Abstract
Background
Infections by viruses including severe acute respiratory syndrome coronavirus 2 could cause organ inflammations such as myocarditis, pneumonia and encephalitis. Innate immunity to viral nucleic acids mediates antiviral immunity as well as inflammatory organ injury. However, the innate immune mechanisms that control viral induced organ inflammations are unclear.
Methods
To understand the role of the E3 ligase TRIM18 in controlling viral myocarditis and organ inflammation, wild-type and Trim18 knockout mice were infected with coxsackievirus B3 for inducing viral myocarditis, influenza A virus PR8 strain and human adenovirus for inducing viral pneumonia, and herpes simplex virus type I for inducing herpes simplex encephalitis. Mice survivals were monitored, and heart, lung and brain were harvested for histology and immunohistochemistry analysis. Real-time PCR, co-immunoprecipitation, immunoblot, enzyme-linked immunosorbent assay, luciferase assay, flow cytometry, over-expression and knockdown techniques were used to understand the molecular mechanisms of TRIM18 in regulating type I interferon (IFN) production after virus infection in this study.
Results
We find that knockdown or deletion of TRIM18 in human or mouse macrophages enhances production of type I IFN in response to double strand (ds) RNA and dsDNA or RNA and DNA virus infection. Importantly, deletion of TRIM18 protects mice from viral myocarditis, viral pneumonia, and herpes simplex encephalitis due to enhanced type I IFN production in vivo. Mechanistically, we show that TRIM18 recruits protein phosphatase 1A (PPM1A) to dephosphorylate TANK binding kinase 1 (TBK1), which inactivates TBK1 to block TBK1 from interacting with its upstream adaptors, mitochondrial antiviral signaling (MAVS) and stimulator of interferon genes (STING), thereby dampening antiviral signaling during viral infections. Moreover, TRIM18 stabilizes PPM1A by inducing K63-linked ubiquitination of PPM1A.
Conclusions
Our results indicate that TRIM18 serves as a negative regulator of viral myocarditis, lung inflammation and brain damage by downregulating innate immune activation induced by both RNA and DNA viruses. Our data reveal that TRIM18 is a critical regulator of innate immunity in viral induced diseases, thereby identifying a potential therapeutic target for treatment.
Similar content being viewed by others
Background
Innate immunity is the first line of defense against invading pathogens including RNA and DNA viruses. Activation of innate immunity requires the recognition of pathogen-associated molecular patterns (PAMPs), such as atypical viral nucleic acids, by pattern-recognition receptors (PRRs) on innate immune cells [1]. Recognition of PAMPs by PRRs activates signaling cascades leading to the production of type I interferon (IFN), which is central to host anti-viral defense by upregulating IFN-stimulated genes (ISGs) that limit virus dissemination and activate adaptive immune responses [2, 3]. Multiple PRRs have been identified that recognize viral RNA and DNA and induce type I IFN, including membrane-bound sensors such as Toll-like receptors (TLRs) [1, 4], cytosolic RNA sensors such as retinoic acid-inducible gene-I (RIG-I)-like receptors (RLRs) [5] and poly(ADP-ribose) polymerase 9 (PARP9) [2: Fig. S3a). We first analyzed TRIM18 gene expression in different mouse immune cells using the Immunological Genome Project (ImmGen) and found TRIM18 was indeed highly expressed in mouse macrophages including peritoneal macrophages, splenic macrophages, alveolar macrophages and microglia macrophages (Additional file 2: Fig. S3b). Next, we isolated mouse peritoneal macrophages (MF PC) and splenic macrophages (MF Sp) and detected high expression of TRIM18 in macrophages from wild-type (WT) mice, while deletion of TRIM18 expression was confirmed by immunoblot analysis (Additional file 2: Fig. S3c). We also investigated if TRIM18 expression was affected by RNA virus or DNA virus infection in mouse bone marrow-derived macrophages (BMDM). We found TRIM18 was induced at both RNA (Additional file 2: Fig. S3d) and protein (Additional file 2: Fig. S3e) levels in mouse BMDM after RNA or DNA virus infection and the induction of TRIM18 was much stronger in mouse BMDM by DNA viruses HSV-1 and adenovirus than that by RNA viruses Flu PR8 and CVB3 (Additional file 2: Fig. S3d, e). Furthermore, TRIM18 had high expression in lung, brain and heart, and low expression in intestine, liver and kidney from WT mice after DNA virus HSV-1 infection (Additional file 2: Fig. S3f). Additionally, KO of TRIM18 did not change expression of surface markers CD11b and F4/80 by flow cytometry (Additional file 2: Fig. S4), indicating that TRIM18 does not affect differentiation markers of mouse macrophages.
To further investigate the role of TRIM18 in response to RNA viruses, we prepared mouse BMDM from WT and TRIM18 KO mice, and stimulated BMDM with dsRNA poly I:C and 5′-triphosphate RNA (5’pppRNA), and measured type I IFN proteins (IFN-α and IFN-β) by ELISA as well as mRNA levels of interferon stimulated gene 15 (ISG15) and ISG56 by qRT-PCR. The results showed that TRIM18 KO BMDM produced much more IFN-α and IFN-β proteins (Fig. 2a, b) and mRNAs of ISG15 and ISG56 (Additional file 2: Fig. S5a, b) than WT BMDM in response to 5’pppRNA and poly I:C stimulation. In addition, we employed two RNA viruses including Flu PR8 and CVB3 to investigate TRIM18 in response to RNA viruses in mouse BMDM. BMDM from WT and TRIM18 KO mice were isolated and infected with RNA viruses Flu PR8 and CVB3. Compared with WT BMDM, TRIM18 KO BMDM produced 2- to threefold more IFN-α and IFN-β proteins (Fig. 2c, d) and twofold more mRNAs of ISG15 and ISG56 (Additional file 2: Fig. S5c, d) post-infection by RNA viruses Flu PR8 and CVB3. Collectively, these data demonstrate a negative role for TRIM18 in regulating production of type I IFN and ISGs in mouse macrophages in response to dsRNA and RNA viruses.
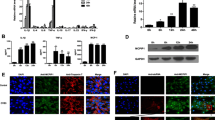
TRIM18 negatively regulates type I IFN production by BMDM upon stimulations of dsRNA and dsDNA or infections with RNA and DNA viruses. a–d ELISA of IFN-α a, c and IFN-β b, d production by BMDM from Trim18+/+ (WT) and Trim18−/− (KO) mice after 12 h of stimulations with 5’pppRNA (0.5 μg/ml), poly I:C (0.5 μg/ml) delivered by Lipofectamine 3000 a, b or infections with RNA viruses including influenza A virus (influenza A virus PR8 strain, Flu PR8) and Coxsackievirus B3 (CVB3) c, d at an MOI of 2 (n = 3 per group). e–h, ELISA of IFN-α e, g) and IFN-β f, h production by BMDM from Trim18+/+ (WT) and Trim18−/− (KO) mice after 12 h of stimulations with dsDNA from HSV-1 virus (HSV60, 2.5 μg/ml) and cGAMP (2.5 μg/ml) delivered by Lipofectamine 3000 e, f or infections with DNA viruses including adenovirus and HSV-1 g, h at an MOI of 2 (n = 3 per group). Each circle represents an individual independent experiment and small solid black lines indicate the average of triplicates for results. **p < 0.01, ***p < 0.001, and p value was calculated by unpaired two-tailed Student’s t test. Mock, wild-type BMDM without stimulation or infection. Data are representative of three independent experiments
To further determine the role of TRIM18 in response to DNA viruses, we prepared mouse BMDM from WT and TRIM18 KO mice, and stimulated BMDM with dsDNA HSV60 and cGAMP (STING stimulator in DNA sensing pathway), and measured type I IFN proteins by ELISA and mRNA levels of ISG15 and ISG56 by RT-qPCR. Consistent with the earlier results, TRIM18 KO BMDM produced much more IFN-α and IFN-β proteins (Fig. 2e, f) and mRNAs of ISG15 and ISG56 (Additional file 2: Fig. S5e, f) than WT BMDM in response to HSV60 and cGAMP stimulation. Furthermore, two DNA viruses including HSV-1 and human adenovirus were chosen to investigate role of TRIM18 in response to DNA viruses in mouse BMDM. BMDM from WT and TRIM18 KO mice were prepared and infected with DNA viruses adenovirus and HSV-1. The results showed TRIM18 KO BMDM produced significantly more IFN-α and IFN-β proteins (Fig. 2g, h) and mRNAs of ISG15 and ISG56 (Additional file 2: Fig. S5g, h) than WT BMDM after infection with DNA viruses. Taken together, these data suggest that TRIM18 is a negative regulator of type I IFN and ISGs productions in mouse macrophages in response to dsDNA and DNA viruses.
