Abstract
Soybean mosaic virus (SMV) is one of the most destructive viral diseases in soybean and causes severe reduction of soybean yield and destroys the seed quality. However, the production of SMV resistant plants by transgenic is the most effective and economical means. Based on our previous yeast two-hybrid assay, the GmVma12 was selected as a strong candidate gene for further function characterization. Here we transformed soybean plants with a construct containing inverted repeat of-GmVma12 sequence to analyze the role of GmVma12 during SMV invasion. Totals of 33 T0 and 160 T1 plants were confirmed as positive transgenic plants through herbicide application, PCR detection and LibertyLink® strip screening. Based on the segregation ratio and Southern Blot data, T1 lines No. 3 and No. 7 were selected to generate T2 plants. After SMV-SC15 inoculation, 41 T1 and 38 T2 plants were identified as highly resistant, and their quantification disease levels were much lower than non-transformed plants. The transcript level of GmVma12 in T2 plants decreased to 70% of non-transformed plants. The expression level of SMV-CP transcript in T2 transgenic plants was lower than that in non-transformed plants and SMV CP protein in T2 plants could not be detected by Enzyme-linked Immunosorbent assay, which indicated that SMV production would be inhibited in transgenic plants. Moreover, coat mottles of T2 seeds were obliterated significantly. In conclusion, inverted repeat of the hairpin structure of GmVma12 interfered with the transcription of GmVma12, which can induce resistance to SMV in soybean. This research lays the foundation for the mechanism of SMV pathogenesis, and provides new ideas for SMV prevention and control.
Similar content being viewed by others
Introduction
Soybean (Glycine max (L.) Merr.) originated from China has been cultivated for more than five thousand years, supplying plant fat and protein in human daily diet. Soybean mosaic virus (SMV) infection is prevalent from north spring soybean production area to the Yangtze River valley area in China (Li et al. 2010). Once SMV infects soybean, leaves are damaged severely, which leads to the reduction of photosynthetic production, nutrient absorption and transport capacity. Ultimately SMV affects the grain traits, resulting in 10–30% yields reduction, or even crop failure (Liao et al. 2002). One potential viable approach to prevent SMV infections in soybeans is by identifying resistance genes and breeding SMV resistant soybean cultivars.
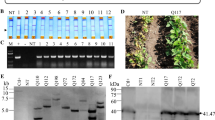
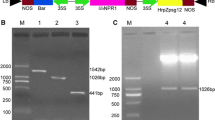
Agrobacterium is a naturally occurring microorganism that can serve as transgenic vector in nature (Hadi et al. 1996). Agrobacterium-mediated transformation, due to many advantages such as low cost, low gene copy number and genetic stability, has become a popular method in crop breeding (Hansen et al. 1999). It was reported that the first transgenic soybean was produced in 1988 using Agrobacterium-mediated transformation and since then, this technology has been widely used in soybean transgenic research (Hinchee et al. 1988). Currently, using agrobacterium transformation to induce RNAi-mediated resistance to SMV is a popular strategy in China. The RNAi strategy was used to trigger robust resistance against three viruses in soybean plants by expressing several short inverted repeat of a portion of the virus sequences (Zhang et al. 2011). The construct containing inverted repeat of SMV-HC-Pro was transformed into five soybean genotypes. The transgenic soybeans demonstrated strong resistant to SMV in transgenic plants (Gao et al. 2015). In addition, the transgenic lines with silenced P3 cistron by RNAi showed significantly increased resistance to multiple potyvirus strains and isolates (Yang et al. 2003; Rong et al. 1996). The segregation ratio of T0 line No. 3 and line No. 7 were both 3:1 (Table 2). This segregation ratio matched with the single copy insertion results of Southern Blot (Fig. 6), which indicated that the progenies of these two lines could generate stable and inheritable transgenic lines.
RNA interference is a natural defense response of plants to viral infections (Tenllado et al. 2003). At present, RNAi has mainly been applied on the coat protein of virus, movement protein, the replicase, and the dominant or recessive disease resistance genes of host plants. Transgenic plants have great differences in virus resistance, and are mainly classified into high resistance, low resistance, susceptible and recovery. The coat protein of Tobacco etch virus (TEV) was transformed into tobacco (Goodwin et al. 1996). After inoculation with TEV, the virus was found in the plants in a short time. However, with the passage of time, the accumulation of viruses in new leaves gradually decreased. This phenomenon is called “recovery”, which was speculated related to transgene copy number(Goodwin et al. 1996). The qRT-PCR and DAS-ELISA assay showed that the silencing efficiency of the endogenous genes was not 100%, which indicated that their expression was inhibited to some extent (Fig. 6). In our experiment, the similar penotype was also found and named delay resistance(DR). The silencing efficiency of GmVma12i and accumulation of SMV in DR plants were also detected (no-show data), which was lower than HR plants. We assumed that the expression of RNAi construction and the efficiency of siRNAs were also important for the resistance phenotype. Moreover, the strength of delay resistance could also be parallel with the SMV accumulation. The severe symptom of SMV in early growth stage might attenuate DR (Furutani et al. 2007).
