Abstract
Background
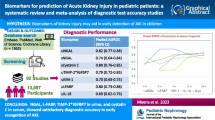
Several biomarkers have been proposed to predict the occurrence of acute kidney injury (AKI); however, their efficacy varies between different trials. The aim of this study was to compare the predictive performance of different candidate biomarkers for AKI.
Methods
In this systematic review, we searched PubMed, Medline, Embase, and the Cochrane Library for papers published up to August 15, 2022. We selected all studies of adults (> 18 years) that reported the predictive performance of damage biomarkers (neutrophil gelatinase-associated lipocalin (NGAL), kidney injury molecule-1 (KIM-1), liver-type fatty acid-binding protein (L-FABP)), inflammatory biomarker (interleukin-18 (IL-18)), and stress biomarker (tissue inhibitor of metalloproteinases-2 × insulin-like growth factor-binding protein-7 (TIMP-2 × IGFBP-7)) for the occurrence of AKI. We performed pairwise meta-analyses to calculate odds ratios (ORs) and 95% confidence intervals (CIs) individually. Hierarchical summary receiver operating characteristic curves (HSROCs) were used to summarize the pooled test performance, and the Grading of Recommendations, Assessment, Development and Evaluations criteria were used to appraise the quality of evidence.
Results
We identified 242 published relevant studies from 1,803 screened abstracts, of which 110 studies with 38,725 patients were included in this meta-analysis. Urinary NGAL/creatinine (diagnostic odds ratio [DOR] 16.2, 95% CI 10.1–25.9), urinary NGAL (DOR 13.8, 95% CI 10.2–18.8), and serum NGAL (DOR 12.6, 95% CI 9.3–17.3) had the best diagnostic accuracy for the risk of AKI. In subgroup analyses, urinary NGAL, urinary NGAL/creatinine, and serum NGAL had better diagnostic accuracy for AKI than urinary IL-18 in non-critically ill patients. However, all of the biomarkers had similar diagnostic accuracy in critically ill patients. In the setting of medical and non-sepsis patients, urinary NGAL had better predictive performance than urinary IL-18, urinary L-FABP, and urinary TIMP-2 × IGFBP-7: 0.3. In the surgical patients, urinary NGAL/creatinine and urinary KIM-1 had the best diagnostic accuracy. The HSROC values of urinary NGAL/creatinine, urinary NGAL, and serum NGAL were 91.4%, 85.2%, and 84.7%, respectively.
Conclusions
Biomarkers containing NGAL had the best predictive accuracy for the occurrence of AKI, regardless of whether or not the values were adjusted by urinary creatinine, and especially in medically treated patients. However, the predictive performance of urinary NGAL was limited in surgical patients, and urinary NGAL/creatinine seemed to be the most accurate biomarkers in these patients. All of the biomarkers had similar predictive performance in critically ill patients.
Trial registration CRD42020207883, October 06, 2020.
Similar content being viewed by others
Background
Acute kidney injury (AKI) is associated with a higher risk of chronic kidney disease (CKD), end-stage renal disease (ESRD), and long-term adverse cardiovascular effects [1, 2]. Due to the lack of effective treatment for impaired kidney function, the best strategy in clinical practice is to identify AKI as early as possible, reverse its cause, and even improve the sequelae. In the past decades, several serum creatinine (SCr)-based classification systems have been proposed to define AKI [3]. Serum creatinine has traditionally served as a surrogate of kidney function, despite its limitations as a diagnostic surrogate of AKI [4]. The limitations of SCr include a lack of steady-state conditions in critically ill patients, and that the determinants of SCr (rate of production, apparent volume of distribution, and rate of elimination) are variable. Therefore, there is an unmet need for other objective measures to help detect AKI in a timely manner. The role of several biomarkers in the early prediction or risk assessment of AKI has been proposed, including kidney tubular damage markers (e.g., neutrophil gelatinase-associated lipocalin (NGAL), kidney injury molecule-1 (KIM-1), liver-type fatty acid-binding protein (L-FABP)) [5,6,7,8,9], inflammation markers (e.g., interleukin-18 (IL-18)) [6, 10, 11], and stress markers (e.g., tissue inhibitor of metalloproteinases-2 and insulin-like growth factor-binding protein-7 (TIMP-2 × IGFBP-7)). The ADQI expert group suggests that routine clinical assessments should be combined with stress, damage, and functional biomarkers to stratify risk, discriminate etiologies, assess severity, plan management, and predict the duration and recovery of AKI [12]. In addition, previous meta-analyses including patients with various clinical scenarios have suggested that these biomarkers hold promise as practical tools in the early prediction of AKI [5, 13,14,15,16,17]. However, few studies have compared the diagnostic accuracy of these AKI biomarkers, and systematic assessments of the quality of evidence, which can provide updated information for clinical guidelines, are lacking. Therefore, the aim of this study was to compare the reported predictive accuracy of AKI biomarkers in various clinical settings and appraise the quality of evidence using a pairwise meta-analysis. The findings of this study may be used to update guidelines and recommendations.
Methods
Search strategy and selection criteria
We conducted this pairwise meta-analysis according to the Preferred Reporting Items of Systematic Reviews and Meta-Analyses (PRISMA) statement [18] and used Cochrane methods [19]. We prospectively submitted the systematic review protocol for registration on PROSPERO [CRD42020207883].
Data sources and search strategy
The primary outcome was incident AKI. Electronic searches were performed on PubMed (Ovid), Medline, Embase, and Cochrane library from inception to August 15, 2022 (Additional file 1: Appendix). We screened references by titles and abstracts and included related studies for further analysis. Reference lists of related studies, systematic reviews, and meta-analyses were manually examined to identify any possible publications relevant to our analysis. Both abstracts and full papers were selected for quality assessment and data synthesis.
Inclusion and exclusion criteria
The inclusion criteria were as follows: (1) clinical studies that included participants over 18 years of age and of any ethnic origin or sex; (2) studies that reported candidate AKI biomarkers including NGAL, KIM-1, L-FABP, IL-18, and TIMP-2 × IGFBP-7; and (3) studies that assessed the occurrence of incident AKI. The exclusion criteria were as follows: (1) studies including patients who had previously received dialysis; (2) studies including pregnant or lactating patients; (3) letters, conference or case reports; and (4) studies that lacked data on sensitivity or specificity of biomarkers to predict the occurrence of AKI. Only regular full papers were selected for quality assessment and data synthesis. We contacted the authors of abstracts for further detailed information, if available.
Study selection and data extraction
Six investigators (Heng-Chih Pan, Terry Ting-Yu Chiou, Chih-Chung Shiao, Che-Hsiung Wu, Hugo You-Hsien Lin, and Ming-Jen Chan) independently reviewed the search results and identified eligible studies. Any resulting discrepancies were resolved by discussion with a seventh investigator (Vin-Cent Wu). All relevant data were independently extracted from the included studies by eight investigators (Heng-Chih Pan, Chih-Chung Shiao, Terry Ting-Yu Chiou, Yih-Ting Chen, Chun-Te Huang, Ya-Fei Yang, Shu-Chen Yu, and Zi-Ming Chen) according to a standardized form. Extracted data included study characteristics (lead author, publication year, population setting, biomarkers, study endpoint, sample size, events, timing of measurements) and participants’ baseline data (mean age (years), gender (%), comorbidities, severity of illness). When available, odds ratios and 95% confidence intervals (CIs) from cohort or case-controlled studies were extracted. Other a priori determined parameters included the type of intensive care unit (ICU) setting (surgical/mixed or medical), criteria used to diagnose AKI and severe AKI, cohort size, and the presence of sepsis. Any disagreements were resolved by discussion with the investigators (Heng-Chih Pan and Vin-Cent Wu).
Quality assessment
The Quality Assessment of Diagnostic Accuracy Studies-2 (QUADAS-2) tool was used to assess the quality of each included study [20, 21]. The following 4 domains were assessed: patient selection, index test, reference standard, and flow and timing. Any disagreements in the quality assessment were resolved by discussion and consensus [15].