Deletion of TRIM18 protects mice from viral myocarditis
Viral myocarditis has been recognized as a cause of congestive heart failure and CVB3 infection is the main cause of viral myocarditis [20, 21]. Since TRIM18 is highly expressed in heart and TRIM18 negatively regulates innate immune response to RNA virus CVB3 in macrophages, we investigated the functional importance of TRIM18 in controlling CVB3 induced myocarditis in vivo. We first intraperitoneally infected both WT and TRIM18 KO mice with the RNA virus CVB3 and checked cardiac histology and functions. The heart histopathology revealed TRIM18 KO mice had significantly reduced cardiac inflammation and infiltration of inflammatory cells compared with WT mice following CVB3 infection (Fig. 3a, b). Additionally, TRIM18 expression was induced in hearts of WT mice with CVB3 infection (Fig. 3c), while TRIM18 induction was stronger in hearts of mice during CVB3 acute infection than that during CVB3 chronic infection (Fig. 3c). It’s reported that another TRIM family member TRIM21 could restrict CVB3 induced cardiac injury by positively regulate IRF3-mediated type I IFN production [68]. We then compared the expressions of TRIM18 and TRIM21 in hearts from WT and TRIM18 KO mice after CVB3 infection. The immunohistochemistry (IHC) data showed that there was high expression of TRIM21 in hearts from WT and TRIM18 KO mice after CVB3 infection (Fig. 3d). However, TRIM18 had higher expression than TRIM21 in heart from CVB3 infected WT mice, while TRIM18 expression was gone in heart from TRIM18 KO mice after CVB3 infection (Fig. 3d). Furthermore, IHC data showed that there were major macrophages and neutrophils, and minor NK cells and T cells in the infiltrated cells in hearts from mice after CVB3 infection for two days (Fig. 3e). In agreement, the echocardiography of WT mice revealed impaired cardiac function (Fig. 3f) as evidenced by decreased ejection fraction (EF) (Fig. 3g) and fractional shortening (FS) (Fig. 3h) when compared with TRIM18 KO mice. Compared to WT mice, TRIM18 KO mice had less heart weight increase during viral myocarditis (Fig. 3i), a marker of cardiac inflammatory edema. Additionally, brain natriuretic peptide (BNP), a marker of heart failure, was dramatically reduced in TRIM18 KO hearts compared with WT hearts (Fig. 3j). Importantly, we found that most of WT mice succumbed to CVB3 infection, while the survival of TRIM18 KO mice was significantly better than that of their WT littermates (Fig. 3k). These data suggested that deficiency of TRIM18 could protect mice from CVB3 induced myocarditis by reducing cardiac inflammation with improved function. To further investigate the mechanisms by which TRIM18 knockout mice reduced CVB3 induced myocarditis, we next checked viral replication and type I IFN protein levels in heart homogenates by plaque-forming assay and ELISA, respectively. We found that the CVB3 viral loads were significantly reduced in hearts from TRIM18 KO mice compared with WT mice at day 2 (D2), day 5 (D5) and day 14 (D14) after CVB3 infection (Fig. 3l). Furthermore, TRIM18 KO mice had higher concentrations of IFN-α (Fig. 3m) and IFN-β (Fig. 3n) in the hearts than did their WT littermates after infection with CVB3. However, WT mice had higher cardiac inflammatory cytokines IL-6 (Fig. 3o), TNF-α (Fig. 3p) and IL-1β (Fig. 3q) than did their TRIM18 KO littermates after CVB3 infection. These data indicate that deletion of TRIM18 protects mice from CVB3 induced myocarditis by improving cardiac function and promoting innate immune activation.
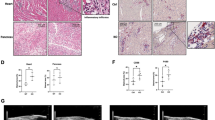
Deletion of TRIM18 protects mice from myocarditis induced by RNA virus CVB3 in vivo. a Hematoxylin and eosin (H&E)-staining of heart sections from age-matched Trim18+/+ (WT) and Trim18−/− (KO) male mice after intraperitoneal infection with CVB3 (1 × 107 PFU per mouse) for 4 days. Scale bars represent 1000 μm for original images and 200 µm for enlarged images. b Histology score analysis of viral myocarditis in heart sections from mice as in (a). c Immunoblot (IB) analysis of TRIM18 expression in hearts from WT mice without or with intraperitoneal CVB3 acute infection (1 × 107 PFU per mouse) or chronic infection (1 × 103 PFU per mouse) for 2 days. The position of protein markers (shown in kDa) is indicated at right. d Immunohistochemistry (IHC) analysis of TRIM18 and TRIM21 expression in hearts from WT and KO male mice after CVB3 infection. e IHC analysis of the infiltrated cells in CVB3 infected hearts using anti-macrophage marker antibody, anti-neutrophil marker antibody, anti-NK cell marker antibody and anti-T cell marker antibody, respectively. Scale bars represent 100 μm in (d, e). f Representative M-mode images of hearts from WT and KO male mice at day 4 after CVB3 infection by echocardiography analysis. g–h Cardiac function analysis of ejection fraction (EF) g and fractional shortening (FS) h of hearts from mice as in (f) (n = 5 per group). i The assessment of heart weight/baseline body weight from WT and KO male mice (n = 5 per group) at day 0 or day 6 after CVB3 infection. j The qRT-PCR analysis of brain natriuretic peptide (BNP) mRNA in the heart of from WT and KO male mice (n = 5 per group) at day 1, day 2 or day 4 after CVB3 infection.; results are presented relative to those of mock mice. k Survival of age-matched WT and KO male mice after intraperitoneal infection with CVB3 (1 × 107 PFU per mouse) (n = 10 per group). l Viral titers in homogenates of hearts from WT and KO male mice at day 2 (D2), day 5 (D5) and day 14 (D14) after CVB3 infection (n = 5 per group for D2 and D5, n = 3 per group for D14). m–q, ELISA of IFN-α (m), IFN-β (n) IL-6 (o), TNF-α (p) and IL-1β q in hearts from mice as in k (n = 5 per group). Error bars indicate standard error of the mean for results in (b, g–j, l–q). NS, not significant (p > 0.05), **p < 0.01, ***p < 0.001, and p value was calculated by unpaired two-tailed Student’s t test and Gehan-Breslow-Wilcoxon test for survival analysis. Data are representative of three independent experiments
Knockout of TRIM18 protects mice from pneumonia and lung injury induced by viral infections
Viral pneumonia is an inflammation of the lungs caused by respiratory viruses, such as influenza virus, adenovirus and SARS-CoV-2 causing the ongoing pandemic of COVID-19 [23, 24]. Interestingly, the public GEO profile database show that patients with SARS-CoV-2 infection have higher expression of TRIM18 (Additional file 2: Fig. S6), we hypothesize that TRIM18 may play crucial roles in controlling pneumonia and lung injury induced by respiratory viruses including SARS-CoV-2. As shown above, TRIM18 was highly expressed in lung and TRIM18 downregulated the innate immune responses to respiratory viruses including RNA virus influenza virus and DNA virus adenovirus in both human and mouse macrophages. Therefore, we investigated if TRIM18 could control viral pneumonia induced by those respiratory viruses including influenza virus and adenovirus in vivo. First, we infected both WT and TRIM18 KO mice intranasally with respiratory RNA virus influenza virus and checked lung inflammation and injury by histology. Indeed, lung histopathology revealed edema, alveolar hemorrhaging, alveolar wall thickness and neutrophil infiltration in lungs from TRIM18 KO mice were less marked than those from WT after influenza virus infection (Fig. 4a, b). Importantly, we found that TRIM18 KO mice were more resistant to influenza virus infection than their WT littermates (Fig. 4c). These results suggested that knockout of TRIM18 protected mice from lung injury and inflammation induced by RNA virus influenza virus in vivo. We further investigated the mechanisms by which deletion of TRIM18 protected mice from pneumonia infected by influenza virus. We determined viral amplification in lungs at day 2 post-infection by plaque-forming assay. We detected significantly less influenza virus loads in TRIM18 KO mice than in their WT littermates (Fig. 4d). We then detected type I IFN production in lungs by ELISA. Compared with WT mice, TRIM18 KO mice produced significantly more IFN-α (Fig. 4e) and IFN-β (Fig. 4f) following influenza virus infection. Additionally, there were increased infiltration cells mainly containing macrophages, neutrophils, and lymphocytes in bronchoalveolar lavage fluid (BALF) of WT mice with Flu PR8 infection, which were dramatically reduced in TRIM18 KO mice (Fig. 4g). These data indicate that TRIM18 deficiency protects mice from pneumonia induced by RNA virus influenza virus through restricting viral replication and promoting innate immune activation in vivo.
