Abstract
Background
Hypoxia contributes to cancer progression through various molecular mechanisms and hepatocellular carcinoma (HCC) is one of the most hypoxic malignancies. Hypoxia-inducible gene domain protein-1a (HIGD1A) is typically induced via epigenetic regulation and promotes tumor cell survival during hypoxia. However, the role of HIGD1A in HCC remains unknown.
Methods
HIGD1A expression was determined in 24 pairs of human HCC samples and para-tumorous tissues. Loss-of-function experiments were conducted both in vivo and in vitro to explore the role of HIGD1A in HCC proliferation and metastasis.
Results
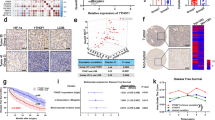
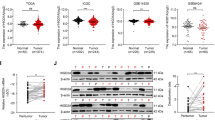
Increased HIGD1A expression was found in HCC tissues and cell lines, which was induced by hypoxia or low-glucose condition. Moreover, HIGD1A knockdown in HCC cells arrested the cell cycle at the G2/M phase and promoted hypoxia-induced cell apoptosis, resulting in great inhibition of cell proliferation, migration, and invasion, as well as tumor xenograft formation. Interestingly, these anti-tumor effects were not observed in normal hepatocyte cell line L02. Further, HIGD1A knockdown suppressed the expression of ornithine decarboxylase 1 (ODC1), a rate-limiting enzyme of polyamine metabolism under c-Myc regulation. HIGD1A was found to bind with the c-Myc promoter region, and its knockdown decreased the levels of polyamine metabolites. Consistently, the inhibitory effect on HCC phenotype by HIGD1A silencing could be reversed by overexpression of c-Myc or supplementation of polyamines.
Conclusions
Our results demonstrated that HIGD1A activated c-Myc–ODC1 nexus to regulate polyamine synthesis and to promote HCC survival and malignant phenotype, implying that HIGD1A might represent a novel therapeutic target for HCC.
Similar content being viewed by others
Background
Hepatocellular carcinoma (HCC), a major type of liver cancer, ranks fourth in global incidence and third in mortality among malignant tumors [1]. In the past decade, despite notable advancements in novel drugs and therapies within the field of liver cancer treatment, along with progress in concepts such as early screening, integrated prevention, and collaboration treatment, the overall prognosis for liver cancer remains dismal [2]. The 5-year overall survival rate is only 12.1%, with merely 30% of patients having the opportunity for surgical resection [3]. Previous studies have indicated that tumor recurrence, metastasis, and drug resistance were the major causes of poor prognosis in HCC patients [4]. Therefore, further exploring the molecular mechanisms that participate in HCC development and progression would be useful for identifying novel drug targets.
HCC is among the most hypoxic malignancies, particularly within areas of tumor necrosis [5]. Moreover, treatments for HCC, such as transarterial chemoembolization that restricts blood supply to inhibit tumor growth, or tyrosine kinase inhibitors that reduce angiogenesis by inhibiting various kinase targets, can further aggravate intratumoral hypoxia [6,7,8]. Many studies revealed that within HCC tissues, the expression levels of hypoxia-inducible factors (such as HIF-1α, HIF-2α) or their target genes (GLUT1, LDHA, CA9, SLC7A1) are significantly elevated in comparison to non-tumorous liver tissue. Moreover, this elevated expression was linked to tumor recurrence, metastasis, and reduced patient survival rates [9, 10]. Functional research demonstrated that the signaling cascade of HIF molecules was involved in HCC cell proliferation, angiogenesis, invasion, and metastasis [11, 34]. Certainly, further experimental validation is needed to confirm this hypothesis.
