Abstract
Background
Noncoding RNAs (ncRNAs), including microRNAs (miRNAs) and long noncoding RNAs (lncRNAs), are pivotal regulators involved in the pathogenic mechanism of multiple coronaviruses. Porcine deltacoronavirus (PDCoV) has evolved multiple strategies to escape the innate immune response of host cells, but whether ncRNAs are involved in this process during PDCoV infection is still unknown.
Results
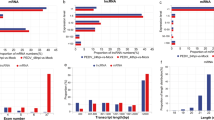
In this study, the expression profiles of miRNAs, lncRNAs and mRNAs in IPEC-J2 cells infected with PDCoV at 0, 12 and 24 hours postinfection (hpi) were identified through small RNA and RNA sequencing. The differentially expressed miRNAs (DEmiRNAs), lncRNAs (DElncRNAs) and mRNAs (DEmRNAs) were screened from the comparison group of IPEC-J2 cells at 0 and 12 hpi as well as the comparison group of IPEC-J2 cells at 12 and 24 hpi. The target genes of these DEncRNAs were predicted. The bioinformatics analysis of the target genes revealed multiple significantly enriched functions and pathways. Among them, the genes that were associated with innate immunity were specifically screened. The expression of innate immunity-related ncRNAs and mRNAs was validated by RT–qPCR. Competing endogenous RNA (ceRNA) regulatory networks among innate immunity-related ncRNAs and their target mRNAs were established. Moreover, we found that the replication of PDCoV was significantly inhibited by two innate immunity-related miRNAs, ssc-miR-30c-3p and ssc-miR-374b-3p, in IPEC-J2 cells.
Conclusions
This study provides a data platform to conduct studies of the pathogenic mechanism of PDCoV from a new perspective and will be helpful for further elucidation of the functional role of ncRNAs involved in PDCoV esca** the innate immune response.
Similar content being viewed by others
Introduction
Porcine deltacoronavirus (PDCoV) is a new porcine enteropathogenic virus. It causes watery diarrhea, dehydration, and high mortality in newborn piglets, which results in significant economic losses to the swine industry [1]. PDCoV is an enveloped RNA virus with a positive-stranded RNA genome that belongs to the genus Deltacoronavirus within the family Coronaviridae [2]. The full genome of PDCoV is approximately 25.4 kb in length and arranged in the following order: 5′ untranslated region (UTR), open reading frame 1a and b (ORF1ab), spike (S), envelope (E), membrane (M), non-structural gene 6 (NS6), nucleocapsid (N), non-structural gene 7 (NS7), and 3′ UTR [2]. PDCoV was initially reported in a molecular surveillance study in Hong Kong (China) in 2012, and then emerged in Ohio (USA) in 2014 [3, 4]. Subsequently, it spread to many countries including Thailand, South Korea, Canada, and China [5,6,7,22,23,24,25,26,43], and then further mapped to Sus scrofa 11.1. The known lncRNAs and mRNAs were obtained by comparing the assembled transcripts with annotated lncRNAs and mRNAs in the reference genome. Then, novel transcripts with length ≥ 200 bp and number of exons ≥2 were further retained from the transcripts that mapped to the reference genome and were located in intergenic regions. Then CNCI and CPC2 software were used to predict the protein-coding potential of novel transcripts. Novel lncRNAs were predicted by removing the novel transcripts that did not pass the protein-coding score test (CPC score < 0, CNCI score < 0), and the remaining transcripts were novel mRNAs. The expression of lncRNAs and mRNAs was calculated by fragments per kilobase of transcript per million mapped reads (FPKM) using RSEM software .