Pre-specified subgroup analysis
We hypothesized that the following factors could have high impacts on patient outcomes observed among different studies: clinical setting (ICU/non-ICU), patient population (surgical versus mixed/medical), whether the studies only included patients with sepsis or not and different AKI criteria (risk, injury, failure, loss, ESRD (RIFLE); Acute Kidney Injury Network (AKIN); Kidney Disease: Improving Global Outcomes (KDIGO)).
Data synthesis and statistical analysis
A 2 by 2 table reporting the patient number of true positive, false positive, true negative, and false negative findings for the cutoff point given by the included studies was used to generate sensitivity, specificity, and diagnostic odds ratio (DOR) for each study. The sensitivity, specificity, and DOR for all of the included studies were combined using a bivariate model. DOR was defined as the endpoint of primary interest in this study because it combines the strengths of sensitivity and specificity with the advantage of accuracy as a single indicator [22]. The sensitivity and specificity were defined as the endpoints of secondary interest in the study. The diagnostic performance for AKI among the 12 different biomarkers was compared using a bivariate model in which the type of biomarker was treated as a categorical covariate. Hierarchical summary receiver operating characteristic curves (HSROCs), which consider the threshold effect [23], were used to illustrate the overall diagnostic performance for each biomarker. The analysis was further stratified by the following pre-specified subgroups: surgical versus mixed/medical patients, ICU/non-ICU patients, sepsis/non-sepsis patients, and different AKI criteria (RIFLE/AKIN/KDIGO). In the subgroup analysis, biomarkers only reported in 1 study could not be compared and were therefore excluded. Potential publication bias was assessed visually using funnel plots. A two-sided P value < 0.05 was considered statistically significant. The bivariate model was conducted using SAS version 9.4 (SAS Institute, Cary, NC) with the “METADAS” macro (version 1.3) which is recommended by the Cochrane Diagnostic Test Accuracy Working Group. The HSROC analysis and funnel plots were performed using R software version 3.6.3 with the “meta4diag” package (version 2.0.8) based on Bayesian inference.
Results
Search results and study characteristics
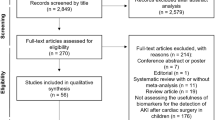
The study selection process is summarized in Additional file 1: Appendix. A total of 23,882 articles were identified through the electronic search, and after excluding duplicate and non-relevant articles, the titles and abstracts of the remaining 1803 articles were screened. A total of 242 studies were eligible for full-text review, of which 110 studies including 38,725 patients reported data on the occurrence of AKI with any one of the biomarkers of interest and were included in the meta-analysis [24,25,26,27,28,140]. In critically ill or surgical patients, the potential benefits of reducing kidney injury-related complications may outweigh the loss caused by over-monitoring the patient, such as related length of stay. Appropriate biomarkers should improve the detection rate of AKI with high sensitivity and good negative predictive value, thus enabling timely initiation of preventive strategies for AKI [141]. Previous investigations have reported that TIMP-2 × IGFBP-7 was a good biomarker to identify patients who will develop AKI and reduce the need for renal replacement therapy [136, 137, 142]. As demonstrated in the present study, NGAL/Cr, L-FABP/Cr, and TIMP-2 × IGFBP-7: custom seemed to have good predictive performance in the setting of critically ill patients, while NGAL/Cr and KIM-1 were the best biomarkers in surgical patients (Tables 4, 5).
In non-critically ill or medical patients, patient stratification for the risk of AKI should be applied to the entire hospital population before any scheduled elective intervention. In order to minimize unnecessary impacts due to these scheduled treatments, the specificity should outweigh the sensitivity [141]. In our study, the clinical performance of TIMP-2 × IGFBP-7 with a cutoff value of 2 was significantly better than that of TIMP-2 × IGFBP-7 with a cutoff value of 0.3 in the medical patients. Urinary NGAL, KIM-1, and serum NGAL seemed to be the best biomarkers in the setting of non-critically ill patients and medical patients (Tables 4, 5).
However, the sensitivity and specificity in the enrolled studies were heterogeneous because they depended on the circumstances and the threshold effects of the biomarkers. Considering the potential threshold effects and the correlation between sensitivity and specificity, HSROC analysis proved the good predictive performance of L-FABP/Cr and the NGAL series (Fig. 1A). There were differences in the applied diagnostic criteria for AKI between the enrolled studies. The subgroup analysis also demonstrated that the relative diagnostic accuracy of the AKI biomarkers remained consistent in the studies using current standard AKI criteria (RIFLE/AKIN/KDIGO) (Table 6). NGAL series seemed to have the best predictive performance for AKI, especially in the high-quality studies and in the studies which were conducted in high-income countries. Other biomarkers outperformed the NGAL series only in low- or moderate-quality studies or in the studies conducted in middle- or low-income countries (Additional file 1: Tables S2-S3). Sensitivity analysis also demonstrated the good predictive performance of serum NGAL, urinary NGAL, and TIMP-2 × IGFBP-7: custom for early onset AKI (AKI developed within 48 h) and severe AKI (stage 2–3 or renal replacement therapy) (Additional file 1: Tables S4-S6). These findings enhance the robustness of the study results.
Although the damage and stress biomarkers in this study had good predictive performance, unlike troponin in acute coronary syndrome, none of the reported biomarkers are completely specific for AKI. Previous studies have reported that NGAL, IL-18, and KIM-1 may be elevated in the setting of sepsis and CKD [143,144,145,146]. Of note, these biomarkers can be used to recruit more homogenous patient populations when implementing a clinical trial [147]. Biomarkers to identify and characterize AKI sub-types are necessary and may have the potential to provide individualized timely etiology-based management of AKI. In addition, considering the complex and multifactorial etiology of AKI, a panel of multiple biomarkers including stress, injury, and kidney reserve biomarkers could provide better discrimination for AKI. Furthermore, more kidney tissue-specific markers may help localize and quantify the severity of AKI and provide a deeper understanding of the pathophysiology of AKI. These biomarkers may offer opportunities for personalized management of AKI and support the call for a refinement of the existing AKI criteria.
Strengths and limitations
The strength of our analysis is the extensive literature search of related studies. We used standard Cochrane protocols and included the largest cumulative study sample size to date in comparison with previous reports. The strength of our meta-analysis also lies in the comprehensive data search with subgroup analyses across several clinical scenarios. We used the GRADE approach to rate the certainty of evidence [148].
Besides limitations in the meta-analysis, there were several limitations in the individual studies. First, most studies had a small sample size, and this contributed to the high heterogeneity of the meta-analysis. Second, our funnel meta-regression and Cochrane Collaboration tool analysis showed significant publication bias (Additional file 1: Appendix). Third, in some scenarios, the limited number of enrolled studies, such as trials focusing on sepsis, made subgroup analysis difficult. Of note, these new biomarkers are most effective in conditions where the time of renal insult is known, for instance, post-cardiac surgery or coronary angiography, compared to situations where the onset of kidney injury is less clear, for instance, in sepsis. To ensure the robustness of the findings, we did not emphasize the diagnostic accuracy of biomarkers extracted from fewer than three articles. Fourth, we did not perform additional analyses to assess the additional predictive value of SCr levels. Most of the included studies did not measure SCr levels with biomarkers to predict AKI. In the literature, SCr has poor predictive performance for AKI due to delayed rise and cannot accurately estimate the timing of injury [118, 127]. Traditionally, the diagnosis of AKI is based on a rise in serum creatinine and the creatinine could be hard to wear two hats, having an administrative role as well as patrolling the beat. Furthermore, the use of SCr as a comparison has several limitations and limits the full interpretation of biomarker performance. For example, SCr may be elevated in pre-renal azotemia, which is not true for renal tissue damage, and biomarkers may not be elevated. On the other hand, in the setting of true renal injury with fluid overload, biomarkers may be elevated but SCr may remain unchanged, which may underestimate the predictive performance of biomarkers [149, 150]. Fifth, the kits for specific biomarker analysis varies among the studies, so it was difficult to determine the optimal cutoff value of biomarkers to predict AKI. Sixth, the occurrence of AKI was diagnosed according to several different criteria in the enrolled studies. However, the KDIGO classification was the mostly commonly used, which has been proposed to provide a uniform definition of AKI, essentially combining the RIFLE and AKIN criteria. Finally, the definition of AKI varied between the studies, and this may have unduly influenced pooled effect estimates. Nonetheless, our conclusions were drawn from studies with different study designs and different clinical scenarios. Further research efforts are certainly needed for the pursuit of better precision medicine, especially with regard to the use of multiple biomarkers. It could be more fruitful to investigate whether different etiologies of AKI (pre-renal versus renal versus obstructive, cardiogenic shock, hypovolemic shock, sepsis-related, etc.) affect the predictive accuracy of biomarkers, and to evaluate whether the efficacy of biomarkers is affected by the severity of AKI. These issues can be incorporated into the design of future randomized controlled trials to evaluate the optimal biomarkers for different clinical settings in order to improve the timely diagnosis of AKI. Moreover, further investigations to improve the diagnosis and manage the underlying mechanisms of AKI may help to mitigate the current high mortality rate of patients with AKI.