Polyamines were essential for cell proliferation and survival, and both the growth of tumor cells and tumor progression require elevated levels of polyamines [35]. Inhibiting polyamine metabolic processes or reducing polyamine metabolic products can exert anti-tumor effects [36]. ODC1 is a rate-limiting enzyme in the downstream polyamine biosynthetic pathway [37]. In liver cancer, the expression and activity of ODC1 were elevated compared to normal tissues [29]. Difluoromethylornithine is a highly effective and specific inhibitor of ODC1 [38] and its treatment resulted in intracellular polyamine depletion, cell cycle arrest [39], thereby inhibiting tumor cell growth. Considering the significant decrease in the expression of c-Myc and ODC1 following HIGD1A silencing in HCC cells, it suggested that the intracellular polyamine metabolic pathway was inhibited, leading to a reduction in polyamine products. This inhibition consequently suppressed HCC cell proliferation and other phenotypes. Based on the above observation, we speculated that the polyamine metabolic products might serve as the effector molecules through which HIGD1A regulated the phenotypes of HCC cells. Therefore, our results further confirmed the proposed strategy that targets c-Myc and ODC1 in HCC treatment.
Conclusions
Current anti-tumor therapies for HCC remain unsatisfactory, predominantly due to their limited efficacy in blocking the high rates of recurrence and metastasis. Our study has revealed that targeted modulation of HIGD1A yields a notable inhibition to the proliferative, migratory, and invasive capacities of HCC cells. This effect is associated with polyamine levels regulated by the c-Myc–ODC1 interplay. Importantly, our study reveals the discrepancies in HIGD1A expressions and functionalities between HCC cells and their normal counterparts. This discernment helps us to identify the underlying mechanisms driving HCC tumorigenesis. Moreover, it implies a promising target for HCC therapeutic intervention, suggesting that the development of HIGD1A inhibitors could be potentially used for the treatment of liver cancer.
Availability of data and materials
All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials. Additional data related to this paper may be requested from the corresponding authors.
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- HIGD1A:
-
Hypoxia-inducible gene domain protein-1a
- ODC1:
-
Ornithine decarboxylase 1
- HIFs:
-
Hypoxia-inducible factors
- CHIP:
-
Chromatin immunoprecipitation
- HPLC:
-
High-Performance Liquid Chromatography
- DNMT1:
-
DNA methyl transferase 1
- CHIP:
-
Chromatin immunoprecipitation
- APH:
-
Aphidicolin
- OCR:
-
Oxygen Consumption Rate
- ECAR:
-
Extracellular Acidification Rate
- EMT:
-
Epithelial-to-mesenchymal transition
- GSEA:
-
Gene set enrichment analysis
- DEGs:
-
Differentially expressed genes
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209–49.
Villanueva A. Hepatocellular Carcinoma. N Engl J Med. 2019;380:1450–62.
Song P, Cai Y, Tang H, Li C, Huang J. The clinical management of hepatocellular carcinoma worldwide: a concise review and comparison of current guidelines from 2001 to 2017. Biosci Trends. 2017;11:389–98.
Bao MH, Wong CC. Hypoxia, Metabolic Reprogramming, and Drug Resistance in Liver Cancer. Cells. 2021;10:1715.
McKeown SR. Defining normoxia, physoxia and hypoxia in tumours-implications for treatment response. Br J Radiol. 2014;87:20130676.
Kudo M. A New Treatment Option for Intermediate-Stage Hepatocellular Carcinoma with High Tumor Burden: Initial Lenvatinib Therapy with Subsequent Selective TACE. Liver Cancer. 2019;8:299–311.
Cucarull B, Tutusaus A, Rider P, Hernáez-Alsina T, Cuño C, García de Frutos P, Morales A et al. Hepatocellular Carcinoma: Molecular Pathogenesis and Therapeutic Advances. Cancers (Basel). 2022;14:621.
Awosika J, Sohal D. A narrative review of systemic treatment options for hepatocellular carcinoma: state of the art review. J Gastrointest Oncol. 2022;13:426–37.
Cao S, Yang S, Wu C, Wang Y, Jiang J, Lu Z. Protein expression of hypoxia-inducible factor-1 alpha and hepatocellular carcinoma: a systematic review with meta-analysis. Clin Res Hepatol Gastroenterol. 2014;38:598–603.
Wada H, Nagano H, Yamamoto H, Yang Y, Kondo M, Ota H, Monden M, et al. Expression pattern of angiogenic factors and prognosis after hepatic resection in hepatocellular carcinoma: importance of angiopoietin-2 and hypoxia-induced factor-1 alpha. Liver Int. 2006;26:414–23.
