Abstract
Background
Intratumoral heterogeneity is the primary challenge in the treatment of glioblastoma (GBM). The presence of glioma stem cells (GSCs) and their conversion between different molecular phenotypes contribute to the complexity of heterogeneity, culminating in preferential resistance to radiotherapy. ARP2/3 (actin-related protein-2/3) complexes (ARPs) are associated with cancer migration, invasion and differentiation, while the implications of ARPs in the phenotype and resistance to radiotherapy of GSCs remain unclear.
Methods
We screened the expression of ARPs in TCGA-GBM and CGGA-GBM databases. Tumor sphere formation assays and limiting dilution assays were applied to assess the implications of ARPC1B in tumorigenesis. Apoptosis, comet, γ-H2AX immunofluorescence (IF), and cell cycle distribution assays were used to evaluate the effect of ARPC1B on radiotherapy resistance. Immunoprecipitation (IP) and mass spectrometry analysis were used to detect ARPC1B-interacting proteins. Immune blot assays were performed to evaluate protein ubiquitination, and deletion mutant constructs were designed to determine the binding sites of protein interactions. The Spearman correlation algorithm was performed to screen for drugs that indicated cell sensitivity by the expression of ARPC1B. An intracranial xenograft GSC mouse model was used to investigate the role of ARPC1B in vivo.
Results
We concluded that ARPC1B was significantly upregulated in MES-GBM/GSCs and was correlated with a poor prognosis. Both in vitro and in vivo assays indicated that knockdown of ARPC1B in MES-GSCs reduced tumorigenicity and resistance to IR treatment, whereas overexpression of ARPC1B in PN-GSCs exhibited the opposite effects. Mechanistically, ARPC1B interacted with IFI16 and HuR to maintain protein stability. In detail, the Pyrin of IFI16 and RRM2 of HuR were implicated in binding to ARPC1B, which counteracted TRIM21-mediated degradation of ubiquitination to IFI16 and HuR. Additionally, the function of ARPC1B was dependent on IFI16-induced activation of NF-κB pathway and HuR-induced activation of STAT3 pathway. Finally, we screened AZD6738, an ataxia telangiectasia mutated and rad3-related (ATR) inhibitor, based on the expression of ARPC1B. In addition to ARPC1B expression reflecting cellular sensitivity to AZD6738, the combination of AZD6738 and radiotherapy exhibited potent antitumor effects both in vitro and in vivo.
Conclusion
ARPC1B promoted MES phenotype maintenance and radiotherapy resistance by inhibiting TRIM21-mediated degradation of IFI16 and HuR, thereby activating the NF-κB and STAT3 signaling pathways, respectively. AZD6738, identified based on ARPC1B expression, exhibited excellent anti-GSC activity in combination with radiotherapy.
Similar content being viewed by others
Background
Glioblastoma (GBM), characterized by wild-type isocitrate dehydrogenase (IDH), is the most aggressive and fatal primary brain tumor in adults [1, 2]. Despite substantial efforts, the standard of care for GBM, maximum surgical resection followed by ionizing radiation (IR) and temozolomide adjuvant therapy, has only a minor clinical benefit [3, 4]. Due to the transcriptional plasticity and heterogeneity of GBM, radiotherapy resistance and eventual recurrence are inevitable [5].
According to bulk expression profiles, there are three phenotypes of GBM, termed proneural (PN), classical (CL) and mesenchymal (MES). GBM patients with the MES phenotype exhibit worse survival and enhanced resistance to radiotherapy than patients with the PN phenotype [6, 61].
Additionally, intrinsic nuclear DNA damage also causes activation of STING and innate immune responses, which are independent of the cGAS signaling pathway. IFI16 together with ATM and PARP-1 mediates the noncanonical activation of STING. Meanwhile, activated STING signaling is further implicated in the activation of NF-κB signaling. In addition to the function of IFI16 in the STING/NF-κB axis, Cindy et al. found that activation of the ATR/CHK1 pathway was related to an increased number of cytoplasmic ssDNA and micronuclei, which further activated the IFI16/STING pathway [58]. Their work revealed the upstream ATR of IFI16, which provided insights for our subsequent drug design and analysis.