Analysis of small RNA-Seq data
High-quality clean reads were first screened from raw reads by removing the low-quality reads and adapter sequences. Then, the clean reads in a size range from 18 to 30 nt were further screened against the GenBank database (Release 209.0) [44] and Rfam (11.0) database [45] to remove small nucleolar RNA, small nuclear RNA, rRNA, tRNA, repeat sequences, exon and intron sequences. Eventually, the existed miRNAs were identified by searching all of the clean reads in the miRbase database [46] (Release 22). The known miRNAs that were still not included in the miRBase database were identified by aligning clean reads to those existed miRNAs of other species which included in miRBase. To distinguish with the existed miRNAs, x (−x) and y (−y) were used to represent the known miRNAs from the 5′ (−5p) and 3′ (−3p) arms of the miRNA precursor in this study. To identify the novel miRNAs, all of the unannotated clean reads were aligned with the swine reference genome (NCBI genome database: Sus scrofa 11.1) and the novel miRNAs were screened according to the results of hairpin structure prediction which performed by miReap software [47]. For gene identification, clean reads were screened from raw reads obtained from RNA sequencing by removing low-quality reads and reads with adapters or unknown nucleobases. Subsequently, the expressed genes and novel genes were identified by genome map**. Briefly, clean reads obtained from RNA sequencing were mapped to a swine reference genome (NCBI genome database: Sus scrofa 11.1), which was performed by HISAT software following the instructions of the software website (http://www.ccb.jhu.edu/software/hisat/index.shtml). The miRNA expression level was computed and normalized to transcripts per million (TPM) [48].
Screening of differentially expressed lncRNAs, mRNAs and miRNAs
The expression analysis of lncRNA, mRNA and miRNA in the two comparison groups were performed using DEGseq2 (http://www.bioconductor.org/packages/release/bioc/html/DESeq2.html) and edgeR (http://www.bioconductor.org/packages/release/bioc/html /edgeR.html). lncRNAs and mRNAs with a false discovery rate (FDR) value < 0.05 and a │log2 (fold change) │ > 1 and miRNAs with a P < 0.05 and a │log2 (fold change) │ > 1 were identified as differentially expressed ncRNAs or mRNAs.
Prediction of lncRNA and miRNA target genes
The potential target genes of DElncRNAs involved in cis- and trans-regulatory effects were predicted. We screened the mRNAs located 10 kb upstream and downstream of lncRNAs as cis target genes. The trans target genes were predicted by correlation analysis between lncRNAs and mRNAs. According to the FPKM values, the Pearson correlation coefficient between the lncRNAs and the mRNAs was calculated, and the threshold for positive correlation was set to Pearson correlation > 0.999.
Three software programs, TargetScan (Version 7.0), RNAhybrid (Version 2.1.2) and miRanda (Version 3.3a), were used to predict the targets of miRNAs. The genes that had reliable binding sites (minimum free energy (MFE) levels < − 10 kcal/mol, P values < 0.05) to miRNAs and commonly existed in 3 prediction results were considered to be credible targets of miRNAs. Targets involved in the innate immune response were annotated to InnateDB (https://www.innatedb.ca/).
RT-qRCR validation
Six miRNAs were randomly selected for RT-qPCR analysis using Bulge-loop™ miRNA RT-qPCR Primer Sets (RiboBio Inc., China) according to the manufacturer’s instructions. The primers for miRNAs and internal standard U6 were designed by RiboBio Inc. (China), and the sequences are covered by a patent. The RT-qPCR examination of lncRNAs and mRNAs was carried out using TB Green® Premix Ex Taq™ II (Tli RNaseH Plus) (Takara Biomedical Technology, China) following the manufacturer’s instructions. Primers for lncRNAs and mRNAs were designed and are listed in Additional file 10: Table S10. The RT-qPCRs were performed on an ABI7500 StepOnePlus Real-Time PCR System (Thermo Fisher Scientific, USA) and run in duplicate. U6 was used as the internal reference gene for miRNA relative expression validation, and GAPDH was used as the internal reference gene for lncRNA and mRNA relative expression validation. The relative expression level of each miRNA, lncRNA and mRNA was calculated using the 2−ΔΔCt method [49, 50].
Function and pathway enrichment analysis
To investigate the function of lncRNAs and miRNAs in IPEC-J2 cells, which were primarily influenced by PDCoV infection, GO functional enrichment analysis and KEGG pathway enrichment analysis of the predicted targets of DElncRNAs and DEmiRNAs were further performed by using of the GOseq R package and KOBAS software (3.0) [51]. The significantly enriched functional GO terms and pathways are represented in this study (P < 0.05).