Conclusion
Based on our pairwise meta-analysis of biomarkers to predict AKI, NGAL series had the best diagnostic accuracy for the prediction of AKI, regardless of whether or not they were adjusted by urinary creatinine, especially in medical patients. However, the predictive performance of urinary NGAL was limited in surgical patients, and NGAL/Cr seemed to be the best biomarkers in these patients. All of the biomarkers had similar predictive performance in critically ill patients. Future pragmatic clinical trials are warranted to evaluate the real-world predictive accuracy of AKI biomarkers.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- AKI:
-
Acute kidney injury
- AKIN:
-
Acute Kidney Injury Network
- CKD:
-
Chronic kidney disease
- CI:
-
Confidence interval
- DOR:
-
Diagnostic odds ratio
- ESRD:
-
End-stage renal disease
- HSROC:
-
Hierarchical summary receiver operating characteristic curve
- ICU:
-
Intensive care unit
- IL-18:
-
Interleukin-18
- KDIGO:
-
Kidney Disease: Improving Global Outcomes
- KIM-1:
-
Kidney injury molecule-1
- L-FABP:
-
Liver-type fatty acid-binding protein
- NGAL:
-
Neutrophil gelatinase-associated lipocalin
- OR:
-
Odds ratio
- PRISMA:
-
Preferred Reporting Items of Systematic Reviews and Meta-Analyses
- RIFLE:
-
Risk, injury, failure, loss, ESRD
- SCr:
-
Serum creatinine
- TIMP-2 × IGFBP-7:
-
Tissue inhibitor of metalloproteinases-2 × insulin-like growth factor-binding protein-7
References
Wu VC, Shiao CC, Chang CH, Huang TM, Lai CF, Lin MC, Chiang WC, Chu TS, Wu KD, Ko WJ, et al. Long-term outcomes after dialysis-requiring acute kidney injury. Biomed Res Int. 2014;2014:365186.
Wu VC, Wu CH, Huang TM, Wang CY, Lai CF, Shiao CC, Chang CH, Lin SL, Chen YY, Chen YM, et al. Long-term risk of coronary events after AKI. J Am Soc Nephrol. 2014;25(3):595–605.
Khwaja A. KDIGO clinical practice guidelines for acute kidney injury. Nephron Clin Pract. 2012;120(4):c179–84.
Dennen P, Douglas IS, Anderson R. Acute kidney injury in the intensive care unit: an update and primer for the intensivist. Crit Care Med. 2010;38(1):261–75.
Susantitaphong P, Siribamrungwong M, Doi K, Noiri E, Terrin N, Jaber BL. Performance of urinary liver-type fatty acid–binding protein in acute kidney injury: a meta-analysis. Am J Kidney Dis. 2013;61(3):430–9.
Zdziechowska M, Gluba-Brzózka A, Poliwczak AR, Franczyk B, Kidawa M, Zielinska M, Rysz J. Serum NGAL, KIM-1, IL-18, L-FABP: new biomarkers in the diagnostics of acute kidney injury (AKI) following invasive cardiology procedures. Int Urol Nephrol. 2020;52(11):2135–43.
Hamasaki H. Urinary liver-type fatty-acid binding protein is a predictor of mortality in individuals with type 2 diabetes. Diabet Med. 2021. https://doi.org/10.1111/dme.14527.
Wang J-J, Chi N-H, Huang T-M, Connolly R, Chen LW, Chueh S-CJ, Kan W-C, Lai C-C, Wu V-C, Fang J-T. Urinary biomarkers predict advanced acute kidney injury after cardiovascular surgery. Crit Care. 2018;22(1):1–13.
Pan H-C, Sun C-Y, Huang TT-M, Huang C-T, Tsao C-H, Lai C-H, Chen Y-M, Wu V-C. Distinct subty** of successful weaning from acute kidney injury requiring renal replacement therapy by consensus clustering in critically ill patients. Biomedicines. 2022;10(7):1628.
Forni L, Darmon M, Ostermann M, Oudemans-van Straaten H, Pettilä V, Prowle J, Schetz M, Joannidis M. Renal recovery after acute kidney injury. Intensive Care Med. 2017;43(6):855–66.
Pan H-C, Huang TT-M, Huang C-T, Sun C-Y, Chen Y-M, Wu V-C. Urinary biomarkers can predict weaning from acute dialysis therapy in critically ill patients. Arch Pathol Lab Med. 2022. https://doi.org/10.5858/arpa.2021-0411-OA.
Chawla LS, Bellomo R, Bihorac A, Goldstein SL, Siew ED, Bagshaw SM, Bittleman D, Cruz D, Endre Z, Fitzgerald RL. Acute kidney disease and renal recovery: consensus report of the Acute Disease Quality Initiative (ADQI) 16 Workgroup. Nat Rev Nephrol. 2017;13(4):241–57.
Shao X, Tian L, Xu W, Zhang Z, Wang C, Qi C, Ni Z, Mou S. Diagnostic value of urinary kidney injury molecule 1 for acute kidney injury: a meta-analysis. PLoS ONE. 2014;9(1):e84131.
Lin X, Yuan J, Zhao Y, Zha Y. Urine interleukin-18 in prediction of acute kidney injury: a systemic review and meta-analysis. J Nephrol. 2015;28(1):7–16.
Liu C, Lu X, Mao Z, Kang H, Liu H, Pan L, Hu J, Wang L, Zhou F. The diagnostic accuracy of urinary [TIMP-2]·[IGFBP7] for acute kidney injury in adults: a PRISMA-compliant meta-analysis. Medicine. 2017;96(27):e7484.
Zhang D, Yuan Y, Guo L, Wang Q. Comparison of urinary TIMP-2 and IGFBP7 cut-offs to predict acute kidney injury in critically ill patients: a PRISMA-compliant systematic review and meta-analysis. Medicine. 2019;98(26):e16232.
Albert C, Zapf A, Haase M, Röver C, Pickering JW, Albert A, Bellomo R, Breidthardt T, Camou F, Chen Z. Neutrophil gelatinase-associated lipocalin measured on clinical laboratory platforms for the prediction of acute kidney injury and the associated need for dialysis therapy: a systematic review and meta-analysis. Am J Kidney Dis. 2020;76(6):826-841.e821.
Higgins JP, Altman DG, Gotzsche PC, Juni P, Moher D, Oxman AD, Savovic J, Schulz KF, Weeks L, Sterne JA. The Cochrane Collaboration’s tool for assessing risk of bias in randomised trials. BMJ (Clin Res Ed). 2011;343:d5928.
Salguero G, Akin E, Templin C, Kotlarz D, Doerries C, Landmesser U, Grote K, Schieffer B. Renovascular hypertension by two-kidney one-clip enhances endothelial progenitor cell mobilization in a p47phox-dependent manner. J Hypertens. 2008;26(2):257–68.
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ (Clin Res Ed). 2003;327(7414):557–60.
Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, Leeflang MM, Sterne JA, Bossuyt PM. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155(8):529–36.
Glas AS, Lijmer JG, Prins MH, Bonsel GJ, Bossuyt PM. The diagnostic odds ratio: a single indicator of test performance. J Clin Epidemiol. 2003;56(11):1129–35.