HuR is reported to counteract miR-330 to promote STAT3 translation [49]. In addition, HuR is responsible for IL-6 mRNA stability, which further activates the JAK1/STAT3 signaling pathway [62]. In terms of the regulation of HuR, Hyeon et al. reported that the function of HuR was regulated by ATM/ATR through Chk1 and Chk2 [59]. It is clear that the upstream regulation of both IFI16 and HuR is implicated in the activation of ATR. Intriguingly, the drug we screened based on the expression profile of ARPC1B, AZD6738, is also coincidentally an inhibitor of ATR.
ATR is implicated in coordinating the DNA damage response (DDR) induced by DNA replication-related stress and cell cycle checkpoints [33]. The activation of ATR is essential for fork stabilization in response to replication stress and adverse stress-induced DNA damage [63]. Therefore, targeting ATR has exhibited promising antitumor effects, and a variety of ATR inhibitors have been developed. AZD6738, an oral inhibitor of ATR, has been studied as a monotherapy or in combination with radiotherapy, chemotherapy or immunotherapy. Our results indicated that AZD6738 in combination with radiotherapy exhibited excellent antitumor activity in vitro and in vivo. As previously discussed, the regulation of IFI16 and HuR is involved in the ATR pathway, whereas AZD6738 has been validated to target ATR-induced activation of IFI16 and HuR. Therefore, AZD6738 has the potential to counteract ARPC1B-induced resistance to radiotherapy in GSCs. We demonstrated that AZD6738 was implicated in the suppression of two downstream molecules of ARPC1B, IFI16 and HuR. That is, in addition to the cellular sensitivity to AZD6738 being reflected by the expression of ARPC1B, AZD6738 also has the potential to inhibit PMT and resistance to radiotherapy by affecting IFI16 and HuR.
Conclusions
In conclusion, we identified ARPC1B, which is significantly upregulated in MES-GBM/GSCs and is correlated with a poor prognosis. ARPC1B promotes PMT and radiotherapy resistance by inhibiting TRIM21-mediated degradation of IFI16 and HuR, thereby activating the NF-κB and STAT3 signaling pathways, respectively. AZD6738 in combination with radiotherapy exhibited potent anti-GSC effects. Our findings expand the understanding of the heterogeneity and plasticity of GBM and provide a potential therapeutic strategy for GBM treatment.
Availability of data and materials
Publicly available datasets were applied in this research. These resources could be found here: TCGA: https://portal.gdc.cancer.gov/; CGGA: http://www.cgga.org.cn/; GEO: https://www.ncbi.nlm.nih.gov/gds/ (Accession number is GSE138794). Other data obtained and/or analyzed during the current study were available from the corresponding authors on reasonable request.
Abbreviations
- ARP2/3:
-
Actin-related protein-2/3
- ARPC1B:
-
Actin related protein 2/3 complex, subunit 1B
- ATR:
-
Ataxia telangiectasia mutated and rad3 related
- CGGA:
-
Chinese Glioma Genome Atlas
- CL:
-
Classical
- DDR:
-
DNA damage response
- EMT:
-
Epithelial-mesenchymal transition
- GBM:
-
Glioblastoma
- GSCs:
-
Glioma stem cells
- IDH:
-
Isocitrate dehydrogenase
- IF:
-
Immunofluorescence
- IHC:
-
Immunohistochemical
- IFI16:
-
γ-Interferon-inducible protein-16
- IR:
-
Ionizing radiation
- MES:
-
Mesenchymal
- MRs:
-
Master regulators
- NPC:
-
Neural progenitor cells
- PMT:
-
PN-to-MES transition
- PN:
-
Proneural
- TCGA:
-
The Cancer Genome Atlas
- TME:
-
Tumor microenvironment
References
Louis D, Perry A, Wesseling P, Brat D, Cree I, Figarella-Branger D, Hawkins C, Ng H, Pfister S, Reifenberger G, et al. The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro Oncol. 2021;23(8):1231–51.
Reifenberger G, Wirsching HG, Knobbe-Thomsen CB, Weller M. Advances in the molecular genetics of gliomas - implications for classification and therapy. Nat Rev Clin Oncol. 2017;14(7):434–52.
Khosla D. Concurrent therapy to enhance radiotherapeutic outcomes in glioblastoma. Ann Transl Med. 2016;4(3):54.