Construction of competing endogenous RNA networks
Innate immunity-related lncRNAs, miRNAs and mRNAs were chosen for analysis. lncRNA-miRNA interactions were predicted by miRanda. Based on the results of the innate immunity-related target genes of DElncRNAs and DEmiRNAs as well as the lncRNA-miRNA interactions, visualization of the lncRNA-miRNA-mRNA interaction network was constructed using Cytoscape software (http://www.cytoscape.org/download.php).
Functional validation of miRNA for the regulation of PDCoV replication
Monolayer IPEC-J2 cells were transfected with miRNA mimic fluorescence controls under different concentrations using Lipofectamine 3000 (Thermo Fisher Scientific, USA). After 24 h, the fluorescence of the cells was observed by fluorescence microscopy (Olympus Corporation, Japan). Subsequently, monolayer IPEC-J2 cells were transfected with mimics and inhibitor as well as the controls of ssc-miR-30c-3p and ssc-miR-374b-3p, respectively, at a concentration of 150 nM. After 24 h, PDCoV at a final MOI of 1.0 was then used for adsorption in the transfected IPEC-J2 cells in an incubator at 37 °C in 5% CO2. The cells were respectively harvested at 24 h after transfection and at 12 hpi, and then the total RNA of each sample was extracted by TRIzol. miRNA RT product and cDNA was respectively obtained using the riboSCRIPT Reverse Transcription Kit (RiboBio Inc., China) and the PrimeScript™ RT reagent Kit (Perfect Real Time) (Takara Biomedical Technology, China) in an Applied Biosystems™ Veriti™ Dx 96-well Fast Thermal Cycler (Thermo Fisher Scientific, USA). The relative expression of ssc-miR-30c-3p and ssc-miR-374b-3p in the cells transfected with mimics or inhibitor as well as the controls of ssc-miR-30c-3p and ssc-miR-374b-3p were detected by RT-qPCR using Bulge-loop™ miRNA RT-qPCR Primer Sets (RiboBio Inc., China) according to the manufacturer’s instructions. The primers for miRNAs and internal standard U6 were designed by RiboBio Inc. (China), and the sequences are covered by a patent. The relative expression of the PDCoV M gene in different transfection groups was detected by RT-qPCR using TB Green® Premix Ex Taq™ II (Tli RNaseH Plus) (Takara Biomedical Technology, China). The RT-qPCRs were performed on an ABI7500 StepOnePlus Real-Time PCR System (Thermo Fisher Scientific, USA). Sequences of forward primer and reverse primer for RT-qPCR of PDCoV M gene are listed in Additional file 10: Table S10.
Availability of data and materials
The data of identification and integrated analysis of circRNAs and miRNAs has not been published, so the sequencing raw data cannot be disclosed temporarily. The raw data of sequencing can be obtained by contacting the corresponding author (E-mail: yanmh81971@126.com). The datasets generated during this study are included in the article and its additional files.
References
Zhang QZ, Yoo DW. Immune evasion of porcine enteric coronaviruses and viral modulation of antiviral innate signaling. Virus Res. 2016;226:128–41.
Feng Y, Xu ZW, Zhu L. Prevalence and phylogenetic analysis of porcine deltacoronavirus in Sichuan province. China Arch Virol. 2020;165(12):2883–9.
Woo PC, Lau SK, Lam CS, Lau CC, Tsang AK, Lau JH, et al. Discovery of seven novel mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J Virol. 2012;86(7):3995–4008.
Wang L, Byrum B, Zhang Y. Detection and genetic characterization of deltacoronavirus in pigs, Ohio, USA. Emerg Infect Dis. 2014;20(7):1227–30.
Janetanakit T, Lumyai M, Bunpapong N, Boonyapisitsopa S, Chaiyawong S, Nonthabenjawan N, et al. Porcine deltacoronavirus, Thailand, 2015. Emerg Infect Dis. 2016;22(4):757–9.