Rutter CM, Gatsonis CA. A hierarchical regression approach to meta-analysis of diagnostic test accuracy evaluations. Stat Med. 2001;20(19):2865–84.
Qian Y, Che L, Yan Y, Lu R, Zhu M, Xue S, Ni Z, Gu L. Urine klotho is a potential early biomarker for acute kidney injury and associated with poor renal outcome after cardiac surgery. BMC Nephrol. 2019;20(1):268.
Prowle JR, Calzavacca P, Licari E, Ligabo EV, Echeverri JE, Bagshaw SM, Haase-Fielitz A, Haase M, Ostland V, Noiri E, et al. Combination of biomarkers for diagnosis of acute kidney injury after cardiopulmonary bypass. Ren Fail. 2015;37(3):408–16.
Lei L, Li LP, Zeng Z, Mu JX, Yang X, Zhou C, Wang ZL, Zhang H. Value of urinary KIM-1 and NGAL combined with serum Cys C for predicting acute kidney injury secondary to decompensated cirrhosis. Sci Rep. 2018;8(1):7962.
van Wolfswinkel ME, Koopmans LC, Hesselink DA, Hoorn EJ, Koelewijn R, van Hellemond JJ, van Genderen PJ. Neutrophil gelatinase-associated lipocalin (NGAL) predicts the occurrence of malaria-induced acute kidney injury. Malar J. 2016;15:464.
Srisawat N, Praditpornsilpa K, Patarakul K, Techapornrung M, Daraswang T, Sukmark T, Khositrangsikun K, Fakthongyoo A, Oranrigsupak P, Praderm L, et al. Neutrophil gelatinase associated lipocalin (NGAL) in leptospirosis acute kidney injury: a multicenter study in Thailand. PLoS ONE. 2015;10(12):e0143367.
Zeng XF, Li JM, Tan Y, Wang ZF, He Y, Chang J, Zhang H, Zhao H, Bai X, **e F, et al. Performance of urinary NGAL and L-FABP in predicting acute kidney injury and subsequent renal recovery: a cohort study based on major surgeries. Clin Chem Lab Med. 2014;52(5):671–8.
Aydogdu M, Gursel G, Sancak B, Yeni S, Sari G, Tasyurek S, Turk M, Yuksel S, Senes M, Ozis TN. The use of plasma and urine neutrophil gelatinase associated lipocalin (NGAL) and Cystatin C in early diagnosis of septic acute kidney injury in critically ill patients. Dis Markers. 2013;34(4):237–46.
Liu S, Che M, Xue S, **e B, Zhu M, Lu R, Zhang W, Qian J, Yan Y. Urinary L-FABP and its combination with urinary NGAL in early diagnosis of acute kidney injury after cardiac surgery in adult patients. Biomarkers. 2013;18(1):95–101.
Wagener G, Minhaz M, Mattis FA, Kim M, Emond JC, Lee HT. Urinary neutrophil gelatinase-associated lipocalin as a marker of acute kidney injury after orthotopic liver transplantation. Nephrol Dial Transplant. 2011;26(5):1717–23.
Makris K, Markou N, Evodia E, Dimopoulou E, Drakopoulos I, Ntetsika K, Rizos D, Baltopoulos G, Haliassos A. Urinary neutrophil gelatinase-associated lipocalin (NGAL) as an early marker of acute kidney injury in critically ill multiple trauma patients. Clin Chem Lab Med. 2009;47(1):79–82.
Constantin JM, Futier E, Perbet S, Roszyk L, Lautrette A, Gillart T, Guerin R, Jabaudon M, Souweine B, Bazin JE, et al. Plasma neutrophil gelatinase-associated lipocalin is an early marker of acute kidney injury in adult critically ill patients: a prospective study. J Crit Care. 2010;25(1):176.e1-176.e6.
Cruz DN, de Cal M, Garzotto F, Perazella MA, Lentini P, Corradi V, Piccinni P, Ronco C. Plasma neutrophil gelatinase-associated lipocalin is an early biomarker for acute kidney injury in an adult ICU population. Intensive Care Med. 2010;36(3):444–51.
de Geus HR, Bakker J, Lesaffre EM, le Noble JL. Neutrophil gelatinase-associated lipocalin at ICU admission predicts for acute kidney injury in adult patients. Am J Respir Crit Care Med. 2011;183(7):907–14.
Endre ZH, Pickering JW, Walker RJ, Devarajan P, Edelstein CL, Bonventre JV, Frampton CM, Bennett MR, Ma Q, Sabbisetti VS, et al. Improved performance of urinary biomarkers of acute kidney injury in the critically ill by stratification for injury duration and baseline renal function. Kidney Int. 2011;79(10):1119–30.
Breidthardt T, Socrates T, Drexler B, Noveanu M, Heinisch C, Arenja N, Klima T, Zusli C, Reichlin T, Potocki M, et al. Plasma neutrophil gelatinase-associated lipocalin for the prediction of acute kidney injury in acute heart failure. Crit Care. 2012;16(1):R2.
Camou F, Oger S, Paroissin C, Guilhon E, Guisset O, Mourissoux G, Pouyes H, Lalanne T, Gabinski C. Plasma neutrophil gelatinase-associated lipocalin (NGAL) predicts acute kidney injury in septic shock at ICU admission. Ann Fr Anesth Reanim. 2013;32(3):157–64.
Doi K, Urata M, Katagiri D, Inamori M, Murata S, Hisagi M, Ono M, Matsubara T, Ishii T, Yahagi N, et al. Plasma neutrophil gelatinase-associated lipocalin in acute kidney injury superimposed on chronic kidney disease after cardiac surgery: a multicenter prospective study. Crit Care. 2013;17(6):R270.
Gaipov A, Solak Y, Turkmen K, Toker A, Baysal AN, Cicekler H, Biyik Z, Erdur FM, Kilicaslan A, Anil M, et al. Serum uric acid may predict development of progressive acute kidney injury after open heart surgery. Ren Fail. 2015;37(1):96–102.
Cuartero M, Betbese AJ, Nunez K, Baldira J, Ordonez-Llanos J. Does whole-blood neutrophil gelatinase-associated lipocalin stratify acute kidney injury in critically ill patients? Dis Markers. 2019;2019:8480925.
Khawaja S, Jafri L, Siddiqui I, Hashmi M, Ghani F. The utility of neutrophil gelatinase-associated lipocalin (NGAL) as a marker of acute kidney injury (AKI) in critically ill patients. Biomark Res. 2019;7:4.
Mosa OF. Prognostic significance of serum NGAL and troponin I against acute kidney injury in Egyptian ICU patients after open heart surgery: a pilot study. Kidney Dis (Basel). 2018;4(4):246–54.
Sun IO, Shin SH, Cho AY, Yoon HJ, Chang MY, Lee KY. Clinical significance of NGAL and KIM-1 for acute kidney injury in patients with scrub typhus. PLoS ONE. 2017;12(4):e0175890.
Ghonemy TA, Amro GM. Plasma neutrophil gelatinase-associated lipocalin (NGAL) and plasma cystatin C (CysC) as biomarker of acute kidney injury after cardiac surgery. Saudi J Kidney Dis Transpl. 2014;25(3):582–8.
Padhy M, Kaushik S, Girish MP, Mohapatra S, Shah S, Koner BC. Serum neutrophil gelatinase associated lipocalin (NGAL) and cystatin C as early predictors of contrast-induced acute kidney injury in patients undergoing percutaneous coronary intervention. Clin Chim Acta. 2014;435:48–52.
de Geus HR, Betjes MG, Schaick R, Groeneveld JA. Plasma NGAL similarly predicts acute kidney injury in sepsis and nonsepsis. Biomark Med. 2013;7(3):415–21.
Haase-Fielitz A, Bellomo R, Devarajan P, Story D, Matalanis G, Dragun D, Haase M. Novel and conventional serum biomarkers predicting acute kidney injury in adult cardiac surgery–a prospective cohort study. Crit Care Med. 2009;37(2):553–60.
Hanson J, Hasan MM, Royakkers AA, Alam S, Charunwatthana P, Maude RJ, Douthwaite ST, Yunus EB, Mantha ML, Schultz MJ, et al. Laboratory prediction of the requirement for renal replacement in acute falciparum malaria. Malar J. 2011;10:217.