Shergalis A, Bankhead A, Luesakul U, Muangsin N, Neamati N. Current Challenges and Opportunities in Treating Glioblastoma. Pharmacol Rev. 2018;70(3):412–45.
Neftel C, Laffy J, Filbin MG, Hara T, Shore ME, Rahme GJ, Richman AR, Silverbush D, Shaw ML, Hebert CM, et al. An Integrative Model of Cellular States, Plasticity, and Genetics for Glioblastoma. Cell. 2019;178(4):835-849 e821.
Verhaak R, Hoadley K, Purdom E, Wang V, Qi Y, Wilkerson M, Miller C, Ding L, Golub T, Mesirov J, et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell. 2010;17(1):98–110.
Xu J, Zhang Z, Qian M, Wang S, Qiu W, Chen Z, Sun Z, **ong Y, Wang C, Sun X, et al. Cullin-7 (CUL7) is overexpressed in glioma cells and promotes tumorigenesis via NF-κB activation. J Exp Clin Cancer Res. 2020;39(1):59.
Minata M, Audia A, Shi J, Lu S, Bernstock J, Pavlyukov M, Das A, Kim S, Shin Y, Lee Y, et al. Phenotypic Plasticity of Invasive Edge Glioma Stem-like Cells in Response to Ionizing Radiation. Cell Rep. 2019;26(7):1893-1905.e1897.
Segerman A, Niklasson M, Haglund C, Bergstrom T, Jarvius M, **e Y, Westermark A, Sonmez D, Hermansson A, Kastemar M, et al. Clonal Variation in Drug and Radiation Response among Glioma-Initiating Cells Is Linked to Proneural-Mesenchymal Transition. Cell Rep. 2016;17(11):2994–3009.
Platten M. Driving mesenchymal transition in glioblastoma. Neuro Oncol. 2020;22(1):1–2.
Bao S, Wu Q, McLendon R, Hao Y, Shi Q, Hjelmeland A, Dewhirst M, Bigner D, Rich J. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature. 2006;444(7120):756–60.
Chen J, Li Y, Yu T, McKay R, Burns D, Kernie S, Parada L. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature. 2012;488(7412):522–6.
Narayanan A, Gagliardi F, Gallotti A, Mazzoleni S, Cominelli M, Fagnocchi L, Pala M, Piras I, Zordan P, Moretta N, et al. The proneural gene ASCL1 governs the transcriptional subgroup affiliation in glioblastoma stem cells by directly repressing the mesenchymal gene NDRG1. Cell Death Differ. 2019;26(9):1813–31.
Wang Z, Shi Y, Ying C, Jiang Y, Hu J. Hypoxia-induced PLOD1 overexpression contributes to the malignant phenotype of glioblastoma via NF-kappaB signaling. Oncogene. 2021;40(8):1458–75.
Carro M, Lim W, Alvarez M, Bollo R, Zhao X, Snyder E, Sulman E, Anne S, Doetsch F, Colman H, et al. The transcriptional network for mesenchymal transformation of brain tumours. Nature. 2010;463(7279):318–25.
Bhat K, Balasubramaniyan V, Vaillant B, Ezhilarasan R, Hummelink K, Hollingsworth F, Wani K, Heathcock L, James J, Goodman L, et al. Mesenchymal differentiation mediated by NF-κB promotes radiation resistance in glioblastoma. Cancer Cell. 2013;24(3):331–46.
Bhat K, Salazar K, Balasubramaniyan V, Wani K, Heathcock L, Hollingsworth F, James J, Gumin J, Diefes K, Kim S, et al. The transcriptional coactivator TAZ regulates mesenchymal differentiation in malignant glioma. Genes Dev. 2011;25(24):2594–609.
Hara T, Chanoch-Myers R, Mathewson N, Myskiw C, Atta L, Bussema L, Eichhorn S, Greenwald A, Kinker G, Rodman C, et al. Interactions between cancer cells and immune cells drive transitions to mesenchymal-like states in glioblastoma. Cancer Cell. 2021;39(6):779-792.e711.