Lee S, Lee C. Complete genome characterization of Korean porcine deltacoronavirus strain KOR/KNU14-04/2014. Genome Announc. 2014;2(6):e01191–14.
Marthaler D, Raymond L, Jiang Y, Collins J, Rossow K, Rovira A. Rapid detection, complete genome sequencing, and phylogenetic analysis of porcine deltacoronavirus. Emerg Infect Dis. 2014;20:1347–50.
Dong N, Fang L, Zeng S, Sun Q, Chen H, **ao S. Porcine deltacoronavirus in mainland China. Emerg Infect Dis. 2015;21(12):2254–5.
Mai K, Feng J, Chen G, Li D, Zhou L, Bai Y, et al. The detection and phylogenetic analysis of porcine deltacoronavirus from Guangdong Province in southern China. Transbound Emerg Dis. 2018;65(1):166–73.
Zhang H, Han F, Shu X, Li Q, Ding Q, Hao C, et al. Co-infection of porcine epidemic diarrhoea virus and porcine deltacoronavirus enhances the disease severity in piglets. Transbound Emerg Dis. 2021. https://doi.org/10.1111/tbed.14144.
Chen L, Zhou Y, Li H. lncRNA, miRNA and lncRNA-miRNA interaction in viral infection. Virus Res. 2018;257:25–32.
Liu W, Ding C. Roles of lncRNAs in viral infections. Front Cell Infect Microbiol. 2017;7:205.
Ma Y, Wang C, Xue M, Fu F, Zhang X, Li L, et al. The coronavirus transmissible gastroenteritis virus evades the type I interferon response through IRE1-mediated manipulation of the microRNA miR-30a-5p/SOCS1/3 axis. J Virol. 2018;92(22):e00728–18.
Gao J, Pan Y, Xu Y, Zhang W, Zhang L, Li X, et al. Unveiling the long non-coding RNA profile of porcine reproductive and respiratory syndrome virus-infected porcine alveolar macrophages.BMC. Genomics. 2021;22:177.
Qi X, Wang T, Xue Q, Zhen Li Z, Yang B, Wang J. MicroRNA expression profling of goat peripheral blood mononuclear cells in response to peste des petits ruminants virus infection. Vet Res. 2018;49(1):62.
Jung K, Miyazaki A, Hu H, Saif LJ. Susceptibility of porcine IPEC-J2 intestinal epithelial cells to infection with porcine deltacoronavirus (PDCoV) and serum cytokine responses of gnotobiotic pigs to acute infection with IPEC-J2 cell culture-passaged PDCoV. Vet Microbiol. 2018;221:49–58.
Brosnahan AJ, Brown DR. Porcine IPEC-J2 intestinal epithelial cells in microbiological investigations. Vet Microbiol. 2012;156:229–37.
Lin H, Li B, Chen L, Ma Z, He K, Fan H. Differential protein analysis of IPEC-J2 cells infected with porcine epidemic diarrhea virus pandemic and classical strains elucidates the pathogenesis of infection. J Proteome Res. 2017;16:2113–20.
**a L, Dai L, Yu Q, Yang Q. Persistent TGEV infection enhances ETEC K88 adhesion by promoting epithelial-mesenchymal transition in intestinal epithelial cells. J Virol. 2017;91(21):e01256–17.
Tang X, Lan T, Wu R, Zhou Z, Chen Y, Sun Y, et al. Analysis of long non-coding RNAs in neonatal piglets at different stages of porcine deltacoronavirus infection. BMC Vet Res. 2019;15(1):111.
Liu J, Wang F, Du L, Li J, Yu T, ** Y, et al. Comprehensive genomic characterization analysis of lncRNAs in cells with porcine delta coronavirus infection. Front Microbiol. 2020;10:3036.
Luo J, Fang L, Dong N, Fang P, Ding Z, Wang D, et al. Porcine deltacoronavirus (PDCoV) infection suppresses RIG-I-mediated interferon-β production. Virology. 2016;495:10–7.