Introcaso G, Nafi M, Bonomi A, L’Acqua C, Salvi L, Ceriani R, Carcione D, Cattaneo A, Sandri MT. Improvement of neutrophil gelatinase-associated lipocalin sensitivity and specificity by two plasma measurements in predicting acute kidney injury after cardiac surgery. Biochem Med (Zagreb). 2018;28(3):030701.
Kim H, Hur M, Lee S, Marino R, Magrini L, Cardelli P, Struck J, Bergmann A, Hartmann O, Di Somma S, et al. Proenkephalin, neutrophil gelatinase-associated lipocalin, and estimated glomerular filtration rates in patients with sepsis. Ann Lab Med. 2017;37(5):388–97.
Ferrari F, Romero-Gonzalez G, Topete LR, Senzolo M, Lorenzin A, Husain-Syed F, Puci MV, Ferraro OE, Muraro E, Serrano-Soto M, et al. Routine adoption of urinary [IGFBP7][TIMP-2] to assess acute kidney injury at any stage 12 hours after intensive care unit admission: a prospective cohort study. Sci Rep. 2019;9(1):16484.
**e Y, Ankawi G, Yang B, Garzotto F, Passannante A, Breglia A, Digvijay K, Ferrari F, Brendolan A, Raffaele B, et al. Tissue inhibitor metalloproteinase-2 (TIMP-2) * IGF-binding protein-7 (IGFBP7) levels are associated with adverse outcomes in patients in the intensive care unit with acute kidney injury. Kidney Int. 2019;95(6):1486–93.
Adler C, Heller T, Schregel F, Hagmann H, Hellmich M, Adler J, Reuter H. TIMP-2/IGFBP7 predicts acute kidney injury in out-of-hospital cardiac arrest survivors. Crit Care. 2018;22(1):126.
Oezkur M, Magyar A, Thomas P, Stork T, Schneider R, Bening C, Stork S, Heuschmann PU, Leyh RG, Wagner M. TIMP-2*IGFBP7 (Nephrocheck(R)) measurements at intensive care unit admission after cardiac surgery are predictive for acute kidney injury within 48 hours. Kidney Blood Press Res. 2017;42(3):456–67.
Wang Y, Zou Z, ** J, Teng J, Xu J, Shen B, Jiang W, Zhuang Y, Liu L, Luo Z, et al. Urinary TIMP-2 and IGFBP7 for the prediction of acute kidney injury following cardiac surgery. BMC Nephrol. 2017;18(1):177.
Finge T, Bertran S, Roger C, Candela D, Pereira B, Scott C, Muller L, Louart B, Lefrant JY. Interest of urinary [TIMP-2] x [IGFBP-7] for predicting the occurrence of acute kidney injury after cardiac surgery: a gray zone Approach. Anesth Analg. 2017;125(3):762–9.
Cuartero M, Ballus J, Sabater J, Perez X, Nin N, Ordonez-Llanos J, Betbese AJ. Cell-cycle arrest biomarkers in urine to predict acute kidney injury in septic and non-septic critically ill patients. Ann Intensive Care. 2017;7(1):92.
Mayer T, Bolliger D, Scholz M, Reuthebuch O, Gregor M, Meier P, Grapow M, Seeberger MD, Fassl J. Urine biomarkers of tubular renal cell damage for the prediction of acute kidney injury after cardiac surgery-a pilot study. J Cardiothorac Vasc Anesth. 2017;31(6):2072–9.
Meersch M, Schmidt C, Van Aken H, Martens S, Rossaint J, Singbartl K, Gorlich D, Kellum JA, Zarbock A. Urinary TIMP-2 and IGFBP7 as early biomarkers of acute kidney injury and renal recovery following cardiac surgery. PLoS ONE. 2014;9(3):e93460.
Dusse F, Edayadiyil-Dudasova M, Thielmann M, Wendt D, Kahlert P, Demircioglu E, Jakob H, Schaefer ST, Pilarczyk K. Early prediction of acute kidney injury after transapical and transaortic aortic valve implantation with urinary G1 cell cycle arrest biomarkers. BMC Anesthesiol. 2016;16:76.
Gunnerson KJ, Shaw AD, Chawla LS, Bihorac A, Al-Khafaji A, Kashani K, Lissauer M, Shi J, Walker MG, Kellum JA, et al. TIMP2*IGFBP7 biomarker panel accurately predicts acute kidney injury in high-risk surgical patients. J Trauma Acute Care Surg. 2016;80(2):243–9.
Wetz AJ, Richardt EM, Wand S, Kunze N, Schotola H, Quintel M, Brauer A, Moerer O. Quantification of urinary TIMP-2 and IGFBP-7: an adequate diagnostic test to predict acute kidney injury after cardiac surgery? Crit Care. 2015;19:3.
Kimmel M, Shi J, Wasser C, Biegger D, Alscher MD, Schanz MB. Urinary [TIMP-2].[IGFBP7]—novel biomarkers to predict acute kidney injury. Am J Nephrol. 2016;43(5):375–82.
Pilarczyk K, Edayadiyil-Dudasova M, Wendt D, Demircioglu E, Benedik J, Dohle DS, Jakob H, Dusse F. Urinary [TIMP-2]*[IGFBP7] for early prediction of acute kidney injury after coronary artery bypass surgery. Ann Intensive Care. 2015;5(1):50.
Hoste EA, McCullough PA, Kashani K, Chawla LS, Joannidis M, Shaw AD, Feldkamp T, Uettwiller-Geiger DL, McCarthy P, Shi J, et al. Derivation and validation of cutoffs for clinical use of cell cycle arrest biomarkers. Nephrol Dial Transplant. 2014;29(11):2054–61.
Cummings JJ, Shaw AD, Shi J, Lopez MG, O’Neal JB. Billings FTt: intraoperative prediction of cardiac surgery-associated acute kidney injury using urinary biomarkers of cell cycle arrest. J Thorac Cardiovasc Surg. 2019;157(4):1545-1553.e1545.
Katagiri D, Doi K, Honda K, Negishi K, Fujita T, Hisagi M, Ono M, Matsubara T, Yahagi N, Iwagami M, et al. Combination of two urinary biomarkers predicts acute kidney injury after adult cardiac surgery. Ann Thorac Surg. 2012;93(2):577–83.
Doi K, Negishi K, Ishizu T, Katagiri D, Fujita T, Matsubara T, Yahagi N, Sugaya T, Noiri E. Evaluation of new acute kidney injury biomarkers in a mixed intensive care unit. Crit Care Med. 2011;39(11):2464–9.
Ferguson MA, Vaidya VS, Waikar SS, Collings FB, Sunderland KE, Gioules CJ, Bonventre JV. Urinary liver-type fatty acid-binding protein predicts adverse outcomes in acute kidney injury. Kidney Int. 2010;77(8):708–14.
Li Y, Zhu M, **a Q, Wang S, Qian J, Lu R, Che M, Dai H, Wu Q, Ni Z, et al. Urinary neutrophil gelatinase-associated lipocalin and L-type fatty acid binding protein as diagnostic markers of early acute kidney injury after liver transplantation. Biomarkers. 2012;17(4):336–42.
Manabe K, Kamihata H, Motohiro M, Senoo T, Yoshida S, Iwasaka T. Urinary liver-type fatty acid-binding protein level as a predictive biomarker of contrast-induced acute kidney injury. Eur J Clin Invest. 2012;42(5):557–63.
Matsui K, Kamijo-Ikemori A, Sugaya T, Yasuda T, Kimura K. Usefulness of urinary biomarkers in early detection of acute kidney injury after cardiac surgery in adults. Circ J. 2012;76(1):213–20.
Khreba NA, Abdelsalam M, Wahab AM, Sanad M, Elhelaly R, Adel M, El-Kannishy G. Kidney injury molecule 1 (KIM-1) as an early predictor for acute kidney injury in post-cardiopulmonary bypass (CPB) in open heart surgery patients. Int J Nephrol. 2019;2019:6265307.