Sun HX, Xu Y, Yang XR, Wang WM, Bai H, Shi RY, Fan J, et al. Hypoxia inducible factor 2 alpha inhibits hepatocellular carcinoma growth through the transcription factor dimerization partner 3/ E2F transcription factor 1-dependent apoptotic pathway. Hepatology. 2013;57:1088–97.
Guo Y, **ao Z, Yang L, Gao Y, Zhu Q, Hu L, Xu Q, et al. Hypoxia-inducible factors in hepatocellular carcinoma (Review). Oncol Rep. 2020;43:3–15.
Ameri K, Rajah AM, Nguyen V, Sanders TA, Jahangiri A, Delay M, Maltepe E, et al. Nuclear localization of the mitochondrial factor HIGD1A during metabolic stress. PLoS One. 2013;8:e62758.
Timón-Gómez A, Bartley-Dier E.L, Fontanesi F, Barrientos A. HIGD-Driven Regulation of Cytochrome c Oxidase Biogenesis and Function. Cells. 2020;9:2620.
Nagao T, Shintani Y, Hayashi T, Kioka H, Kato H, Nishida Y, Takashima S, et al. Higd1a improves respiratory function in the models of mitochondrial disorder. Faseb J. 2020;34:1859–71.
Ameri K, Jahangiri A, Rajah AM, Tormos KV, Nagarajan R, Pekmezci M, Maltepe E, et al. HIGD1A Regulates Oxygen Consumption, ROS Production, and AMPK Activity during Glucose Deprivation to Modulate Cell Survival and Tumor Growth. Cell Rep. 2015;10:891–9.
Chen B, Xu F, Gao Y, Hu G, Zhu K, Lu H, Zhao G, et al. DNA damage-induced translocation of mitochondrial factor HIGD1A into the nucleus regulates homologous recombination and radio/chemo-sensitivity. Oncogene. 2022;41:1918–30.
**e Z, Shen S, Huang K, Wang W, Liu Z, Zhang H, Zhang X et al. Mitochondrial HIGD1A inhibits hepatitis B virus transcription and replication through the cellular PNKD‐NF‐κB‐NR2F1 nexus. J Med Virol. 2023;95:e28749.
An HJ, Ryu M, Jeong HJ, Kang M, Jeon HM, Lee JO, Lee H, et al. Higd-1a regulates the proliferation of pancreatic cancer cells through a pERK/p27(KIP1)/pRB pathway. Cancer Lett. 2019;461:78–89.
Xu Z, Sun J, Mao Y, Chen Y, Zhang T, Qin Y, Hua D. HIG1 domain family member 1A disrupts proliferation, migration, and invasion of colon adenocarcinoma cells. Bioengineered. 2021;12:10501–11.
Kicmal T.M, Tate PM, Dial CN, Esin JJ, Mounce BC. Polyamine Depletion Abrogates Enterovirus Cellular Attachment. J Virol. 2019;93:e01054–19.
Yu D, Shi X, Meng G, Chen J, Yan C, Jiang Y, Ding Y, et al. Kidney-type glutaminase (GLS1) is a biomarker for pathologic diagnosis and prognosis of hepatocellular carcinoma. Oncotarget. 2015;6:7619–31.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Mesirov JP, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–50.
Song JW, Lam SM, Fan X, Cao WJ, Wang SY, Tian H, Shui G, et al. Omics-Driven Systems Interrogation of Metabolic Dysregulation in COVID-19 Pathogenesis. Cell Metab. 2020;32:188-202.e185.
Yu C, Liu R, **e C, Zhang Q, Yin Y, Bi K, Li Q. Quantification of free polyamines and their metabolites in biofluids and liver tissue by UHPLC-MS/MS: application to identify the potential biomarkers of hepatocellular carcinoma. Anal Bioanal Chem. 2015;407:6891–7.
Li T, **an WJ, Gao Y, Jiang S, Yu QH, Zheng QC, Zhang Y. Higd1a Protects Cells from Lipotoxicity under High-Fat Exposure. Oxid Med Cell Longev. 2019;2019:6051262.
Ameri K, Maltepe E. HIGD1A-mediated dormancy and tumor survival. Mol Cell Oncol. 2015;2:e1030537.