Knockout of TRIM18 protects mice from pneumonia and lung injury induced by RNA virus Flu PR8 or DNA virus adenovirus in vivo. a Hematoxylin and eosin (H&E)-staining of lung sections from age- and sex-matched Trim18+/+ (WT) and Trim18−/− (KO) mice left infected (Mock) or infected for 4 days by intranasal infection of Flu PR8 virus (1 × 105 PFU per mouse). b Histology score analysis of viral pneumonia in lung sections from mice as in (a). c Survival of age- and sex-matched WT and KO mice after intranasal infection with Flu PR8 virus (1 × 105 PFU per mouse) (n = 10 per group). d Plaque assay of Flu PR8 virus titers in the lung of WT and KO mice infected for 2 days by intranasal infection of Flu PR8 virus (n = 5 per group). e–f, ELISA of and IFN-α e and IFN-β f in BALF samples from mice (n = 5 per group) as in (d). g Quantification of cell numbers in BALF samples from mice (n = 3 per group) as in (d). h Hematoxylin and eosin (H&E)-staining of lung sections from WT and KO mice left infected (Mock) or infected for 4 days by intranasal infection of adenovirus (1 × 108 PFU per mouse). i, Histology score analysis of viral pneumonia in lung sections from mice as in (h). j Viral titers in homogenates of lung from WT and KO mice (n = 5 per group) at day 2 of intranasal infection with adenovirus (1 × 108 PFU per mouse). k, l ELISA of IFN-α k and IFN-β l in BALF samples from mice as in j (n = 5 per group). m Quantification of cell numbers in BALF samples from mice (n = 3 per group) as in (j). Scale bars represent 400 μm for images in a and (h). Error bars indicate standard error of the mean for results in (b, d–g, i–m). NS, not significant (p > 0.05), **P < 0.01 and ***P < 0.001, and p value was calculated by unpaired two-tailed Student’s t test and Gehan-Breslow-Wilcoxon test for survival analysis. Data are representative of three independent experiments
Next, we evaluated the importance of TRIM18 in controlling pneumonia following infection by respiratory DNA virus adenovirus in vivo. Both WT and TRIM18 KO mice were infected intranasally with adenovirus, which is normally transmitted by the nasal route and targets the lungs for pneumonia. Lung histopathology revealed much-reduced edema, alveolar hemorrhage, alveolar wall thickness, and neutrophil infiltrations in TRIM18 KO mice compared to the lung pathology in WT mice (Fig. 4h, i), indicating the importance of TRIM18 in promoting adenovirus induced pneumonia and lung inflammation. To further investigate the underlying mechanisms, we measured adenovirus replication and type I IFN production in lung by plaque-forming assay and ELISA, respectively. We found that the adenovirus loads were dramatically reduced in TRIM18 KO mice compared with WT mice (Fig. 4j). Additionally, there was significantly more IFN-α (Fig. 4k) and IFN-β (Fig. 4l) in the BALF from TRIM18 KO mice than that from WT mice at day 2 post infection. Compared with TRIM18 KO mice, there were much more infiltration cells mainly consisted of macrophages, neutrophils, and lymphocytes in BALF of WT mice with adenovirus infection (Fig. 4m). Collectively, these findings demonstrate that knockout of TRIM18 protects mice from pneumonia and lung injury induced by viral infections through enhancing activation of innate immunity in vivo.
Deficiency of TRIM18 protects mice from encephalitis induced by HSV-1 infection
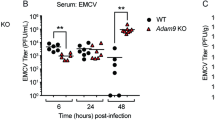
Herpes simplex encephalitis (HSE) is caused by HSV-1 infection of the brain and is the most common cause of sporadic fatal encephalitis worldwide[22]. Given that the high expression of TRIM18 in brain and the critical role of TRIM18 in regulating innate immune response to HSV-1 in macrophages, we further investigated if TRIM18 plays role in controlling brain damage and inflammation induced by DNA virus HSV-1 in vivo. We challenged WT and TRIM18 KO mice intravenously with HSV-1 virus and checked brain damage and inflammation by histology. The brain histopathology revealed much-reduced demyelination, necrosis, and inflammatory cell infiltration in TRIM18 KO mice as compared to the brain pathology in WT mice after HSV-1 infection (Fig. 5a, b). Compared with WT mice, TRIM18 KO mice had significantly higher survival rates (Fig. 5c). These results indicated that knockout of TRIM18 protected mice from brain damage and inflammation induced by HSV-1 virus in vivo. To further investigate the underlying mechanisms, we measured HSV-1 replication in brain and type I IFN production in serum by plaque-forming assay and ELISA, respectively. We detected significantly less HSV-1 virus in brain of TRIM18 KO mice than in WT mice on day 2 after infection (Fig. 5d). Moreover, TRIM18 KO mice had higher concentrations of IFN-α (Fig. 5e) and IFN-β (Fig. 5f) in the serum than did their WT littermates after HSV-1 infection. Collectively, these data demonstrate deficiency of TRIM18 protects mice from encephalitis induced by HSV-1 by enhancing innate immune activation.