Zhang Z, Xu J, Chen Z, Wang H, Xue H, Yang C, Guo Q, Qi Y, Guo X, Qian M, et al. Transfer of MicroRNA via Macrophage-Derived Extracellular Vesicles Promotes Proneural-to-Mesenchymal Transition in Glioma Stem Cells. Cancer Immunol Res. 2020;8(7):966–81.
Chen Z, Wang H, Zhang Z, Xu J, Qi Y, Xue H, Gao Z, Zhao R, Wang S, Zhang S, et al. Cell surface GRP78 regulates BACE2 via lysosome-dependent manner to maintain mesenchymal phenotype of glioma stem cells. J Exp Clin Cancer Res. 2021;40(1):20.
Rotty J, Wu C, Bear J. New insights into the regulation and cellular functions of the ARP2/3 complex. Nat Rev Mol Cell Biol. 2013;14(1):7–12.
Goley E, Welch M. The ARP2/3 complex: an actin nucleator comes of age. Nat Rev Mol Cell Biol. 2006;7(10):713–26.
Molinie N, Gautreau A. The Arp2/3 Regulatory System and Its Deregulation in Cancer. Physiol Rev. 2018;98(1):215–38.
Kazazian K, Go C, Wu H, Brashavitskaya O, Xu R, Dennis J, Gingras A, Swallow C. Plk4 Promotes Cancer Invasion and Metastasis through Arp2/3 Complex Regulation of the Actin Cytoskeleton. Cancer Res. 2017;77(2):434–47.
Cheng Z, Wei W, Wu Z, Wang J, Ding X, Sheng Y, Han Y, Wu Q. ARPC2 promotes breast cancer proliferation and metastasis. Oncol Rep. 2019;41(6):3189–200.
Choi J, Lee Y, Yoon Y, Kim C, Park S, Kim S, Doo Kim N, Cho Han D, Kwon B. Pimozide suppresses cancer cell migration and tumor metastasis through binding to ARPC2, a subunit of the Arp2/3 complex. Cancer Sci. 2019;110(12):3788–801.
Mondaca J, Uzair I, Castro Guijarro A, Flamini M, Sanchez A. Molecular Basis of LH Action on Breast Cancer Cell Migration and Invasion via Kinase and Scaffold Proteins. Front Cell Dev Biol. 2020;8:630147.
Wei Z, Wang R, Yin X, Zhang L, Lei Y, Zhang Y, Li Y, Wu J, Bu Y, ** G, et al. PRR11 induces filopodia formation and promotes cell motility via recruiting ARP2/3 complex in non-small cell lung cancer cells. Genes Dis. 2022;9(1):230–44.
Frentzas S, Simoneau E, Bridgeman V, Vermeulen P, Foo S, Kostaras E, Nathan M, Wotherspoon A, Gao Z, Shi Y, et al. Vessel co-option mediates resistance to anti-angiogenic therapy in liver metastases. Nat Med. 2016;22(11):1294–302.
Liu T, Zhu C, Chen X, Wu J, Guan G, Zou C, Shen S, Chen L, Cheng P, Cheng W, et al. Dual role of ARPC1B in regulating the network between tumor-associated macrophages and tumor cells in glioblastoma. Oncoimmunology. 2022;11(1):2031499.
Ji H, Zhao H, ** J, Liu Z, Gao X, Wang F, Dong J, Yan X, Zhang J, Wang N, et al. Novel Immune-Related Gene-Based Signature Characterizing an Inflamed Microenvironment Predicts Prognosis and Radiotherapy Efficacy in Glioblastoma. Front Genet. 2021;12:736187.
Liu J, Lu J, Li W. A Comprehensive Prognostic and Immunological Analysis of a Six-Gene Signature Associated With Glycolysis and Immune Response in Uveal Melanoma. Front Immunol. 2021;12:738068.
Cimprich K, Cortez D. ATR: an essential regulator of genome integrity. Nat Rev Mol Cell Biol. 2008;9(8):616–27.
Forment J, O’Connor M. Targeting the replication stress response in cancer. Pharmacol Ther. 2018;188:155–67.
Suzuki T, Hirokawa T, Maeda A, Harata S, Watanabe K, Yanagita T, Ushigome H, Nakai N, Maeda Y, Shiga K, et al. ATR inhibitor AZD6738 increases the sensitivity of colorectal cancer cells to 5-fluorouracil by inhibiting repair of DNA damage. Oncol Rep. 2022;47(4):78.
