officer R, Nash R, Morrison L (2017) Microplastics in sewage sludge: effects of treatment. Environ Sci Technol 51(2):810–818. https://doi.org/10.1021/acs.est.6b04048
Murphy F, Ewins C, Carbonnier F, Quinn B (2016) Wastewater treatment works (WwTW) as a source of microplastics in the aquatic environment. Environ Sci Technol 50(11):5800–5808. https://doi.org/10.1021/acs.est.5b05416
Nizzetto L, Futter M, Langaas S (2016) Are agricultural soils dumps for microplastics of urban origin? Environ Sci Technol 50(20):10777–10779. https://doi.org/10.1021/acs.est.6b04140
Qiao Y, Hu R, Chen D, Wang L, Wang Z, Yu H, Fu Y, Li C, Dong Z, Weng Y-X, Du W (2022) Fluorescence-activated droplet sorting of PET degrading microorganisms. J Hazard Mater 424:127417. https://doi.org/10.1016/j.jhazmat.2021.127417
Rochman CM, Hoh E, Hentschel BT, Kaye S (2013) Long-term field measurement of sorption of organic contaminants to five types of plastic pellets: implications for plastic marine debris. Environ Sci Technol 47(3):1646–1654. https://doi.org/10.1021/es303700s
Scé F, Cano I, Martin C, Beobide G, Castillo Ó, de Pedro I (2019) Comparing conventional and microwave-assisted heating in PET degradation mediated by imidazolium-based halometallate complexes. New J Chem 43(8):3476–3485. https://doi.org/10.1039/C8NJ06090H
Sorolla-Rosario D, Llorca-Porcel J, Pérez-Martínez M, Lozano-Castelló D, Bueno-López A (2022) Study of microplastics with semicrystalline and amorphous structure identification by TGA and DSC. J Environ Chem Eng 10(1):106886. https://doi.org/10.1016/j.jece.2021.106886
Su L, Woodard Ronald W, Chen J, Wu J (2013) Extracellular location of thermobifida fusca cutinase expressed in Escherichia coli BL21(DE3) without mediation of a signal peptide. Appl Environ Microbiol 79(14):4192–4198. https://doi.org/10.1128/AEM.00239-13
Sun J, Dai X, Wang Q, van Loosdrecht MCM, Ni BJ (2019) Microplastics in wastewater treatment plants: detection, occurrence and removal. Water Res 152:21–37. https://doi.org/10.1016/j.watres.2018.12.050
Sun Q, Ren S-Y, Ni H-G (2020) Incidence of microplastics in personal care products: an appreciable part of plastic pollution. Sci Total Environ 742:140218. https://doi.org/10.1016/j.scitotenv.2020.140218
Talvitie J, Mikola A, Koistinen A, Setälä O (2017) Solutions to microplastic pollution—removal of microplastics from wastewater effluent with advanced wastewater treatment technologies. Water Res 123:401–407. https://doi.org/10.1016/j.watres.2017.07.005
Thakur B, Singh J, Singh J, Angmo D, Vig AP (2023) Biodegradation of different types of microplastics: molecular mechanism and degradation efficiency. Sci Total Environ 877:162912. https://doi.org/10.1016/j.scitotenv.2023.162912
Tournier V, Topham CM, Gilles A, David B, Folgoas C, Moya-Leclair E, Kamionka E, Desrousseaux ML, Texier H, Gavalda S, Cot M, Guémard E, Dalibey M, Nomme J, Cioci G, Barbe S, Chateau M, André I, Duquesne S, Marty A (2020) An engineered PET depolymerase to break down and recycle plastic bottles. Nature 580(7802):216–219. https://doi.org/10.1038/s41586-020-2149-4
van der Laan LJW, Bosker T, Peijnenburg WJGM (2023) Deciphering potential implications of dietary microplastics for human health. Nat Rev Gastroenterol Hepatol 20(6):340–341. https://doi.org/10.1038/s41575-022-00734-3
Walter T, Augusta J, Müller R-J, Widdecke H, Klein J (1995) Enzymatic degradation of a model polyester by lipase from Rhizopus delemar. Enzyme Microb Technol 17(3):218–224. https://doi.org/10.1016/0141-0229(94)00007-E
Wang Y-H, Chen H-H, Huang Z, Li X-J, Zhou N, Liu C, Jiang C-Y, Li D-F, Liu S-J (2021) PapA, a peptidoglycan-associated protein, interacts with OmpC and maintains cell envelope integrity. Environ Microbiol 23(2):600–612. https://doi.org/10.1111/1462-2920.15038
Wei R, Oeser T, Then J, Kühn N, Barth M, Schmidt J, Zimmermann W (2014) Functional characterization and structural modeling of synthetic polyester-degrading hydrolases from Thermomonospora curvata. AMB Express 4(1):44. https://doi.org/10.1186/s13568-014-0044-9
Wei W, Huang Q-S, Sun J, Wang J-Y, Wu S-L, Ni B-J (2019) Polyvinyl chloride microplastics affect methane production from the anaerobic digestion of waste activated sludge through leaching toxic bisphenol-A. Environ Sci Technol 53(5):2509–2517. https://doi.org/10.1021/acs.est.8b07069
Wei R, von Haugwitz G, Pfaff L, Mican J, Badenhorst CPS, Liu W, Weber G, Austin HP, Bednar D, Damborsky J, Bornscheuer UT (2022) Mechanism-based design of efficient PET hydrolases. ACS Catal 12(6):3382–3396. https://doi.org/10.1021/acscatal.1c05856