Deficiency of TRIM18 protects mice from brain damage induced by DNA virus HSV-1 in vivo. a Hematoxylin and eosin (H&E)-staining of brain sections from Trim18+/+ (WT) and Trim18−/− (KO) mice left infected (Mock) or infected for 4 days by intravenous infection of HSV-1 (1 × 107 PFU per mouse). Scale bars represent 1000 μm for original images and 200 µm for enlarged images. b Histology score analysis of viral encephalitis in brain sections from mice as in (a). c Survival of WT and KO mice (n = 10 per group) after intravenous injection of HSV-1 (1 × 107 PFU per mouse). d Viral titers in homogenates of brains from WT and KO mice (n = 5 per group) after intravenous injection of HSV-1 (1 × 107 PFU per mouse). e, f, ELISA of IFN-α e and IFN-β f in serum obtained from WT and KO mice (n = 5 per group) at 12 h after intravenous injection of HSV-1. Error bars indicate standard error of the mean for results in (b, d–f). NS, not significant (p > 0.05), **P < 0.01 and ***P < 0.001, and p value was calculated by unpaired two-tailed Student’s t test and Gehan-Breslow-Wilcoxon test for survival analysis. Data are representative of three independent experiments.
TRIM18 recruits PPM1A to inactivate TBK1 blocking TBK1 from interactions with its upstream adaptors during virus infection
To determine the molecular mechanisms by which the enhanced production of type I IFN and innate immune activation were achieved in BMDM from TRIM18 KO mice, we isolated BMDM from WT and TRIM18 KO mice and infected the cells without or with CVB3 and adenovirus for 1 h, then assessed activation of the transcription factors IRF3 by immunoblot analysis. We found that phosphorylation of IRF3 in TRIM18 KO BMDM was enhanced relative to WT BMDM after infection with CVB3 or adenovirus (Fig. 6a). Typically, cytosolic RNA or DNA is sensed by RIG-I-like receptors or DNA sensor such as cGAS, which then activate downstream MAVS and STING, respectively. MAVS or STING recruits downstream TBK1 to phosphorylate and activate IRF3 for inducing type I IFN. To further dissect the role of TRIM18 in these different IFN-I induction pathways, we examined the effect of TRIM18 on the IFN-β luciferase reporter activated by these components including MDA5, MAVS, TBK1, IKKi and cGAS/STING. Overexpression of TRIM18 reduced the IFN-β luciferase reporter activation by MDA5 and MAVS and to a greater extent by TBK1 and cGAS/STING (Additional file 2: Fig. S7a–d). However, TRIM18 did not inhibit downstream IKKi dependent IFN-β luciferase reporter activation (Additional file 2: Fig. S7e), indicating that TRIM18 targets the pathway at nodes between TBK1 and IKKi. To further investigate how TRIM18 regulates such signaling molecules, we used immunoprecipitation with an antibody specific to TRIM18 to identify TRIM18-interacting proteins in lysates of BMDM, followed by protein sequencing by liquid chromatography–mass spectrometry. Among a group of TRIM18-interacting proteins, we identified protein phosphatase, magnesium-dependent 1A (PPM1A; formerly called PP2C) (Additional file 1: Table S1). PPM1A has previously been reported to silence cytosolic RNA sensing and antiviral defense through direct dephosphorylation of TBK1 [6e) showed that the C-terminal SPRY (Sp1A kinase and Ryanodine receptors) domain of TRIM18 bound to PPM1A (Fig. 6f). To further investigate whether TRIM18 recruits PPM1a to target and dephosphorylate TBK1, we isolated BMDM from WT and TRIM18 KO mice and infected the cells without or with CVB3 and adenovirus for 1 h, then assessed phosphorylation of TBK1 by immunoblot analysis. We found that phosphorylation of TBK1 in TRIM18 KO BMDM was significantly enhanced relative to that in WT BMDM after infection with CVB3 or adenovirus (Fig. 6g), suggesting that TRIM18 indeed recruited PPM1A to dephosphorylate TBK1 for TBK1 inactivation. Additionally, we found there was more PPM1A expression in WT BMDM than TRIM18 KO BMDM without and with CVB3 or adenovirus infection (Fig. 6b, g). To further investigate the outcomes of the recruitment and stability of PPM1A by TRIM18, we isolated BMDM from WT and TRIM18 KO mice and infected the cells without or with CVB3 and adenovirus for 5 h, then evaluated the interaction between TBK1 and its upstream adaptors MAVS or STING by co-immunoprecipitation and immunoblot analysis. We found that there was no interaction between TBK1 and MAVS or STING in BMDM from either WT or TRIM18 KO mice that had not been infected (Fig. 6h). However, after infection with CVB3 or adenovirus, interaction between TBK1 and MAVS or STING in both WT and TRIM18 KO BMDM was evident, but significantly enhanced in the TRIM18 KO BMDM relative to levels in WT BMDM (Fig. 6h), indicating that the recruitment of PPM1A by TRIM18 blocked interaction between TBK1 and its upstream adaptors MAVS or STING in macrophages after virus infection. Collectively, these data suggest that TRIM18 recruits PPM1A to dephosphorylate TBK1 for its inactivation and blocks interaction of TBK1 with its upstream adaptors MAVS and STING for signal transduction during virus infection.
TRIM18 stabilizes PPM1A by mediating K63-linked ubiquitination
Because TRIM18 is an E3 ubiquitin ligase and less PPM1A is seen in TRIM18 KO cells, we surmised that TRIM18 may mediate the stability of PPM1A. Consequently, we next determined whether TRIM18 was responsible for the ubiquitination of PPM1A ex vivo. We isolated BMDM from WT and TRIM18 KO mice and infected those cells with CVB3 or adenovirus for 6 h. Cell lysates were then prepared and analyzed for the ubiquitination of PPM1A. As results, PPM1A was modified via K63-mediated ubiquitination in BMDM from WT mice but not TRIM18 KO mice (Fig. 7a). Additionally, there was more PPM1A expression in WT BMDM that TRIM18 KO BMDM (Fig. 7a), which demonstrated the crucial role of K63-mediated ubiquitination in mediating stability of PPM1A. To investigate whether the ubiquitination of PPM1A was dependent on the binding site of TRIM18 with PPM1A, we transfected the HEK293T cells to co-express Myc-tagged PPM1A and HA-tagged full-length TRIM18, or truncated TRIM18 lacking the binding site of PPM1A (T18-∆SPRY). We then prepared cell lysates and incubated them for 5 min at 90 °C with 1% SDS (sodium dodecyl sulfate) to disrupt protein–protein interactions, followed by immunoprecipitation of Myc-tagged PPM1A. Immunoblot analysis of HA or ubiquitin demonstrated that the ubiquitination of PPM1A was strongly enhanced by overexpression of TRIM18 but not by overexpression of T18-∆SPRY (Fig. 7b). Immunoblot analysis of K63-linked ubiquitin further demonstrated that TRIM18 induced ubiquitination of PPM1A by K63-mediated linkage (Fig. 7b). Together, these data indicate that TRIM18 targets PPM1A and induces its ubiquitination for protein stability by K63-linkage.