Patin EC, Dillon MT, Nenclares P, Grove L, Soliman H, Leslie I, Northcote D, Bozhanova G, Crespo-Rodriguez E, Baldock H, et al. Harnessing radiotherapy-induced NK-cell activity by combining DNA damage-response inhibition and immune checkpoint blockade. J Immunother Cancer. 2022;10(3):e004306.
Kundu K, Cardnell RJ, Zhang B, Shen L, Stewart CA, Ramkumar K, Cargill KR, Wang J, Gay CM, Byers LA. SLFN11 biomarker status predicts response to lurbinectedin as a single agent and in combination with ATR inhibition in small cell lung cancer. Transl Lung Cancer Res. 2021;10(11):4095–105.
George SL, Parmar V, Lorenzi F, Marshall LV, Jamin Y, Poon E, Angelini P, Chesler L. Novel therapeutic strategies targeting telomere maintenance mechanisms in high-risk neuroblastoma. J Exp Clin Cancer Res. 2020;39(1):78.
Wilson Z, Odedra R, Wallez Y, Wijnhoven P, Hughes A, Gerrard J, Jones G, Bargh-Dawson H, Brown E, Young L, et al. ATR Inhibitor AZD6738 (Ceralasertib) Exerts Antitumor Activity as a Monotherapy and in Combination with Chemotherapy and the PARP Inhibitor Olaparib. Cancer Res. 2022;82(6):1140–52.
Kim R, Kwon M, An M, Kim S, Smith S, Loembé A, Mortimer P, Armenia J, Lukashchuk N, Shah N, et al. Ann Oncol. 2022;33(2):193–203.
Yu H, Ding J, Zhu H, **g Y, Zhou H, Tian H, Tang K, Wang G, Wang X. LOXL1 confers antiapoptosis and promotes gliomagenesis through stabilizing BAG2. Cell Death Differ. 2020;27(11):3021–36.
Gyori B, Venkatachalam G, Thiagarajan P, Hsu D, Clement M. OpenComet: an automated tool for comet assay image analysis. Redox Biol. 2014;2:457–65.
Han S, Zhang C, Li Q, Dong J, Liu Y, Huang Y, Jiang T, Wu A. Tumour-infiltrating CD4(+) and CD8(+) lymphocytes as predictors of clinical outcome in glioma. Br J Cancer. 2014;110(10):2560–8.
Yang W, Soares J, Greninger P, Edelman E, Lightfoot H, Forbes S, Bindal N, Beare D, Smith J, Thompson I, et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 2013;41:D955-961.
Mir S, De Witt Hamer P, Krawczyk P, Balaj L, Claes A, Niers J, Van Tilborg A, Zwinderman A, Geerts D, Kaspers G, et al. In silico analysis of kinase expression identifies WEE1 as a gatekeeper against mitotic catastrophe in glioblastoma. Cancer Cell. 2010;18(3):244–57.
Li D, Wu R, Guo W, **e L, Qiao Z, Chen S, Zhu J, Huang C, Huang J, Chen B, et al. STING-Mediated IFI16 Degradation Negatively Controls Type I Interferon Production. Cell Rep. 2019;29(5):1249-1260.e1244.
Dunphy G, Flannery S, Almine J, Connolly D, Paulus C, Jønsson K, Jakobsen M, Nevels M, Bowie A, Unterholzner L. Non-canonical Activation of the DNA Sensing Adaptor STING by ATM and IFI16 Mediates NF-κB Signaling after Nuclear DNA Damage. Mol Cell. 2018;71(5):745-760.e745.
Guha A, Waris S, Nabors L, Filippova N, Gorospe M, Kwan T, King P. The versatile role of HuR in Glioblastoma and its potential as a therapeutic target for a multi-pronged attack. Adv Drug Deliv Rev. 2022;181:114082.
Mubaid S, Ma J, Omer A, Ashour K, Lian X, Sanchez B, Robinson S, Cammas A, Dormoy-Raclet V, Di Marco S, et al. HuR counteracts miR-330 to promote STAT3 translation during inflammation-induced muscle wasting. Proc Natl Acad Sci U S A. 2019;116(35):17261–70.