Tu Y, Wang H, Sun R, Ni Y, Ma L, Xv F, Hu X, Jiang L, Wu A, Chen X, et al. Urinary netrin-1 and KIM-1 as early biomarkers for septic acute kidney injury. Ren Fail. 2014;36(10):1559–63.
Parikh CR, Abraham E, Ancukiewicz M, Edelstein CL. Urine IL-18 is an early diagnostic marker for acute kidney injury and predicts mortality in the intensive care unit. J Am Soc Nephrol. 2005;16(10):3046–52.
Parikh CR, Jani A, Melnikov VY, Faubel S, Edelstein CL. Urinary interleukin-18 is a marker of human acute tubular necrosis. Am J Kidney Dis. 2004;43(3):405–14.
Han WK, Wagener G, Zhu Y, Wang S, Lee HT. Urinary biomarkers in the early detection of acute kidney injury after cardiac surgery. Clin J Am Soc Nephrol. 2009;4(5):873–82.
Liangos O, Tighiouart H, Perianayagam MC, Kolyada A, Han WK, Wald R, Bonventre JV, Jaber BL. Comparative analysis of urinary biomarkers for early detection of acute kidney injury following cardiopulmonary bypass. Biomarkers. 2009;14(6):423–31.
Naggar GFE, Srogy HAE, Fathy SM. Kidney Injury Molecule -1 (KIM-1): an early novel biomarker for Acute Kidney Injury (AKI) in critically—ill patients. Life Sci J. 2012;9(4):3937–43.
Nickolas TL, Schmidt-Ott KM, Canetta P, Forster C, Singer E, Sise M, Elger A, Maarouf O, Sola-Del Valle DA, O’Rourke M, et al. Diagnostic and prognostic stratification in the emergency department using urinary biomarkers of nephron damage: a multicenter prospective cohort study. J Am Coll Cardiol. 2012;59(3):246–55.
Vaidya VS, Waikar SS, Ferguson MA, Collings FB, Sunderland K, Gioules C, Bradwin G, Matsouaka R, Betensky RA, Curhan GC, et al. Urinary biomarkers for sensitive and specific detection of acute kidney injury in humans. Clin Transl Sci. 2008;1(3):200–8.
Nisula S, Yang R, Poukkanen M, Vaara ST, Kaukonen KM, Tallgren M, Haapio M, Tenhunen J, Korhonen AM, Pettila V, et al. Predictive value of urine interleukin-18 in the evolution and outcome of acute kidney injury in critically ill adult patients. Br J Anaesth. 2015;114(3):460–8.
Nickolas TL, O’Rourke MJ, Yang J, Sise ME, Canetta PA, Barasch N, Buchen C, Khan F, Mori K, Giglio J, et al. Sensitivity and specificity of a single emergency department measurement of urinary neutrophil gelatinase-associated lipocalin for diagnosing acute kidney injury. Ann Intern Med. 2008;148(11):810–9.
Cho E, Yang HN, Jo SK, Cho WY, Kim HK. The role of urinary liver-type fatty acid-binding protein in critically ill patients. J Korean Med Sci. 2013;28(1):100–5.
Park HS, Kim JW, Lee KR, Hong DY, Park SO, Kim SY, Kim JY, Han SK. Urinary neutrophil gelatinase-associated lipocalin as a biomarker of acute kidney injury in sepsis patients in the emergency department. Clin Chim Acta. 2019;495:552–5.
Perry TE, Muehlschlegel JD, Liu KY, Fox AA, Collard CD, Shernan SK, Body SC, Investigators CG. Plasma neutrophil gelatinase-associated lipocalin and acute postoperative kidney injury in adult cardiac surgical patients. Anesth Analg. 2010;110(6):1541–7.
Shapiro NI, Trzeciak S, Hollander JE, Birkhahn R, Otero R, Osborn TM, Moretti E, Nguyen HB, Gunnerson K, Milzman D, et al. The diagnostic accuracy of plasma neutrophil gelatinase-associated lipocalin in the prediction of acute kidney injury in emergency department patients with suspected sepsis. Ann Emerg Med. 2010;56(1):52-59.e51.
Thanakitcharu P, Jirajan B. Determination of urinary neutrophil gelatinase-associated lipocalin (NGAL) cut-off level for early detection of acute kidney injury in Thai adult patients undergoing open cardiac surgery. J Med Assoc Thai. 2014;97(Suppl 11):S48–55.
Valette X, Savary B, Nowoczyn M, Daubin C, Pottier V, Terzi N, Seguin A, Fradin S, Charbonneau P, Hanouz JL, et al. Accuracy of plasma neutrophil gelatinase-associated lipocalin in the early diagnosis of contrast-induced acute kidney injury in critical illness. Intensive Care Med. 2013;39(5):857–65.
Varela CF, Greloni G, Schreck C, Bratti G, Medina A, Marenchino R, Pizarro R, Belziti C, Rosa-Diez G. Assessment of fractional excretion of urea for early diagnosis of cardiac surgery associated acute kidney injury. Ren Fail. 2015;37(10):327–31.
Chen TH, Chang CH, Lin CY, Jenq CC, Chang MY, Tian YC, Hung CC, Fang JT, Yang CW, Wen MS, et al. Acute kidney injury biomarkers for patients in a coronary care unit: a prospective cohort study. PLoS ONE. 2012;7(2):e32328.
Nisula S, Yang R, Kaukonen KM, Vaara ST, Kuitunen A, Tenhunen J, Pettila V, Korhonen AM, Group FS. The urine protein NGAL predicts renal replacement therapy, but not acute kidney injury or 90-day mortality in critically ill adult patients. Anesth Analg. 2014;119(1):95–102.
Maisel AS, Wettersten N, van Veldhuisen DJ, Mueller C, Filippatos G, Nowak R, Hogan C, Kontos MC, Cannon CM, Muller GA, et al. Neutrophil gelatinase-associated lipocalin for acute kidney injury during acute heart failure hospitalizations: the AKINESIS study. J Am Coll Cardiol. 2016;68(13):1420–31.
Matsa R, Ashley E, Sharma V, Walden AP, Keating L. Plasma and urine neutrophil gelatinase-associated lipocalin in the diagnosis of new onset acute kidney injury in critically ill patients. Crit Care. 2014;18(4):R137.
Munir MU, Khan DA, Khan FA, Shahab Naqvi SM. Rapid detection of acute kidney injury by urinary neutrophil gelatinase-associated lipocalin after cardiopulmonary bypass surgery. J Coll Physicians Surg Pak. 2013;23(2):103–6.
Onk OA, Onk D, Ozcelik F, Gunay M, Turkmen K. Risk factors for acute kidney injury after coronary artery bypass surgery and its detection using neutrophil gelatinase-associated lipocalin. Cardiorenal Med. 2016;6(3):216–29.
Md Ralib A, Mat Nor MB, Pickering JW. Plasma neutrophil gelatinase-associated lipocalin diagnosed acute kidney injury in patients with systemic inflammatory disease and sepsis. Nephrology (Carlton). 2017;22(5):412–9.
Yang CH, Chang CH, Chen TH, Fan PC, Chang SW, Chen CC, Chu PH, Chen YT, Yang HY, Yang CW, et al. Combination of urinary biomarkers improves early detection of acute kidney injury in patients with heart failure. Circ J. 2016;80(4):1017–23.
Ueta K, Watanabe M, Iguchi N, Uchiyama A, Shirakawa Y, Kuratani T, Sawa Y, Fu**o Y. Early prediction of acute kidney injury biomarkers after endovascular stent graft repair of aortic aneurysm: a prospective observational study. J Intensive Care. 2014;2:45.
Chang CH, Yang CH, Yang HY, Chen TH, Lin CY, Chang SW, Chen YT, Hung CC, Fang JT, Yang CW, et al. Urinary biomarkers improve the diagnosis of intrinsic acute kidney injury in coronary care units. Medicine (Baltimore). 2015;94(40):e1703.
Hjortrup PB, Haase N, Treschow F, Moller MH, Perner A. Predictive value of NGAL for use of renal replacement therapy in patients with severe sepsis. Acta Anaesthesiol Scand. 2015;59(1):25–34.