TRIM18 induces ubiquitination of PPM1A by K63 linkage. a Immunoblot (IB) analysis of TRIM18 (fourth blot), PPM1A (fifth blot) and β-actin (bottom blot) in WT and TRIM18 KO BMDM, the abundance (top blot), total ubiquitination (second blot), and K63-mediated ubiquitination (third blot) of PPM1A in those cells, infected for 4 h with CVB3 or adenovirus at an MOI of 5, assessed after immunoprecipitation with anti-PPM1A antibody. b Immunoblot analysis (with anti-Myc) of the abundance (top), total ubiquitination (second blot), and K63-linked ubiquitination (third blot) of Myc-tagged PPM1A in HEK293T cells transfected with empty vector or expression vector for full length HA-tagged TRIM18 (HA-T18 full), truncation T18-∆SPRY (losing binding site of PPM1A), assessed after immunoprecipitation with anti-Myc antibody; immunoblot analysis of whole-cell lysates with anti-HA (fourth blot), anti-Myc (fifth blot) and anti-β-actin (bottom). c Immunoblot analysis (with anti-Myc) of the abundance (top), total ubiquitination (second blot), and K63-linked ubiquitination (third blot) of Myc-tagged PPM1A in HEK293T cells transfected with HA-TRIM18 and Myc-PPM1A wild-type (WT) and its mutations including K9RK12R, K118RK119R, K156R, K296R, K303RK304R, and K354R, assessed after immunoprecipitation with anti-Myc antibody; immunoblot analysis of whole-cell lysates with anti-Myc (fourth blot), anti-HA (fifth blot) and anti-β-actin (bottom). d Luciferase assay in HEK293T cells transfected with IFN-β reporter, Flag-tagged plasmid expressing TBK1 along with Myc-tagged empty vector or PPM1A wild-type (WT) and its mutations including K9RK12R, K118RK119R, K156R, K296R, K303RK304R, and K354R. Renilla luciferase RL-TK was used as the internal control. e Immunoblot analysis (with anti-Myc) of the abundance (top), and K63-linked ubiquitination (second blot) of Myc-tagged PPM1A in HEK293T cells transfected with HA-TRIM18, Flag-TBK1 and Myc-PPM1A wild-type (WT) and its ubiquitination losing mutant K9/12/296/354R, assessed after immunoprecipitation with anti-Myc antibody; immunoblot analysis of whole-cell lysates with anti-phosphorylated (p-) TBK1 (third blot), anti-TBK1 (fourth blot) and anti-β-actin (bottom). f Luciferase assay in HEK293T cells transfected with IFN-β reporter, Flag-tagged plasmid expressing TBK1 along with Myc-tagged empty vector or PPM1A wild-type (WT) and its ubiquitination losing mutant K9/12/296/354R. Renilla luciferase RL-TK was used as the internal control. NS, not significant (p > 0.05), ***p < 0.001, and p value was calculated by unpaired two-tailed Student’s t test. The position of protein markers (shown in kDa) is indicated on the right. Data are representative of three independent experiments
PPM1A contains twenty-two lysine residues. Thirteen of these (Lys9, Lys12, Lys98, Lys118, Lys119, Lys156, Lys172, Lys208, Lys296, Lys303, Lys304, Lys310, and Lys354) were predicted to be possible ubiquitination sites with high scores by the BDM-PUB program (Additional file 1: Table S2). To determine the ubiquitination sites in PPM1A, we chose nine predicted ubiquitination sites with scores of more than one (Lys9, Lys12, Lys118, Lys119, Lys156, Lys296, Lys303, Lys304, and Lys354) and replaced each of those lysine residues individually with arginine. We co-expressed HA-tagged TRIM18 with Myc-tagged WT PPM1A and its mutants (K9RK12R, K118RK119R, K156R, K296R, K303RK304R, and K354R) in HEK293T cells and detected their expression and ubiquitination. TRIM18 was expressed similarly and could promote the stability of WT PPM1A and its mutants (K118RK119R, K156R, and K303RK304R), but not PPM1A mutants (K9RK12R, K296R and K354R) (Fig. 7c). In agreement, the total and K63-linked ubiquitination of PPM1A WT or its mutants (K118RK119R, K156R, and K303RK304R) was strongly enhanced by overexpression of TRIM18 (Fig. 7c). However, the total and K63-linked ubiquitination of PPM1A mutants (K9RK12R, K296R and K354R) by TRIM18 was absent (Fig. 7c). Furthermore, we examined the effect of PPM1A WT and its mutants on the IFN-β luciferase reporter activated by TBK1. Overexpression of PPM1A WT or its mutants (K118RK119R, K156R, and K303RK304R) reduced the IFN-β luciferase reporter activation by TBK1 (Fig. 7d). However, PPM1A mutants (K9RK12R, K296R and K354R) did not inhibit TBK1 mediated IFN-β luciferase reporter activation (Fig. 7d), indicating that the four lysine residues of PPM1A (Lys9, Lys12, Lys296, and Lys354) mediated its key effect on TBK1 triggered signaling activation. Next, we constructed PPM1A mutant harboring all those four lysine mutations (PPM1A K9/12/296/354R) to investigate if TRIM18 still induce ubiquitination of PPM1A mutant K9/12/296/354R. Indeed, TRIM18 could induce K63-linked ubiquitination of WT PPM1A, while the K63-mediated ubiquitination of PPM1A mutant K9/12/296/354R were completely blocked (Fig. 7e). The inhibition of PPM1A ubiquitination dramatically reduced its ability to dephosphorylate TBK1 and triggered much stronger phosphorylation and activation of TBK1 (Fig. 7e), thereby boosted TBK1 mediated IFN-β luciferase reporter activation by luciferase assay (Fig. 7f). Collectively, these data showed that TRIM18 targeted PPM1A and induced the K63-linked ubiquitination of PPM1A for maintaining its stability and four lysine residues (Lys9, Lys12, Lys296, and Lys354) were critical sites for TRIM18-mediated ubiquitination and regulation of PPM1A.
Discussion
The elucidation of immune regulatory mechanisms is critical to understanding how the host constrains adventitious inflammation to maintain immune homeostasis. In the present study, we demonstrate an essential role of E3 ubiquitin ligase TRIM18 in serving as negative regulator of antiviral innate immunity against organ inflammations induced by RNA and DNA viruses (Fig. 8). TRIM18 was shown to reduce the type I IFN response by cytosolic dsRNA and dsDNA, which linked TRIM18 to both RNA and DNA sensing pathways. Deficiency of TRIM18 in both human and mouse macrophages potentiated type I IFN induction by RNA and DNA viruses. Functionally, knockout of TRIM18 protected mice from viral myocarditis, viral pneumonia, and herpes simplex encephalitis in vivo. Mechanistically, we demonstrated that TRIM18 recruited the protein phosphatase PPM1A to dephosphorylate TBK1, which inactivated TBK1 and blocked interactions of TBK1 with its upstream adaptors MAVS and STING, dampening type I IFN mediated antiviral signaling during virus infection. Furthermore, TRIM18 promoted K63-linked polyubiquitination of PPM1A for maintaining its stability. Importantly, ablation of TRIM18 led to enhanced antiviral cytokine production and clearance of both RNA and DNA viruses in vivo, underscoring its physiologic function. This work demonstrates that TRIM18 serves as an immunological rheostat to safeguard against inappropriate innate immune responses to cytosolic viral RNA and DNA in human and mouse macrophages. Since we only have TRIM18 global knockout mice but not macrophage specific TRIM18 knockout mice, we could not exclude the potential roles of TRIM18 in cells other than macrophages in antiviral innate immunity against RNA and DNA viruses. Because TRIM18 expression is induced by RNA and DNA virus infection and TRIM18 has high expression in lung, heart and brain of mice with HSV-1 infection, we speculate that the regulatory machinery of TRIM18 in antiviral innate immunity may exist in other cells types such as lung epithelial cells, cardiomyocytes or neurons.