Grillo E, Ravelli C, Corsini M, Zammataro L, Mitola S. Protein domain-based approaches for the identification and prioritization of therapeutically actionable cancer variants. Biochim Biophys Acta Rev Cancer. 2021;1876(2):188614.
Shi D, Grossman S. Ubiquitin becomes ubiquitous in cancer: emerging roles of ubiquitin ligases and deubiquitinases in tumorigenesis and as therapeutic targets. Cancer Biol Ther. 2010;10(8):737–47.
Song Y, Wu X, Xu Y, Zhu J, Li J, Zou Z, Chen L, Zhang B, Hua C, Rui H, et al. HPV E7 inhibits cell pyroptosis by promoting TRIM21-mediated degradation and ubiquitination of the IFI16 inflammasome. Int J Biol Sci. 2020;16(15):2924–37.
Guha A, Ahuja D, Das Mandal S, Parasar B, Deyasi K, Roy D, Sharma V, Willard B, Ghosh A, Ray PS. Integrated Regulation of HuR by Translation Repression and Protein Degradation Determines Pulsatile Expression of p53 Under DNA Damage. iScience. 2019;15:342–59.
Foote K, Nissink J, McGuire T, Turner P, Guichard S, Yates J, Lau A, Blades K, Heathcote D, Odedra R, et al. Discovery and Characterization of AZD6738, a Potent Inhibitor of Ataxia Telangiectasia Mutated and Rad3 Related (ATR) Kinase with Application as an Anticancer Agent. J Med Chem. 2018;61(22):9889–907.
Sheng H, Huang Y, **ao Y, Zhu Z, Shen M, Zhou P, Guo Z, Wang J, Wang H, Dai W, et al. ATR inhibitor AZD6738 enhances the antitumor activity of radiotherapy and immune checkpoint inhibitors by potentiating the tumor immune microenvironment in hepatocellular carcinoma. J Immunother Cancer. 2020;8(1):e000340.
Kim S, Smith S, Mortimer P, Loembé A, Cho H, Kim K, Smith C, Willis S, Irurzun-Arana I, Berges A, et al. Phase I Study of Ceralasertib (AZD6738), a Novel DNA Damage Repair Agent, in Combination with Weekly Paclitaxel in Refractory Cancer. Clin Cancer Res. 2021;27(17):4700–9.
Ashworth A. ATR Inhibitors and Paclitaxel in Melanoma. Clin Cancer Res. 2021;27(17):4667–8.
Orvain C, Lin Y, Jean-Louis F, Hocini H, Hersant B, Bennasser Y, Ortonne N, Hotz C, Wolkenstein P, Boniotto M, et al. Hair follicle stem cell replication stress drives IFI16/STING-dependent inflammation in hidradenitis suppurativa. J Clin Invest. 2020;130(7):3777–90.
Kim HH, Abdelmohsen K, Gorospe M. Regulation of HuR by DNA Damage Response Kinases. J Nucleic Acids. 2010;2010:981487.
Xu J, Zhang J, Zhang Z, Gao Z, Qi Y, Qiu W, Pan Z, Guo Q, Li B, Zhao S, et al. Hypoxic glioma-derived exosomes promote M2-like macrophage polarization by enhancing autophagy induction. Cell Death Dis. 2021;12(4):373.
Almine J, O’Hare C, Dunphy G, Haga I, Naik R, Atrih A, Connolly D, Taylor J, Kelsall I, Bowie A, et al. IFI16 and cGAS cooperate in the activation of STING during DNA sensing in human keratinocytes. Nat Commun. 2017;8:14392.
Li S, Bao X, Wang D, You L, Li X, Yang H, Bian J, Wang Y, Yang Y. APOBEC3B and IL-6 form a positive feedback loop in hepatocellular carcinoma cells. Sci China Life Sci. 2017;60(6):617–26.
Lecona E, Fernandez-Capetillo O. Targeting ATR in cancer. Nat Rev Cancer. 2018;18(9):586–95.
Acknowledgements
We are grateful to Dr. Frederick F. Lang and Dr. Krishna P.L. Bhat (The University of Texas MD Anderson Cancer Center, Houston, TX) for providing GSC cell lines used in our study.