Zhu X, Wang D, Zhou J, Pan T, Chen J, Yang Y, et al. Porcine deltacoronavirus nsp5 antagonizes typeI interferon signaling by cleaving STAT2. J Virol. 2017;91(10):e00003–17.
Zhu X, Fang L, Wang D, Yang Y, Chen J, Ye X, et al. Porcine deltacoronavirus nsp5 inhibits interferon-βproduction through the cleavage of NEMO. Virology. 2017;502:33–8.
Fang P, Fang L, Ren J, Hong Y, Liu X, Zhao Y, et al. Porcine deltacoronavirus accessory protein NS6antagonizes interferon beta production by interfering with the binding of RIG-I/MDA5 to double-strandedRNA. J Virol. 2018;92(15):e00712–8.
Liu X, Fang P, Fang L, Hong Y, Zhu X, Wang D, et al. Porcine deltacoronavirus nsp15 antagonizes interferon-β production independently of its endoribonuclease activity. Mol Immunol. 2019;114:100–7.
Fang P, Fang L, **a S, Ren J, Zhang J, Bai D, et al. Porcine deltacoronavirus accessory protein NS7a antagonizes IFN-β production by competing with TRAF3 and IRF3 for binding to IKKε. Front Cell Infect Microbiol. 2020;10:257.
Bai D, Fang L, **a S, Ke W, Wang J, Wu X, et al. Porcine deltacoronavirus (PDCoV) modulates calcium influx to favor viral replication. Virology. 2020;539:38–48.
Xu Z, Zhang Y, Cao Y. The roles of apoptosis in swine response to viral infection and pathogenesis of swine enteropathogenic coronaviruses. Front Vet Sci. 2020;7:572425.
Kong L, Zhang Y, Ye ZQ, Liu XQ, Zhao SQ, Wei L, et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007;35(Web Server issue):W345–9.
Sun L, Luo H, Bu D, Zhao G, Yu K, Zhang C, et al. Utilizing sequence intrinsic composition to classify protein-coding and long noncoding transcripts. Nucleic Acids Res. 2013;41(17):e166.
Augustino SMA, Xu Q, Liu X, Mi S, Shi L, Liu Y, et al. Integrated analysis of lncRNAs and mRNAs reveals key trans-target genes associated with ETEC-F4ac adhesion phenotype in porcine small intestine epithelial cells. BMC Genomics. 2020;21:780.
Jiang S, Li FQ, Li XL, Wang LL, Zhang L, Lu C, et al. Transcriptome analysis of PK-15 cells in innate immune response to porcine deltacoronavirus infection. PLoS One. 2019;14(10):e0223177.
Mair KH, Sedlak C, Käser T, Pasternak A, Levast B, Gerner W, et al. The porcine innate immune system: an update. Dev Comp Immunol. 2014;45(2):321–43.
Kanehisa M, Goto S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30.
Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505(7483):344–52.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504.
Li N, Huang K, Chen Y, Huang Z, Zhang Y, Leng C, et al. MicroRNA ssc-miR-124a exhibits antiviral activity against porcine reproductive and respiratory syndrome virus via suppression of host genes CD163. Vet Microbiol. 2021;261:109216.
Huang S, Cheng A, Cui M, Pan Y, Wang M, Huang J, et al. Duck Tembusu virus promotes the expression of suppressor of cytokine signaling 1 by downregulating miR-148a-5p to facilitate virus replication. Infect Genet Evol. 2020;85:104392.
Ma Y, Wang C, Xue M, Fu F, Zhang X, Li L, et al. The coronavirus transmissible gastroenteritis virus evades the type I interferon response through IRE1α-mediated manipulation of the microRNA miR-30a-5p/SOCS1/3 axis. J Virol. 2018;92(22):e00728–18.
Zheng L, Li X, Yan M, Ren W, Zhang L, Lu C, et al. Isolation, identification and biological characteristics analysis of porcine deltacoronavirus TJ1. China Anim Husbandry Vet Med. 2018;45(1):219–24 (in Chinese).