Wu L, Ning D, Zhang B, Li Y, Zhang P, Shan X, Zhang Q, Brown MR, Li Z, Van Nostrand JD, Ling F, **ao N, Zhang Y, Vierheilig J, Wells GF, Yang Y, Deng Y, Tu Q, Wang A, Acevedo D, Agullo-Barcelo M, Alvarez PJJ, Alvarez-Cohen L, Andersen GL, de Araujo JC, Boehnke KF, Bond P, Bott CB, Bovio P, Brewster RK, Bux F, Cabezas A, Cabrol L, Chen S, Criddle CS, Deng Y, Etchebehere C, Ford A, Frigon D, Sanabria J, Griffin JS, Gu AZ, Habagil M, Hale L, Hardeman SD, Harmon M, Horn H, Hu Z, Jauffur S, Johnson DR, Keller J, Keucken A, Kumari S, Leal CD, Lebrun LA, Lee J, Lee M, Lee ZMP, Li Y, Li Z, Li M, Li X, Ling F, Liu Y, Luthy RG, Mendonça-Hagler LC, de Menezes FGR, Meyers AJ, Mohebbi A, Nielsen PH, Ning D, Oehmen A, Palmer A, Parameswaran P, Park J, Patsch D, Reginatto V, de los Reyes FL, Rittmann BE, Noyola A, Rossetti S, Shan X, Sidhu J, Sloan WT, Smith K, de Sousa OV, Stahl DA, Stephens K, Tian R, Tiedje JM, Tooker NB, Tu Q, Van Nostrand JD, de los Cobos Vasconcelos D, Vierheilig J, Wagner M, Wakelin S, Wang A, Wang B, Weaver JE, Wells GF, West S, Wilmes P, Woo SG, Wu L, Wu JH, Wu L, ** C, **ao N, Xu M, Yan T, Yang Y, Yang M, Young M, Yue H, Zhang B, Zhang P, Zhang Q, Zhang Y, Zhang T, Zhang Q, Zhang W, Zhang Y, Zhou H, Zhou J, Wen X, Curtis TP, He Q, He Z, Brown MR, Zhang T, He Z, Keller J, Nielsen PH, Alvarez PJJ, Criddle CS, Wagner M, Tiedje JM, He Q, Curtis TP, Stahl DA, Alvarez-Cohen L, Rittmann BE, Wen X, Zhou J, Global Water Microbiome C (2019) Global diversity and biogeography of bacterial communities in wastewater treatment plants. Nat Microbiol 4(7):1183–1195. https://doi.org/10.1038/s41564-019-0426-5
Xu A, Zhou J, Blank LM, Jiang M (2023) Future focuses of enzymatic plastic degradation. Trends Microbiol 31(7):668–671. https://doi.org/10.1016/j.tim.2023.04.002
Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Kimura Y, Oda K (2016) A bacterium that degrades and assimilates poly(ethylene terephthalate). Science 351(6278):1196–1199. https://doi.org/10.1126/science.aad6359
Zhang B, Ning D, Van Nostrand JD, Sun C, Yang Y, Zhou J, Wen X (2020a) Biogeography and assembly of microbial communities in wastewater treatment plants in China. Environ Sci Technol 54(9):5884–5892. https://doi.org/10.1021/acs.est.9b07950
Zhang L, **e Y, Liu J, Zhong S, Qian Y, Gao P (2020b) An overlooked entry pathway of microplastics into agricultural soils from application of sludge-based fertilizers. Environ Sci Technol 54(7):4248–4255. https://doi.org/10.1021/acs.est.9b07905
Zhou Y, Lu Z, Wang X, Selvaraj JN, Zhang G (2018) Genetic engineering modification and fermentation optimization for extracellular production of recombinant proteins using Escherichia coli. Appl Microbiol Biotechnol 102(4):1545–1556. https://doi.org/10.1007/s00253-017-8700-z
Zhu X, Qi J, Cheng L, Zhen G, Lu X, Zhang X (2022) Depolymerization and conversion of waste-activated sludge to value-added bioproducts by fungi. Fuel 320:123890. https://doi.org/10.1016/j.fuel.2022.123890
Acknowledgements
The authors are indebted to Dr. Shuangjiang Liu at the Institute of Microbiology, Chinese Academy of Sciences for providing the Comamonas testosteroni CNB-1 strain, and the pBBR1MCS2pfer plasmid. The authors are supported by the National Key R & D Program of China (2019YFA0905500), the National Natural Science Foundation of China (21978129, 31961133017), the European Union’s Horizon 2020 research and innovation program under grant agreement no. 870294 for the project MIX-UP, the Natural Science Foundation of Jiangsu Province of China for Excellent Young Scholars (BK20211591), and the Jiangsu Synergetic Innovation Center for Advanced Bio-Manufacture (XTB2203).
Author information
Authors and Affiliations
Contributions
Anming Xu and Weiliang Dong conceived and designed the experiments. Zhanqing Cao, Wei **a, Shilei Wu and Jiale Ma performed the experiments. Zhanqing Cao, Shilei Wu, and **aoli Zhou analyzed the data. Anming Xu and **ujuan Qian wrote the draft manuscript. Weiliang Dong and Min Jiang supervised the study. All authors revised and proofread the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
There are no conflicts to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cao, Z., **a, W., Wu, S. et al. Bioengineering Comamonas testosteroni CNB-1: a robust whole-cell biocatalyst for efficient PET microplastic degradation. Bioresour. Bioprocess. 10, 94 (2023). https://doi.org/10.1186/s40643-023-00715-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40643-023-00715-7