Funding
This work was supported by grants from the National Natural Science Foundation of China (Nos. 82273195; 81874083; 82072776; 82273286; 82072775; 82203419;), Natural Science Foundation of Shandong Province of China (Nos. ZR2019BH057; ZR2020QH174; ZR2021LSW025), the **an Science and Technology Bureau of Shandong Province (2021GXRC029), Key Clinical Research Project of Clinical Research Center of Shandong University (2020SDUCRCA011) and Taishan Pandeng Scholar Program of Shandong Province (No. tspd20210322).
Author information
Authors and Affiliations
Contributions
GL and XG supervised the project. ZJG, JYX and YF designed the research and performed all experiment. ZJG, JYX and YF completed the basic experiment part. ZJG, JYX, YF and HZW performed statistical analysis. ZPZ, MYQ, PZ, LD, JS and RRZ helped to revise the manuscript. GL, XG and HX helped advise on this research design. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The study was authorized by the Ethics Committee of Qilu Hospital. The animal experiments in this study were approved by the Ethics Committee of Qilu Hospital (No. DWLL-2021–082).
Consent for publication
All authors give consent for the publication of the manuscript in Journal of Experimental & Clinical Cancer Research.
Competing interests
The authors declare no potential conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Figure S1.
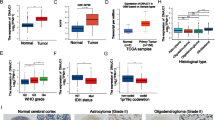
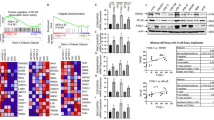
(A) The expression of ARPs among mesenchymal (MES), proneural (PN), and classical (CL) phenotypes in TCGA GBM. (B) Kaplan–Meier curves visualizing the overall survival of CCGA-GBM patients stratified according to expression of ARPs. (C) The expression of ARPs among MES, PN, and CL phenotypes in CGGA GBM. (D) GSEA exhibited a positive correlation between ARPC1B expression and MES phenotypes, and a negative correlation with PN phenotypes. (E) Single-cell RNA sequencing of GSE138794 visualizing the expression of ARPs other than ARPC1B. Fig. S2. (A) Correlation analysis of ARPC1B with CD44, YKL-40, OLIG2 and SOX2 in TCGA-GBM and CGGA-GBM, respectively. (B) Western blot analysis of ARPC1B and SOX2 protein levels in GSC 8-11 overexpressing ARCP1B. (C, D) Representative images and quantification of tumor sphere formation of GSC 8-11 (C) and GSC 11 (D) transduced with vector or ARPC1B. Scale bar, 100μm. (E) The protein expression of ARPC1B in GSC 20 and GSC 267 under different IR dose treatments. Fig. S3. (A) Flow cytometric analysis showing the effect of ARPC1B overexpression on the apoptosis in IR-treated (6Gy) GSC 8-11 cells. The right panels showing the quantification of apoptosis rate. (B) Representative images and quantification of comet assay showing the effect of ARPC1B overexpression on DNA damage of GSC 8-11 with IR treatment (6 Gy). Scale bar, 20μm. (C) Representative images and quantification of γ-H2AX IF staining showing the effect of ARPC1B knockdown on DNA damage of GSC 267 and GSC 20 with IR treatment (6 Gy). Scale bar, 40μm. (D) Representative images and quantification of γ-H2AX IF staining in GSC 8-11. Scale bar, 40μm. (E) Cell-cycle analysis of GSC 267, GSC 20 and GSC 8-11 in different treatment groups. The proportions of cells arrested in G2/M phase were quantified (right panel). Fig. S4. (A, B) Bioluminescence imaging of tumor size on day 7 in sh-control, sh-ARPC1B#1 and sh-ARPC1B#2 GSC 267 (A) or GSC 20 (B) xenograft nude mice in indicated groups. The right panel shows the quantification of photon counts of GSC 267 and GSC 20 xenografts. (C, D) Bioluminescence imaging of tumor size on day 7 (C) and day 30 (D) in Vector or ARPC1B-transfected GSC 8-11 xenograft nude mice receiving or exempt from IR treatment. The right panel showing the quantification of photon counts of GSC 8-11 xenografts. Fig. S5. (A) Representative images and quantification of IHC staining for CD44 in sections of non-IR GSC 267 xenografts (upper, scale bar, 200μm), and TUNEL