Schematic illustration of TRIM18 serving as a negative regulator in antiviral innate immunity against organ inflammations induced by RNA and DNA viruses. In WT macrophages after infections with DNA and RNA viruses, TRIM18 interacts with PPM1A and induces its K63-linked ubiquitination for maintaining stability of PPM1A, thereby further dephosphorylating TBK1 for its inactivation and blocking the interactions of TBK1 with its upstream adaptors STING and MAVS leading to dramatic reduction of type I IFN, which promotes viral myocarditis and more massive inflammations in lung and brain. In contrast, in TRIM18 KO macrophages post infections with DNA and RNA viruses, PPM1A could not maintain its stability without TRIM18, and loss its strong ability to inactivate TBK1 resulting in significant induction of type I IFN, which restricts viral myocarditis, inflammations in lung and brain
Accumulating evidence suggests that members of the TRIM family play versatile roles in antiviral immunity [57, 58]. A previous systematic analysis of 75 human TRIM proteins indicated that nearly half of TRIM proteins serve as positive regulators of antiviral responses [58]. For example, TRIM4 and TRIM25 promote type I IFN production and NF-κB activity by regulating the ubiquitination of RIG-I in RNA sensing pathway [59, 60], while TRIM23-mediated K27-linked ubiquitination of NEMO promotes TLR3- and RIG-I/MDA5-mediated antiviral and inflammatory responses [61]. Additionally, TRIM32 and TRIM56 positively regulate DNA virus-induced type I IFN signaling by targeting STING for K63-linked ubiquitination [62, 63]. Furthermore, TRIM9 short isoform preferentially promotes DNA and RNA virus-induced production of type I interferon by recruiting GSK3β to TBK1 [64]. On contrast, TRIM38 is shown to negatively regulate TLR3/4-and RIG-I-mediated IFN-β production by promoting K48-linked polyubiquitination and proteasomal degradation of NAP1 [65]. However, these studies lack key in vivo data without using gene knockout mice. In our study, we demonstrated an essential role of TRIM18 in controlling viral myocarditis, viral pneumonia and herpes simplex encephalitis through downregulating innate immune activation against RNA and DNA viruses both in vitro and in vivo. We have previously demonstrated TRIM29 is a negative regulator in antiviral innate immunity against respiratory RNA virus by promoting K48-linked polyubiquitination and proteasomal degradation of NEMO in alveolar macrophages [38] and TRIM29 promotes DNA virus infection by targeting STING for K48-linked polyubiquitination and proteasomal degradation in airway epithelial cells and dendritic cells and targeting TAB2 for degradation in natural killer cells both in vitro and in vivo [39, 66]. Additionally, TRIM31 promotes activation of antiviral innate immunity against RNA viruses by inducing K63-linked polyubiquitination of MAVS using TRIM31 knockout mice [67]. Furthermore, TRIM21 could positively regulate IRF3-mediated type I IFN production through interaction with MAVS, thereby restricting RNA virus CVB3 replication and cardiac injury [68]. We compared the expression of TRIM21 and TRIM18 in CVB3 infected hearts and found TRIM18 expressed higher than TRIM21. We speculate that TRIM21 inhibits CVB3 induced myocarditis by serving as positive regulator in type I IFN production pathway, while TRIM18 promotes myocarditis induced by CVB3 through negatively regulating type I IFN production in antiviral innate immunity. Although TRIM29, TRIM31 and TRIM21 have essential roles in antiviral innate immunity in vivo, they only function their antiviral immunity against either RNA virus or DNA virus by targeting different substrates in different innate immune cells. Here, we identified TRIM18 as crucial negative regulator in antiviral immunity against both RNA and DNA viruses by recruiting PPM1A to inactivate TBK1 and block interactions of TBK1 with its upstream adaptors MAVS and STING for signal transduction in macrophages both in vitro and in vivo. It is reported that there are sex differences in immune responses that underlie COVID-19 disease outcomes [69] and men have higher COVID-19 mortality than women [70, 71]. Interestingly, TRIM18 gene is found on the X chromosome and mutations of TRIM18 gene are responsible for a rare genetic disease called X-linked Opitz G/BBB Syndrome (XLOS) [27]. Importantly, we found that male mice were more susceptible to CVB3 infection than females. Additionally, the public GEO profile database show that patients with SARS-CoV-2 infection have higher levels of TRIM18, while we demonstrated that WT mice with high expression of TRIM18 were more vulnerable to both RNA and DNA virus infections. Therefore, we hypothesize that TRIM18 may be a key biology factor associated with men’s higher risk of COVID-19-associated mortality, although testing this is beyond the scope of the current study.
TBK1 is a key adaptor protein shared by both RNA and DNA sensing pathways and is crucial for the activation of IRF3 and subsequent type I IFN induction [7, 72]. TBK1 is regulated by posttranslational modifications such as ubiquitination, phosphorylation, and acetylation. RNF128 promotes TBK1 activation by inducing its K63-linked polyubiquitination [73]. Likewise, TRIP, NLRP4-DTX4, Siglec1-TRIM27, and TRAF3IP3 also promote TBK1 degradation, although via K48-linked polyubiquitination [74,75,76,77,78]. It was reported that GSK3β, PPM1B, PP4 and PPM1A could modulate TBK1 activity by altering the TBK1 phosphorylation state [56, 79,80,81]. Additionally, HDAC9 deacetylates TBK1 and enhances TBK1 activation [82]. Although those above reports illustrate that TBK1 is under tight multi-layered control, the cellular regulatory mechanisms remain incompletely understood. Our data from IFN-β luciferase reporter and IP-MS assays suggest TRIM18 targets both RNA and DNA sensing pathways at TBK1 or its upstream nodes. In the macrophage steady state, TRIM18 interacts with PPM1A, which is reported to dephosphorylate and inactivate TBK1. Importantly, we detect interaction of TRIM18 with both PPM1A and TBK1 after virus infection, which drove us to further investigate the molecular mechanisms by which TRIM18 targets TBK1 to dampen type I IFN production in virus infection. Finally, we find TRIM18 serves as a negative regulator in antiviral innate immunity against RNA and DNA viruses through three novel mechanisms. First, TRIM18 recruits protein phosphatase PPM1A, which then interacts with and dephosphorylates TBK1 for its inactivation. Second, TRIM18 interacts with TBK1 and blocks its interactions with upstream adaptors MAVS and STING for preventing signal transduction during virus infection. Third, TRIM18 promotes the stability of PPM1A by inducing its K-63 linked polyubiquitination during virus infection. Furthermore, we found four lysine residues (Lys9, Lys12, Lys296, and Lys354) of PPM1A were critical sites for TRIM18-dependent ubiquitination and stability of PPM1A. Moreover, these four lysine residues of PPM1A mediated its inhibition of TBK1 triggered type I IFN production. Therefore, this work expands the regulatory landscape of the cytosolic sensing by RNA and DNA receptors and uncovers a function of TRIM18 in modulating innate immunity not previously appreciated.
Conclusions
In summary, we have identified TRIM18 as a novel negative regulator of viral myocarditis, lung inflammation and brain damage by downregulating innate immune activation against both RNA and DNA viruses. Our results underscore the pivotal role of TRIM18 in controlling both RNA and DNA virus infections, where TRIM18 provides a safeguard against aberrant and excessive type I IFN production as well as potential tissue damage and autoimmunity diseases. Importantly, the ongoing COVID-19 pandemic serves as a reminder that the new emerging RNA viruses remain a significant public health threat [25]. Thus, our findings may provide an opportunity for boosting protective antiviral immunity by inhibiting TRIM18. Our work may also be beneficial in the design of better pharmacological antagonists to improve the vaccine efficacy against both RNA and DNA viruses including SARS-CoV-2.
Availability of data and materials
All data relevant to the study are included in the article and in additional files. The reagents used in this publication are available from the corresponding author on reasonable request.
References:
Takeuchi O, Akira S. Pattern recognition receptors and inflammation. Cell. 2010;140(6):805–20.
Wu J, Chen ZJ. Innate immune sensing and signaling of cytosolic nucleic acids. Annu Rev Immunol. 2014;32:461–88.
Lin R, et al. Editorial: sensing DNA in antiviral innate immunity. Front Immunol. 2021;12: 644310.