Chen YT, Jenq CC, Hsu CK, Yu YC, Chang CH, Fan PC, Pan HC, Wu IW, Cherng WJ, Chen YC. Acute kidney disease and acute kidney injury biomarkers in coronary care unit patients. BMC Nephrol. 2020;21(1):207.
Wybraniec MT, Chudek J, Bozentowicz-Wikarek M, Mizia-Stec K. Prediction of contrast-induced acute kidney injury by early post-procedural analysis of urinary biomarkers and intra-renal Doppler flow indices in patients undergoing coronary angiography. J Interv Cardiol. 2017;30(5):465–72.
Sinkala M, Zulu M, Kaile T, Simakando M, Chileshe C, Kafita D, Nkhoma P. Performance characteristics of kidney injury molecule-1 in relation to creatinine, urea, and microalbuminuria in the diagnosis of kidney disease. Int J Appl Basic Med Res. 2017;7(2):94–9.
Torregrosa I, Montoliu C, Urios A, Andres-Costa MJ, Gimenez-Garzo C, Juan I, Puchades MJ, Blasco ML, Carratala A, Sanjuan R, et al. Urinary KIM-1, NGAL and L-FABP for the diagnosis of AKI in patients with acute coronary syndrome or heart failure undergoing coronary angiography. Heart Vessels. 2015;30(6):703–11.
Tekce BK, Uyeturk U, Tekce H, Uyeturk U, Aktas G, Akkaya A. Does the kidney injury molecule-1 predict cisplatin-induced kidney injury in early stage? Ann Clin Biochem. 2015;52(Pt 1):88–94.
Torregrosa I, Montoliu C, Urios A, Elmlili N, Puchades MJ, Solis MA, Sanjuan R, Blasco ML, Ramos C, Tomas P, et al. Early biomarkers of acute kidney failure after heart angiography or heart surgery in patients with acute coronary syndrome or acute heart failure. Nefrologia. 2012;32(1):44–52.
Matsui K, Kamijo-Ikemori A, Hara M, Sugaya T, Kodama T, Fujitani S, Taira Y, Yasuda T, Kimura K. Clinical significance of tubular and podocyte biomarkers in acute kidney injury. Clin Exp Nephrol. 2011;15(2):220–5.
Parikh CR, Coca SG, Thiessen-Philbrook H, Shlipak MG, Koyner JL, Wang Z, Edelstein CL, Devarajan P, Patel UD, Zappitelli M, et al. Postoperative biomarkers predict acute kidney injury and poor outcomes after adult cardiac surgery. J Am Soc Nephrol. 2011;22(9):1748–57.
Wang C, Zhang J, Han J, Yang Q, Liu J, Liang B. The level of urinary IL-18 in acute kidney injury after cardiopulmonary bypass. Exp Ther Med. 2017;14(6):6047–51.
Haase-Fielitz A, Bellomo R, Devarajan P, Bennett M, Story D, Matalanis G, Frei U, Dragun D, Haase M. The predictive performance of plasma neutrophil gelatinase-associated lipocalin (NGAL) increases with grade of acute kidney injury. Nephrol Dial Transplant. 2009;24(11):3349–54.
Waskowski J, Pfortmueller CA, Schenk N, Buehlmann R, Schmidli J, Erdoes G, Schefold JC. (TIMP2) x (IGFBP7) as early renal biomarker for the prediction of acute kidney injury in aortic surgery (TIGER). A single center observational study. PLoS ONE. 2021;16(1):e0244658.
Imoto Y, Wakasaki A, Izumida K, Shimada H, Ohkubo K, Kawano Y, Ishikura H, Matsunaga A. Analysis of the diagnostic capabilities of urinary neutrophil gelatinase-associated lipocalin and serum procalcitonin for acute kidney injury at the early stage of critical care intensive care unit admission. J Clin Lab Anal. 2021;35(7):e23852.
Lee TH, Lee CC, Chen JJ, Fan PC, Tu YR, Yen CL, Kuo G, Chen SW, Tsai FC, Chang CH. Assessment of cardiopulmonary bypass duration improves novel biomarker detection for predicting postoperative acute kidney injury after cardiovascular surgery. J Clin Med. 2021;10(13):2741.
Szymanowicz W, Danilowicz-Szymanowicz L, Karolak W, Kowalik MM, Lango R. Brain and muscle oxygen saturation combined with kidney injury biomarkers predict cardiac surgery related acute kidney injury. Diagnostics (Basel). 2021;11(9):1591.
Zhen XW, Song NP, Ma LH, Ma LN, Guo L, Yang XD. Calprotectin and neutrophil gelatinase-associated lipocalin as biomarkers of acute kidney injury in acute coronary syndrome. Am J Med Sci. 2021;361(6):736–43.
Obata Y, Kamijo-Ikemori A, Inoue S. Clinical utility of urinary biomarkers for prediction of acute kidney injury and chronic renal dysfunction after open abdominal aortic aneurysm repair. Int J Nephrol Renovasc Dis. 2021;14:371–84.
Qiu ZL, Yan BQ, Zhao R, Xu DW, Shen K, Deng XQ, Lu SQ. Combination of hepcidin with neutrophil gelatinase-associated lipocalin for prediction of the development of sepsis-induced acute kidney injury. Clin Chim Acta. 2021;523:38–44.
Pode Shakked N, de Oliveira MHS, Cheruiyot I, Benoit JL, Plebani M, Lippi G, Benoit SW, Henry BM. Early prediction of COVID-19-associated acute kidney injury: are serum NGAL and serum Cystatin C levels better than serum creatinine? Clin Biochem. 2022;102:1–8.
Vogel MJ, Mustroph J, Staudner ST, Leininger SB, Hubauer U, Wallner S, Meindl C, Hanses F, Zimmermann M, Maier LS, et al. Kidney injury molecule-1: potential biomarker of acute kidney injury and disease severity in patients with COVID-19. J Nephrol. 2021;34(4):1007–18.
Orhon Ergun M, Zengin SU, Mustafayeva A, Umuroglu T. Neutrophil gelatinase associated lipocalin in predicting postoperative acute kidney injury in elderly. Ir J Med Sci. 2022;191(3):1297–303.
Pilarczyk K, Panholzer B, Huenges K, Salem M, Jacob T, Cremer J, Haneya A. Prediction of Acute kidney injury by Cystatin C and [TIMP-2]*[IGFBP7] after thoracic aortic surgery with moderate hypothermic circulatory arrest. J Clin Med. 2022;11(4):1024.
Okuda H, Obata Y, Kamijo-Ikemori A, Inoue S. Quantitative and qualitative analyses of urinary L-FABP for predicting acute kidney injury after emergency laparotomy. J Anesth. 2022;36(1):38–45.
Pei Y, Zhou G, Wang P, Shi F, Ma X, Zhu J. Serum cystatin C, kidney injury molecule-1, neutrophil gelatinase-associated lipocalin, klotho and fibroblast growth factor-23 in the early prediction of acute kidney injury associated with sepsis in a Chinese emergency cohort study. Eur J Med Res. 2022;27(1):39.
Jahaj E, Vassiliou AG, Pratikaki M, Gallos P, Mastora Z, Dimopoulou I, Orfanos SE, Orfanos P, Lagiou P, Kotanidou A. Serum neutrophil gelatinase-associated lipocalin (NGAL) could provide better accuracy than creatinine in predicting acute kidney injury development in critically ill patients. J Clin Med. 2021;10(22):5379.
Sampaio de Souza Garms D, Cardoso Eid KZ, Burdmann EA, Marcal LJ, Antonangelo L, Dos Santos A, Ponce D. The role of urinary biomarkers as diagnostic and prognostic predictors of acute kidney injury associated with vancomycin. Front Pharmacol. 2021;12:705636.
Irqsusi M, Beckers J, Wiesmann T, Talipov I, Ramzan R, Rastan AJ, Vogt S. Urinary TIMP-2 and IGFBP-7 protein levels as early predictors of acute kidney injury after cardiac surgery. J Card Surg. 2022;37(4):717–24.