The vacuolar-type ATPase (VATP) is prevalent in the endometrium of eukaryotic and animal cells. This enzyme functions as a proton pump and plays an important role in the membrane system, including membrane trafficking and intracellular pH regulation (Beyenbach et al. 2006; Forgac et al. 2007). VATP contains multiple subunits that belong to the heterologous polyproteinases. In intracellular vacuolar transport, which is mediated by organelles such as the endoplasmic reticulum, Golgi apparatus, and lysosomes, VATP transports newly generated secretory proteins and receptor proteins to the plasma membrane (Bonifacino et al. 2004). Meanwhile, VATP also plays a key role in other intracellular transport pathways, such as coordinating the interaction between intracellular organ receptors and extracellular donor organs, organ fusion, budding, and secretion (Baars et al. 2007; Marshansky et al. 2008; 2014). In the Mannose-6-phosphate (M6P) pathway, VATP is involved in the synthesis and transport of hydrolases, and regulates the transport from the Golgi apparatus or the endoplasmic reticulum to lysosomes (Maxfield et al. 2004; Coutinho et al. 2012). Until now, there are no reports about the function of Vma12 related to disease resistance. It was obvious that Tianlong No.1 gained the resistance to SMV by knocking down Gmvma12, which implicates GmVma12 plays an important role in the invasion of SMV as a host factor.
In the future study, the stable homozygous line with stable resistance to SMV could be screened and used to study the role of GmVma12 during SMV infection, replication and transport. Moreover, it will be interesting to investigate the variation of ultrastructure in different stages after SMV inoculation, analyze the changes of expression of pathogenesis-related proteins and search for related pathways to clarify how GmVma12 functions in the process of SMV invasion.
Availability of data and materials
All data generated or analyzed during this study are included in the figures and tables. Any material used in this study is available for research purposes upon request.
References
Baars TL, Petri S, Peters C, Mayer A (2007) Role of the V-ATPase in regulation of the vacuolar fission fusion equilibrium. Mol Biol Cell 18(10):3873–3882. https://doi.org/10.1091/mbc.e07-03-0205
Beyenbach KW, Wieczorek H (2006) The V-type H+ ATPase: molecular structure and function, physiological roles and regulation. J Exp Biol 209(4):577–589. https://doi.org/10.1242/jeb.02014
Bonas U, Lahaye T (2002) Plant disease resistance triggered by pathogen-derived molecules: refined models of specific recognition. Curr Opin Microbiol 5(1):44–50. https://doi.org/10.1016/s1369-5274(02)00284-9
Bonifacino JS, Glick BS (2004) The mechanisms of vesicle budding and fusion. Cell 116(2):153–166. https://doi.org/10.1016/s0092-8674(03)01079-1
Coutinho MF, Prata MJ, Alves S (2012) Mannose-6-phosphate pathway: a review on its role in lysosomal function and dysfunction. Molecular genetics and metabolism. Mol Genet Metab 105(4):542–550. https://doi.org/10.1016/j.ymgme.2011.12.012->
Forgac M (2007) Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nat Rev Mol Cell Bio 8(11):917. https://doi.org/10.1038/nrm2272
Fraser RSS (1990) The genetics of resistance to plant viruses. Annu Rev Phytopathol 28(1):179–200. https://doi.org/10.1146/annurev.phyto.28.1.179
Fraser RSS, Van Loon LC (1985) Genes for resistance to plant viruses. Crit Rev Plant Sci 1986 3(3):257–294. https://doi.org/10.1080/07352688609382212
Furutani N, Yamagishi N, Hidaka S, Shizukawa Y, Kanematsu S, Kosaka Y (2007) Soybean mosaic virus resistance in transgenic soybean caused by post-transcriptional gene silencing. Breeding Sci 57(2):123–128. https://doi.org/10.1270/jsbbs.57.123
Gao L, Ding X, Liao LKW, Zhong Y, Ren R, Liu Z, Adhimoolam K, Zhi H (2015) Characterization of Soybean mosaic virus resistance derived from inverted repeat-SMV-HC-Pro genes in multiple soybean cultivars. Theor Appl Genet 128(8):1489–1505. https://doi.org/10.1007/s00122-015-2522-0