Jiang S, Li F, Li X, Wang L, Zheng L, Zhang L, et al. Establishment and preliminary application of TaqMan fluorescence quantitative PCR assay for detecting porcine deltacoronavirus. Prog in Vet Med. 2019;40(10):10–7 (in Chinese).
Pertea M, Pertea GM, Antonescu CM, Chang T-C, Mendell JT, Salzberg SL. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33(3):290–5.
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Ostell J, Pruitt KD, et al. GenBank. Nucleic Acids Res. 2018;46(D1):D41–7.
Griffiths-Jones S, Bateman A, Marshall M, Khanna A, Eddy SR. Rfam: an RNA family database. Nucleic Acids Res. 2003;31(1):439–41.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006;34(Database issue):D140–4.
Xu LN, Ling YH, Wang YQ, Wang ZY, Hu BJ, Zhou ZY, et al. Identification of differentially expressed microRNAs between bacillus thuringiensis Cry1Ab-resistant and -susceptible strains of Ostrinia furnacalis. Sci Rep. 2015;5:15461.
Jirak P, Wernly B, Lichtenauer M, Franz M, Knost T, Abusamrah T, et al. Next-generation sequencing analysis of circulating micro-RNA expression in response to parabolic flight as a spaceflight analogue. NPJ Microgravity. 2020;6(1):31.
Schmittgen TD, Lee EJ, Jiang J, Sarkar A, Yang L, Elton TS, et al. Real-time PCR quantifcation of precursor and mature microRNA. Methods. 2008;44:31–8.
Schmittgen TD, Livak KJ. Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc. 2008;3(6):1101–8.
Shi LL, Zhang N, **e XM, Chen YJ, Wang R, Shen L, et al. Transcriptome profile of rat genes in injured spinal cord at different stages by RNA-sequencing. BMC Genomics. 2017;18(1):173.
Acknowledgments
We would like to acknowledge Research Innovation Team of Swine Disease Lab at Tian** Institute of Animal Husbandry and Veterinary Medicine, Tian** Academy of Agricultural Sciences, and State Key Laboratory of Veterinary Biotechnology at Harbin Veterinary Research Institute, Chinese Academy of Agricultural Sciences for their assistance and support for the conduct of this research project.
Funding
This work is supported by State Key Laboratory of Veterinary Biotechnology Foundation (grant number: SKLVBF202113); Key Project of Advanced Manufacturing Technology for High Quality Veterinary Drugs (grant number: 17ZXGSNC00080); Key Project of Tian** Science and Technology Support (grant number: 19YFZCSN00570); Tian** Modern Agricultural Industry Technology System Project (grant number: ITTPRS2021003); the Central Public-interest Scientific Institution Basal Research Fund, National Data Center of Animal Health.
Author information
Authors and Affiliations
Contributions
SJ: Data analysis, Validation experiments, Manuscript preparation; JFC, XLL, WKR, FXL, TW, CL, ZMD and XXT: Methodology; LZ, LLW, CL, and JJC: Software, Figure; MHY and LF: Supervision, Design, revision. MHY and LF are co-correspondence authors, which contributed equally to this work. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1.
Overview of the RNA sequencing data.
Additional file 2.
Overview of the small RNA sequencing data.
Additional file 3.
Differentially expressed lncRNAs, miRNAs and mRNAs.
Additional file 4.
Prediction results of the target genes of DElncRNAs.
Additional file 5.
Prediction results of the target genes of DEmiRNAs.
Additional file 6.
Innate immunity-related target genes of DElncRNAs.
Additional file 7.
Innate immunity-related target genes of DEmiRNAs.
Additional file 8.
Enrichment analysis result of DElncRNAs' target genes.
Additional file 9.
Enrichment analysis result of DEmiRNAs' target genes.
Additional file 10.
Primers used for qRT-PCR examination in this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Jiang, S., Chen, J., Li, X. et al. Identification and integrated analysis of lncRNAs and miRNAs in IPEC-J2 cells provide novel insight into the regulation of the innate immune response by PDCoV infection. BMC Genomics 23, 486 (2022). https://doi.org/10.1186/s12864-022-08722-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-022-08722-2