Guray U, Coteli C, Korkmaz A, Gulkan B, Kosem A, Turker Duyuler P, Ucar Elalmis O. Value of point-of-care neutrophil gelatinase associated lipocalin in early diagnosis of acute kidney injury in patients with left ventricular systolic dysfunction after coronary angiography. Scand J Clin Lab Invest. 2021;81(7):552–6.
Tan D, Zhao L, Peng W, Wu FH, Zhang GB, Yang B, Huo WQ. Value of urine IL-8, NGAL and KIM-1 for the early diagnosis of acute kidney injury in patients with ureteroscopic lithotripsy related urosepsis. Chin J Traumatol. 2022;25(1):27–31.
Lakhal K, Bigot-Corbel E, Sacchetto E, Chabrun F, Senage T, Figueres L, Leroy M, Legrand A, Rozec B. Early recognition of cardiac surgery-associated acute kidney injury: lack of added value of TIMP2 IGFBP7 over short-term changes in creatinine (an observational pilot study). BMC Anesthesiol. 2021;21(1):244.
Sahu AK, Goel PK, Khanna R, Kumar S, Kapoor A, Tewari S, Garg N. Neutrophil gelatinase-associated lipocalin as a marker for contrast-induced nephropathy in patients undergoing percutaneous coronary intervention: a prospective observational analysis. Indian J Nephrol. 2022;32(3):247–55.
Joslin J, Wilson H, Zubli D, Gauge N, Kinirons M, Hopper A, Pile T, Ostermann M. Recognition and management of acute kidney injury in hospitalised patients can be partially improved with the use of a care bundle. Clin Med. 2015;15(5):431.
Shiao CC, Wu PC, Wu VC, Lin JH, Pan HC, Yang YF, Lai TS, Huang TM, Wu CH, Yang WS. Nationwide epidemiology and prognosis of dialysis-requiring acute kidney injury (NEP-AKI-D) study: Design and methods. Nephrology. 2016;21(9):758–64.
Ostermann M, Zarbock A, Goldstein S, Kashani K, Macedo E, Murugan R, Bell M, Forni L, Guzzi L, Joannidis M. Recommendations on acute kidney injury biomarkers from the acute disease quality initiative consensus conference: a consensus statement. JAMA Netw Open. 2020;3(10):e2019209.
Meersch M, Schmidt C, Hoffmeier A, Van Aken H, Wempe C, Gerss J, Zarbock A. Prevention of cardiac surgery-associated AKI by implementing the KDIGO guidelines in high risk patients identified by biomarkers: the PrevAKI randomized controlled trial. Intensive Care Med. 2017;43(11):1551–61.
Bonventre JV, Vaidya VS, Schmouder R, Feig P, Dieterle F. Next-generation biomarkers for detecting kidney toxicity. Nat Biotechnol. 2010;28(5):436–40.
Parikh CR, Thiessen-Philbrook H, Garg AX, Kadiyala D, Shlipak MG, Koyner JL, Edelstein CL, Devarajan P, Patel UD, Zappitelli M. Performance of kidney injury molecule-1 and liver fatty acid-binding protein and combined biomarkers of AKI after cardiac surgery. Clin J Am Soc Nephrol. 2013;8(7):1079–88.
Oh D-J. A long journey for acute kidney injury biomarkers. Ren Fail. 2020;42(1):154–65.
van Duijl T, Soonawala D, de Fijter J, Ruhaak L, Cobbaert C. Rational selection of a biomarker panel targeting unmet clinical needs in kidney injury. Clin Proteom. 2021;18(1):1–14.
Göcze I, Jauch D, Götz M, Kennedy P, Jung B, Zeman F, Gnewuch C, Graf BM, Gnann W, Banas B. Biomarker-guided intervention to prevent acute kidney injury after major surgery: the prospective randomized BigpAK study. In.: LWW; 2018.
Smith GL, Lichtman JH, Bracken MB, Shlipak MG, Phillips CO, DiCapua P, Krumholz HM. Renal impairment and outcomes in heart failure: systematic review and meta-analysis. J Am Coll Cardiol. 2006;47(10):1987–96.
Devarajan P. Emerging biomarkers of acute kidney injury. Acute Kidney Inj. 2007;156:203–12.
Rovcanin B, Savic Vujovic K, Obradovic D, Duric D, Prostran M. Evaluation of novel biomarkers of acute kidney injury: the possibilities and limitations. Curr Med Chem. 2016;23(19):1981–97.
Schrezenmeier E, Barasch J, Budde K, Westhoff T, Schmidt-Ott K. Biomarkers in acute kidney injury–pathophysiological basis and clinical performance. Acta Physiol. 2017;219(3):556–74.
Parikh CR, Moledina DG, Coca SG, Thiessen-Philbrook HR, Garg AX. Application of new acute kidney injury biomarkers in human randomized controlled trials. Kidney Int. 2016;89(6):1372–9.
Guyatt GH, Oxman AD, Kunz R, Woodcock J, Brozek J, Helfand M, Alonso-Coello P, Glasziou P, Jaeschke R, Akl EA, et al. GRADE guidelines: 7. Rating the quality of evidence–inconsistency. J Clin Epidemiol. 2011;64(12):1294–302.
Parikh CR, Mansour SG. Perspective on clinical application of biomarkers in AKI. J Am Soc Nephrol. 2017;28(6):1677–85.
Hasson D, Menon S, Gist KM. Improving acute kidney injury diagnostic precision using biomarkers. Pract Lab Med. 2022;30:e00272.
Acknowledgements
The authors acknowledge the administrative support of the Chang Gung Memorial Hospital Clinical Trial Center—which is funded by the Taiwanese Ministry of Health and Welfare (grants MOHW110-TDU-B-212-124005, MOHW111-TDU-B-212-134005). The authors thank Shu-Chen Yu, Zi-Ming Chen, all participants of NSARF and CAKs/TCTC. (Details of the members of CAKs can be downloaded here: http://links.lww.com/MD/B298.) We also thank Alfred Hsing-Fen Lin who serves in Raising Statistics Consultant Inc. for his statistical assistance during the completion of this manuscript.
Funding
The authors greatly appreciate the Second Core Lab in National Taiwan University Hospital for technical assistance. This study was supported by Ministry of Science and Technology (MOST) of the Republic of China (Taiwan) [grant number, MOST 106-2314-B-182A-064, MOST 107-2314-B-182A-138, MOST 107-2314-B-002-026-MY3, MOST 108-2314-B-182A-027, MOST108-2314-B-002-058, MOST 110-2314-B-002-241, MOST 110-2314-B-002-239, MOST 111-2314-B-182A-074-MY3], National Science and Technology Council (NSTC) [grant number, NSTC 109-2314-B-002-174-MY3, 110-2314-B-002-124-MY3, 111-2314-B-002-046, 111-2314-B-002-058], National Health Research Institutes [PH-102-SP-09], Chang Gung Memorial Foundation [CMRPG-2G0361-3, CMRPG-2H0161-5, CMRPG-2J0261, CMRPG-2K0091-3], National Taiwan University Hospital [109-S4634, PC-1246, PC-1309, VN109-09, UN109-041, UN110-030, 111-FTN0011], the Taiwan Ministry of Health and Welfare (grant number: PMRPG-2L0011, MOHW 110-TDU-B-212-124005, MOHW111-TDU-B-212-134005) and Mrs. Hsiu-Chin Lee Kidney Research Fund. The funders had no role in study design, decision to publish, data collection and analysis, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
VCW chaired the group, conceived and designed the study, performed statistical analysis, and contributed to data collection, data interpretation, and critical revision of the manuscript. HCP, YY, TYC, CCS, CHW, CTH, TJW, and JYC conducted a literature search. HWL, SYC, TMH, YFY, YHL, MJC, CYS, YTC, and YCC performed statistical analysis. HCP, SYY, TYU, and VCW wrote the manuscript and performed a critical review of the manuscript. All authors contributed to subsequent drafts and examined the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1:
Supplementary appendix.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Pan, HC., Yang, SY., Chiou, T.TY. et al. Comparative accuracy of biomarkers for the prediction of hospital-acquired acute kidney injury: a systematic review and meta-analysis. Crit Care 26, 349 (2022). https://doi.org/10.1186/s13054-022-04223-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13054-022-04223-6