Goldbach R, Bucher E, Prins M (2003) Resistance mechanisms to plant viruses: an overview. Virus Res 92(2):207–212. https://doi.org/10.1016/s0168-1702(02)00353-2
Goodwin J, Chapman K, Swaney S, Parks TD, Wernsman EA, Dougherty WG (1996) Genetic and biochemical dissection of transgenic RNA-mediated virus resistance. Plant Cell 8(1):95–105. https://doi.org/10.2307/3870071
Hadi MZ, McMullen MD, Finer JJ (1996) Transformation of 12 different plasmids into soybean via particle bombardment. Plant Cell Rep 15(7):500–505. https://doi.org/10.1007/s002990050062
Hansen G, Wright MS (1999) Recent advances in the transformation of plants. Trends Plant Sci 4(6):226–231. https://doi.org/10.1016/s1360-1385(99)01412-0
Hinchee MAW, Connor-Ward DV, Newell CA, McDonnell Raymond E, Sato Shirley J, Gasser Charles S, Fischhoff David A, Re Diane B, Fraley Robert T, Horsch Robert B (1988) Production of transgenic soybean plants using Agrobacterium-mediated DNA transfer. Biotechnology 6(8):915–922. https://doi.org/10.1038/nbt0888-915
Höfgen R, Willmitzer L (1988) Storage of competent cells for Agrobacterium transformation. Nucleic Acids Res 16(20):9877. https://doi.org/10.1093/nar/16.20.9877
Johansen IE, Lund OS, Hjulsager CK, Laursen J (2001) Recessive resistance in pisum sativum and potyvirus pathotype resolved in a gene-for-cistron correspondence between host and virus. J Virol 75(14):6609–6614. https://doi.org/10.1128/jvi.75.14.6609-6614.2001
Karimi M, Inzé D, Depicker A (2002) Gateway™ vectors for agrobacterium -mediated plant transformation. Trends Plant Sci 7:193–195. https://doi.org/10.1016/s1360-1385(02)02251-3
Li K, Yang QH, Zhi HJ, Gai JY (2010) Identification and distribution of soybean mosaic virus strains in southern China. Plant Dis 94:351–357. https://doi.org/10.1094/PDIS-94-3-0351
Liao L, Chen P, Buss GR, Yang Q, Tolin SA (2002) Inheritance and allelism of resistance to soybean mosaic virus in Zao18 soybean from China. J Hered 93:447–452. https://doi.org/10.1093/jhered/93.6.447
Lim HS, Ko TS, Hobbs HA, Lambert KN, Yu JM, Mccoppin NK, Korban SS, Hartman GL, Domier LL (2007) Soybean mosaic virus helper component-protease alters leaf morphology and reduces seed production in transgenic soybean plants. Phytopathology 97:366–372. https://doi.org/10.1094/phyto-97-3-0366
Luan HX, Liao WL, Niu HP, Cui XY, Chen X, Zhi HJ (2019) Comprehensive analysis of soybean mosaic virus P3 protein interactors and hypersensitive response-like lesion-inducing protein function. Int J Mol Sci 20:3388. https://doi.org/10.3390/ijms20143388
Marshansky V, Futai M (2008) The V-type H+-ATPase in vesicular trafficking: targeting, regulation and function. Curr Opin Cell Biol 20:415–426. https://doi.org/10.1016/j.ceb.2008.03.015
Marshansky V, Rubinstein JL, Grüber G (2014) Eukaryotic V-ATPase: novel structural findings and functional insights. Biochim Biophys Acta 1837:857–879. https://doi.org/10.1016/j.bbabio.2014.01.018
Maxfield FR, Mcgraw TE (2004) Endocytic recycling. Nat Rev Mol Cell Biol 5:121. https://doi.org/10.1038/nrm1315
Olhoft P, Somers D (2001) L-cysteine increases Agrobacterium-mediated T-DNA delivery into soybean cotyledonary-node cells. Plant Cell Rep 20:706–711. https://doi.org/10.1007/s002990100379
Olhoft P, Lin K, Galbraith J, Nielsen N, Somers D (2001) The role of thiol compounds in increasing Agrobacterium -mediated transformation of soybean cotyledonary-node cells. Plant Cell Rep 20:731–737. https://doi.org/10.1007/s002990100388
Olhoft PM, Flagel LE, Donovan CM, Somers DA (2003) Efficient soybean transformation using hygromycin B selection in the cotyledonary-node method. Planta 216:723–735. https://doi.org/10.1007/s00425-002-0922-2
Robaglia C, Caranta C (2006) Translation initiation factors: a weak link in plant RNA virus infection. Trends Plant Sci 11:40–45. https://doi.org/10.1016/j.tplants.2005.11.004