Alexopoulou L, et al. Recognition of double-stranded RNA and activation of NF-kappaB by Toll-like receptor 3. Nature. 2001;413(6857):732–8.
Kato H, et al. RIG-I-like receptors: cytoplasmic sensors for non-self RNA. Immunol Rev. 2011;243(1):91–8.
**ng J, et al. Identification of poly(ADP-ribose) polymerase 9 (PARP9) as a noncanonical sensor for RNA virus in dendritic cells. Nat Commun. 2021;12(1):2681.
Sun L, et al. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science. 2013;339(6121):786–91.
Unterholzner L, et al. IFI16 is an innate immune sensor for intracellular DNA. Nat Immunol. 2010;11(11):997–1004.
Zhang Z, et al. The helicase DDX41 senses intracellular DNA mediated by the adaptor STING in dendritic cells. Nat Immunol. 2011;12(10):959–65.
Yoneyama M, et al. Shared and unique functions of the DExD/H-box helicases RIG-I, MDA5, and LGP2 in antiviral innate immunity. J Immunol. 2005;175(5):2851–8.
Kato H, et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature. 2006;441(7089):101–5.
Pichlmair A, et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5’-phosphates. Science. 2006;314(5801):997–1001.
Ablasser A, Hur S. Regulation of cGAS- and RLR-mediated immunity to nucleic acids. Nat Immunol. 2020;21(1):17–29.
Hur S. Double-stranded RNA sensors and modulators in innate immunity. Annu Rev Immunol. 2019;37:349–75.
Loo YM, Gale M Jr. Immune signaling by RIG-I-like receptors. Immunity. 2011;34(5):680–92.
Yoo JS, et al. Sensing viral invasion by RIG-I like receptors. Curr Opin Microbiol. 2014;20:131–8.
Liu S, et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science. 2015;347(6227):aaa2630.
Chen K, et al. Regulation of type I interferon signaling in immunity and inflammation: a comprehensive review. J Autoimmun. 2017;83:1–11.
Zhang E, et al. Mechanisms involved in controlling RNA virus-induced intestinal inflammation. Cell Mol Life Sci. 2022;79(6):313.
Yajima T, Knowlton KU. Viral myocarditis: from the perspective of the virus. Circulation. 2009;119(19):2615–24.
Tschöpe C, et al. Myocarditis and inflammatory cardiomyopathy: current evidence and future directions. Nat Rev Cardiol. 2021;18(3):169–93.
Marcocci ME, et al. Herpes simplex virus-1 in the brain: the dark side of a sneaky infection. Trends Microbiol. 2020;28(10):808–20.
Ruuskanen O, et al. Viral pneumonia. Lancet. 2011;377(9773):1264–75.
Pagliano P, et al. Characteristics of viral pneumonia in the COVID-19 era: an update. Infection. 2021;49(4):607–16.
Wang D, et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA. 2020. https://doi.org/10.1001/jama.2020.1585.
Hatakeyama S. TRIM family proteins: roles in autophagy, immunity, and carcinogenesis. Trends Biochem Sci. 2017;42(4):297–311.
Quaderi NA, et al. Opitz G/BBB syndrome, a defect of midline development, is due to mutations in a new RING finger gene on Xp22. Nat Genet. 1997;17(3):285–91.
Suzuki M, et al. MID1 and MID2 are required for Xenopus neural tube closure through the regulation of microtubule organization. Development. 2010;137(14):2329–39.
Lancioni A, et al. Lack of Mid1, the mouse ortholog of the Opitz syndrome gene, causes abnormal development of the anterior cerebellar vermis. J Neurosci. 2010;30(8):2880–7.
Nakamura T, et al. Novel role of Rac-Mid1 signaling in medial cerebellar development. Development. 2017;144(10):1863–75.
Pfirrmann T, et al. Hedgehog-dependent E3-ligase Midline1 regulates ubiquitin-mediated proteasomal degradation of Pax6 during visual system development. Proc Natl Acad Sci U S A. 2016;113(36):10103–8.
Collison A, et al. The E3 ubiquitin ligase midline 1 promotes allergen and rhinovirus-induced asthma by inhibiting protein phosphatase 2A activity. Nat Med. 2013;19(2):232–7.
Krauss S, et al. Translation of HTT mRNA with expanded CAG repeats is regulated by the MID1-PP2A protein complex. Nat Commun. 2013;4:1511.
Matthes F, et al. Inhibition of the MID1 protein complex: a novel approach targeting APP protein synthesis. Cell Death Discov. 2018;4:4.
Demir U, et al. Metformin anti-tumor effect via disruption of the MID1 translational regulator complex and AR downregulation in prostate cancer cells. BMC Cancer. 2014;14:52.
Köhler A, et al. A hormone-dependent feedback-loop controls androgen receptor levels by limiting MID1, a novel translation enhancer and promoter of oncogenic signaling. Mol Cancer. 2014;13:146.
Perry J, et al. A short pseudoautosomal region in laboratory mice. Genome Res. 2001;11(11):1826–32.
**ng J, et al. Identification of a role for TRIM29 in the control of innate immunity in the respiratory tract. Nat Immunol. 2016;17(12):1373–80.
**ng J, et al. TRIM29 promotes DNA virus infections by inhibiting innate immune response. Nat Commun. 2017;8(1):945.
**ng J, et al. TRIM29 negatively regulates the type I IFN production in response to RNA virus. J Immunol. 2018;201(1):183–92.
Zhang Z, et al. The E3 ubiquitin ligase TRIM21 negatively regulates the innate immune response to intracellular double-stranded DNA. Nat Immunol. 2013;14(2):172–8.
**ng J, et al. Differential inhibition of macrophage activation by lymphocytic choriomeningitis virus and pichinde virus is mediated by the Z protein N-terminal domain. J Virol. 2015;89(24):12513–7.
**ng J, et al. The Z proteins of pathogenic but not nonpathogenic arenaviruses inhibit RIG-I-like receptor-dependent interferon production. J Virol. 2015;89(5):2944–55.
**ng J, et al. DHX15 is required to control RNA virus-induced intestinal inflammation. Cell Rep. 2021;35(12): 109205.
**ng J, et al. Herpes simplex virus 1-encoded tegument protein VP16 abrogates the production of beta interferon (IFN) by inhibiting NF-kappaB activation and blocking IFN regulatory factor 3 to recruit its coactivator CBP. J Virol. 2013;87(17):9788–801.
**ng J, et al. Herpes simplex virus 1 tegument protein US11 downmodulates the RLR signaling pathway via direct interaction with RIG-I and MDA-5. J Virol. 2012;86(7):3528–40.
Zhang A, et al. EphA2 phosphorylates NLRP3 and inhibits inflammasomes in airway epithelial cells. EMBO Rep. 2020;21(7): e49666.
Cai Z, et al. Involvement of endoplasmic reticulum stress-mediated C/EBP homologous protein activation in coxsackievirus B3-induced acute viral myocarditis. Circ Heart Fail. 2015;8(4):809–18.
Lupfer C, et al. Receptor interacting protein kinase 2-mediated mitophagy regulates inflammasome activation during virus infection. Nat Immunol. 2013;14(5):480–8.
Kallewaard NL, et al. Tissue-specific deletion of the coxsackievirus and adenovirus receptor protects mice from virus-induced pancreatitis and myocarditis. Cell Host Microbe. 2009;6(1):91–8.
Soberman RJ, et al. CD200R1 supports HSV-1 viral replication and licenses pro-inflammatory signaling functions of TLR2. PLoS ONE. 2012;7(10): e47740.
Ashar HK, et al. The role of extracellular histones in influenza virus pathogenesis. Am J Pathol. 2018;188(1):135–48.