staining in sections of IR treated GSC 267 xenografts (lower, scale bar, 200μm). (B) Representative images and quantification of IHC staining for CD44 in sections of non-IR GSC 8-11 xenografts (upper, scale bar, 200μm), and TUNEL staining in sections of IR treated GSC 8-11 xenografts (lower, scale bar, 200μm). (C) Representative images of H&E staining in sections from indicated xenografts. Scale bar, 400μm. (D) Western blotting analysis of protein levels of DBN1, ACTN4, FLNA, and CORO1C upon knockdown of ARPC1B in GSC 20 and GSC 267, or overexpression of ARPC1B in GSC 8-11. Fig. S6. (A) Co-IF staining exhibiting the distribution of ARPC1B with IFI16 or HuR in GSC 20. Scale bar, 5μm. (B) The protein levels of IFI16 and HuR after overexpression of ARPC1B in GSC 8-11. (C) The mRNA expression of IFI16 and HuR assessed by qRT-PCR assay. (D) The protein levels of IFI16 and HuR in sh-control or sh-ARPC1B-GSCs treated with 100μg/ml CHX for indicated times. (E) Representative images and quantification of IHC staining for ARPC1B, HuR and IFI16 in different groups of GSC 267 xenograft sections (scale bar, 200μm). Fig. S7. (A) Western blotting analysis showing the effect of TRIM21 knockdown on the protein levels of IFI16 and HuR in sh-ARPC1B GSCs. (B) Western blotting analysis showing the effect of ARPC1B knockdown in GSC 20 and GSC 267, or ARPC1B overexpression in GSC 8-11 on the protein levels of STAT3, p-STAT3, P65, and p-P65. (C) Western blotting analysis of protein levels of ARPC1B, IFI16, HuR, P65, p-P65, STAT3, p-STAT3 and, CD44 in GSC 20 treated with indicated interventions. (D) Western blotting analysis showing that knockdown of TRIM21 could reverse the effect of ARPC1B inhibition on IFI16 and HuR ubiquitination. GSCs were pretreated with MG132 (10 µM) for 6 hours before cell lysates were collected. Fig. S8. (A) Representative images of tumor sphere formation of GSC 267 and GSC 20 treated with indicated interventions. Scale bar, 100μm. (B) Representative images of flow cytometry assays showing apoptosis of GSC 267 and GSC 20 treated with indicated interventions. (C) Representative images of comet assays showing DNA damage of GSC 267 and GSC 20 treated with indicated interventions. Scale bar, 20μm. Fig. S9. (A) The correlation between the ARPC1B expression and drug sensitivity assessed by Spearman algorithm. (B) CCK-8 assay in GSC 267 and GSC 20 treated with different concentrations of AZD6738 for 48 h. (C) Representative images and quantification of apoptosis assays (upper panel) and comet assays (lower panel, scale bar, 20μm) for GSC 20 and GSC 267 upon treatment with the indicated interventions. (D) The representative images and quantification of apoptosis assays (upper panel) and comet assays (lower panel, scale bar, 20μm) for GSC 8-11 upon treatment with the indicated interventions. The right panels are the quantification of apoptosis rate and DNA damage, respectively. (E) Representative images and quantification of TUNEL staining in sections of GSC 267 xenografts for different groups. Scale bar, 200μm. (F) Bioluminescence imaging of tumor size on day 7 in GSC 267 xenograft nude mice treated with the indicated interventions. (G) The quantification of photon counts on day 7 of the GSC 267 xenografts. Supplementary Materials and Methods.
Additional file 2: Table S1.
Correlation analysis of ARPC1B with PN and MES marker genes (Correlation). Table S2. Mass spectrometry result of proteins related with ARPC1B. Table S3. The correlation between the ARPC1B expression and drug sensitivity assessed by Spearman algorithm (correlation).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Gao, Z., Xu, J., Fan, Y. et al. ARPC1B promotes mesenchymal phenotype maintenance and radiotherapy resistance by blocking TRIM21-mediated degradation of IFI16 and HuR in glioma stem cells. J Exp Clin Cancer Res 41, 323 (2022). https://doi.org/10.1186/s13046-022-02526-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13046-022-02526-8