Hogarty MD, Norris MD, Davis K, Liu X, Evageliou NF, Hayes CS, Haber M, et al. ODC1 is a critical determinant of MYCN oncogenesis and a therapeutic target in neuroblastoma. Cancer Res. 2008;68:9735–45.
Ye Z, Zeng Z, Shen Y, Yang Q, Chen D, Chen Z, Shen S. ODC1 promotes proliferation and mobility via the AKT/GSK3β/β-catenin pathway and modulation of acidotic microenvironment in human hepatocellular carcinoma. Onco Targets Ther. 2019;12:4081–92.
Xu H, Liu R, He B, Bi CW, Bi K, Li Q. Polyamine Metabolites Profiling for Characterization of Lung and Liver Cancer Using an LC-Tandem MS Method with Multiple Statistical Data Mining Strategies: Discovering Potential Cancer Biomarkers in Human Plasma and Urine. Molecules. 2016;21:1040.
Cheng Z, Wang G, Zhu W, Luo C, Guo Z. LEF1-AS1 accelerates tumorigenesis in glioma by sponging miR-489-3p to enhance HIGD1A. Cell Death Dis. 2020;11:690.
Timón-Gómez A, Garlich J, Stuart RA, Ugalde C, Barrientos A. Distinct Roles of Mitochondrial HIGD1A and HIGD2A in Respiratory Complex and Supercomplex Biogenesis. Cell Rep. 2020;31:107607.
Huang K, Liu Z, **e Z, Li X, Zhang H, Chen Y, Zhang X, et al. HIGD2A silencing impairs hepatocellular carcinoma growth via inhibiting mitochondrial function and the MAPK/ERK pathway. J Transl Med. 2023;21:253.
Chen YC, Taylor EB, Dephoure N, Heo JM, Tonhato A, Papandreou I, Rutter J, et al. Identification of a protein mediating respiratory supercomplex stability. Cell Metab. 2012;15:348–60.
Igarashi K, Kashiwagi K. Modulation of cellular function by polyamines. Int J Biochem Cell Biol. 2010;42:39–51.
Wallace HM, Fraser AV. Inhibitors of polyamine metabolism: review article. Amino Acids. 2004;26:353–65.
Dang CV. c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol Cell Biol. 1999;19:1–11.
Alexiou GA, Lianos GD, Ragos V, Galani V, Kyritsis AP. Difluoromethylornithine in cancer: new advances. Future Oncol. 2017;13:809–19.
Chang L, Li Z, Guo H, Zhang W, Lan W, Wang J, Zhao P et al. Function of Polyamines in Regulating Cell Cycle Progression of Cultured Silkworm Cells. Insects. 2021;12:624.
Acknowledgments
We would like to thank Prof. Chuanjiang Li (Division of Hepatobiliopancreatic Surgery, Department of General Surgery, Nanfang Hospital, Southern Medical University) for providing the clinical samples.
Funding
This study was supported by grants from the National Key Research and Development Program of China (No.2022YFC2303603), the National Natural Science Foundation of China (82172257), and the Health Care Major Project of Guangzhou (202206080001).
Author information
Authors and Affiliations
Contributions
HYL, XYZ, and JLH conceived and designed the study. HXZ, XRL, ZYL, ZML, KYH, YRW, YC, LYL, LYW, and ZLX performed the experiments, data collection, and statistical analysis. HXZ, XRL, and ZYL analyzed the data and wrote the manuscript. XYZ and HYL contributed to data interpretation and critical revision of the manuscript. All authors reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was approved by the Southern Medical University of Nanfang Hospital (Approval No. 2019-029) and informed consent was obtained from all patients. All animal protocols were approved by the Animal Ethics Committee of the Nanfang Hospital of Southern Medical University (Permit No. IACUC-LAC-20221009-006).
Consent for publication
All authors read and approved the final manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article has been updated to correct the missing affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Zhang, H., Li, X., Liu, Z. et al. Elevated expression of HIGD1A drives hepatocellular carcinoma progression by regulating polyamine metabolism through c-Myc–ODC1 nexus. Cancer Metab 12, 7 (2024). https://doi.org/10.1186/s40170-024-00334-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40170-024-00334-6