Ishikawa H, et al. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature. 2009;461(7265):788–92.
Sun Q, et al. The specific and essential role of MAVS in antiviral innate immune responses. Immunity. 2006;24(5):633–42.
Stelzer, G. et al. (2016) The genecards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinformatics 54, 1.30.1–1.30.33.
**ang W, et al. PPM1A silences cytosolic RNA sensing and antiviral defense through direct dephosphorylation of MAVS and TBK1. Sci Adv. 2016;2(7): e1501889.
Rajsbaum R, et al. TRIMmunity: the roles of the TRIM E3-ubiquitin ligase family in innate antiviral immunity. J Mol Biol. 2014;426(6):1265–84.
Versteeg GA, et al. The E3-ligase TRIM family of proteins regulates signaling pathways triggered by innate immune pattern-recognition receptors. Immunity. 2013;38(2):384–98.
Gack MU, et al. TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature. 2007;446(7138):916–20.
Yan J, et al. TRIM4 modulates type I interferon induction and cellular antiviral response by targeting RIG-I for K63-linked ubiquitination. J Mol Cell Biol. 2014;6(2):154–63.
Arimoto K, et al. Polyubiquitin conjugation to NEMO by triparite motif protein 23 (TRIM23) is critical in antiviral defense. Proc Natl Acad Sci U S A. 2010;107(36):15856–61.
Zhang J, et al. TRIM32 protein modulates type I interferon induction and cellular antiviral response by targeting MITA/STING protein for K63-linked ubiquitination. J Biol Chem. 2012;287(34):28646–55.
Tsuchida T, et al. The ubiquitin ligase TRIM56 regulates innate immune responses to intracellular double-stranded DNA. Immunity. 2010;33(5):765–76.
Qin Y, et al. TRIM9 short isoform preferentially promotes DNA and RNA virus-induced production of type I interferon by recruiting GSK3β to TBK1. Cell Res. 2016;26(5):613–28.
Zhao W, et al. Tripartite motif-containing protein 38 negatively regulates TLR3/4- and RIG-I-mediated IFN-β production and antiviral response by targeting NAP1. J Immunol. 2012;188(11):5311–8.
Dou Y, et al. Identification of the E3 Ligase TRIM29 as a Critical Checkpoint Regulator of NK Cell Functions. J Immunol. 2019;203(4):873–80.
Liu B, et al. The ubiquitin E3 ligase TRIM31 promotes aggregation and activation of the signaling adaptor MAVS through Lys63-linked polyubiquitination. Nat Immunol. 2017;18(2):214–24.
Liu H, et al. TRIM21 restricts coxsackievirus B3 replication, cardiac and pancreatic injury via interacting with MAVS and positively regulating IRF3-mediated type-I interferon production. Front Immunol. 2018;9:2479.
Takahashi T, et al. Sex differences in immune responses that underlie COVID-19 disease outcomes. Nature. 2020;588(7837):315–20.
Sharma G, et al. Sex differences in mortality from COVID-19 pandemic: are men vulnerable and women protected? JACC Case Rep. 2020;2(9):1407–10.
Bischof E, et al. Clinical trials for COVID-19 should include sex as a variable. J Clin Invest. 2020;130(7):3350–2.
Fitzgerald KA, et al. IKKepsilon and TBK1 are essential components of the IRF3 signaling pathway. Nat Immunol. 2003;4(5):491–6.
Song G, et al. E3 ubiquitin ligase RNF128 promotes innate antiviral immunity through K63-linked ubiquitination of TBK1. Nat Immunol. 2016;17(12):1342–51.
Zhang M, et al. TRAF-interacting protein (TRIP) negatively regulates IFN-β production and antiviral response by promoting proteasomal degradation of TANK-binding kinase 1. J Exp Med. 2012;209(10):1703–11.
Cui J, et al. NLRP4 negatively regulates type I interferon signaling by targeting the kinase TBK1 for degradation via the ubiquitin ligase DTX4. Nat Immunol. 2012;13(4):387–95.
Lin M, et al. USP38 inhibits type I interferon signaling by editing TBK1 ubiquitination through NLRP4 signalosome. Mol Cell. 2016;64(2):267–81.
Zheng Q, et al. Siglec1 suppresses antiviral innate immune response by inducing TBK1 degradation via the ubiquitin ligase TRIM27. Cell Res. 2015;25(10):1121–36.
Deng M, et al. TRAF3IP3 negatively regulates cytosolic RNA induced anti-viral signaling by promoting TBK1 K48 ubiquitination. Nat Commun. 2020;11(1):2193.
Lei CQ, et al. Glycogen synthase kinase 3β regulates IRF3 transcription factor-mediated antiviral response via activation of the kinase TBK1. Immunity. 2010;33(6):878–89.
Zhao Y, et al. PPM1B negatively regulates antiviral response via dephosphorylating TBK1. Cell Signal. 2012;24(11):2197–204.
Zhan Z, et al. Phosphatase PP4 negatively regulates type I IFN production and antiviral innate immunity by dephosphorylating and deactivating TBK1. J Immunol. 2015;195(8):3849–57.
Li X, et al. Methyltransferase Dnmt3a upregulates HDAC9 to deacetylate the kinase TBK1 for activation of antiviral innate immunity. Nat Immunol. 2016;17(7):806–15.
Acknowledgements
We thank Dr. Dorothy E. Lewis (Houston Methodist) for critical reading and editing of this manuscript. We also acknowledge the excellent services from the flow cytometry core and the pathology core at Houston Methodist Hospital in Houston, TX.
Funding
The work is supported by the following grants from the National Institutes of Health: R01AI155488 (to Z.Z.); R01AI080779 (to X.C.L.); and R01DE027879 (to T.C.C). J.X. is supported by the American Heart Association Career Development Award 20CDA35260116 (**ng). T.C.C is also supported by an Endowment from the Stowers Family Foundation.
Author information
Authors and Affiliations
Contributions
MF and JX designed and performed most of the experiments; AZ, YD, WL, JW, LJM and TCC helped with some of the experiments; JX and ZZ supervised the project; JX, TCC, XCL, and ZZ wrote and/or edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All animal protocols were performed according to the guidelines and approved by the Institutional Animal Care and Use Committee of Houston Methodist.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Table S1.
PPM1A is in the TRIM18-binding protein complex. Table S2. The potential ubiquitination sites at the PPM1A molecule. Table S3. Primers for qRT-PCR and PCR used in this study.
Additional file 2: Fig S1.
TRIM18 inhibits production of type I IFN by human THP-1 macrophages after stimulation with dsRNA and dsDNA, but not LPS. Fig S2. TRIM18 negatively regulates IFN-α production in human THP-1 macrophages after infection with RNA and DNA viruses. Fig S3. Trim18 gene targeting and TRIM18 expression in mouse macrophages and different tissues. Fig S4. TRIM18 does not affect expression of differentiation markers CD11b and F4/80 in mouse splenic macrophages. Fig S5. Knockout of TRIM18 enhances production of ISG15 and ISG56 in BMDM in response to dsRNA and dsDNA stimulations or infection with RNA and DNA viruses. Fig S6. TRIM18 is induced in human patients with SARS-CoV infection. Fig S7. TRIM18 inhibits IFN-β reporter activation mediated by overexpression of MDA5, MAVS, TBK1 and cGAS/STING, but not IKKi.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Fang, M., Zhang, A., Du, Y. et al. TRIM18 is a critical regulator of viral myocarditis and organ inflammation. J Biomed Sci 29, 55 (2022). https://doi.org/10.1186/s12929-022-00840-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12929-022-00840-z