Rong D, Purcell V, Collins GB, Ghabrial SA (1996) Production of transgenic soybean lines expressing the bean pod mottle virus coat protein precursor gene. Plant Cell Rep 15:746–750. https://doi.org/10.1007/s002990050112
Senda M, Masuta C, Ohnishi S, Goto K, Kasai A, Sano T, Hong JS, MacFarlane S (2004) Patterning of virus-infected Glycine max seed coat is associated with suppression of endogenous silencing of chalcone synthase genes. Plant Cell 16:807–818. https://doi.org/10.1105/tpc.019885
Soosaar JLM, Burchsmith TM, Dineshkumar SP (2005) Mechanisms of plant resistance to viruses. Nat Rev Microbiol 3:789–798
Tenllado F, Martínez-García B, Vargas M, Díaz-Ruíz JR (2003) Crude extracts of bacterially expressed dsRNA can be used to protect plants against virus infections. BMC Biotechnol 3:3. https://doi.org/10.1186/1472-6750-3-3
Truniger V, Aranda MA (2009) Recessive resistance to plant viruses. Adv Virus Res 75:119. https://doi.org/10.1016/s0065-3527(09)07504-6
Wang A, Krishnaswamy S (2012) Eukaryotic translation initiation factor 4E-mediated recessive resistance to plant viruses and its utility in crop improvement. Mol Plant Pathol 13:795–803. https://doi.org/10.1111/j.1364-3703.2012.00791.x
Yang X, Niu L, Zhang W, Yang J, **ng G, He H, Guo D, Du Q, Qian X, Yao Li Q, Dong Y (2018) RNAi-mediated SMV P3 cistron silencing confers significantly enhanced resistance to multiple Potyvirus strains and isolates in transgenic soybean. Plant Cell Rep 37:103–114. https://doi.org/10.1007/s00299-017-2186-0
Zhang X, Sato S, Ye X, Dorrance AE, Morris TJ, Clemente TE, Qu F (2011) Robust RNAi-based resistance to mixed infection of three viruses in soybean plants expressing separate short hairpins from a single transgene. Phytopathology 101:1264–1269. https://doi.org/10.1094/phyto-02-11-0056
Acknowledgements
We thank Keshun Yu and **angnan Guan for critical review and English edition.
Funding
This work was supported by the Fund of Transgenic Breeding for Soybean Resistance to Soybean mosaic virus (2016ZX08004-004), the Fundamental Research Funds for the Central Universities (KYT201801), Program for Changjiang Scholars and Innovative Research Team in University (PCSIRT_17R55), the National Natural Science Foundation of China (31571690), the National Science Foundation of China (31760392), the National Soybean Industrial Technology System of China (CARS-004), Jiangsu Collaborative Innovation Center for Modern Crop Production (JCIC-MCP), and the National Key R&D Program of China (2017YFD0101501).
Author information
Authors and Affiliations
Contributions
Funding acquisition, YS and HZ; investigation, HL, WL and HN; methodology, YS and TH; supervision, HL and WL; writing—original draft, HL and WL; writing—review & editing, HL.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 1: Fig. S1.
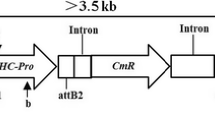
Schematic diagram of T-DNA region of recombinant plasmid pB7GWIWG2(II)-GmVma12i,Fig. S2. The morphology of Fig. S3. The transcript levels of two isforms of GmVma12 by qRT-PCR in T1 RNAi silenced transgenic plants qRT-PCR. X-axes indicate T1 transgenic plants and non-transformed (Mock) plants. Data are expressed as the means of three biological replicates with error bars indicating the SD (n = 3). Asterisks denote significant difference from mock, as determined by the t- test, p < 0.001. Each result is representative of three biological repeats. wild type and GmVma12 transgenic plants at (a) true leaf period stage and (b) pod setting stage. Table S1. Primer sequence. Table S2. Components in medium used for soybean transformation. Table S3. DAS-ELISA analyses of T2 plants at 30 dpi of SMV infection
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Luan, H., Liao, W., Song, Y. et al. Transgenic plant generated by RNAi-mediated knocking down of soybean Vma12 and soybean mosaic virus resistance evaluation. AMB Expr 10, 62 (2020). https://doi.org/10.1186/s13568-020-00997-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13568-020-00997-6