Abstract
During neurogenesis, generation, migration and integration of the correct numbers of each neuron sub-type depends on complex molecular interactions in space and time. MicroRNAs represent a key control level allowing the flexibility and stability needed for this process. Insight into the role of this regulatory pathway in the brain is still limited. We performed a sequential experimental approach using postnatal olfactory bulb neurogenesis in mice, starting from global expression analyses to the investigation of functional interactions between defined microRNAs and their targets. Deep sequencing of small RNAs extracted from defined compartments of the postnatal neurogenic system demonstrated that the miR-200 family is specifically induced during late neuronal differentiation stages. Using in vivo strategies we interfered with the entire miR-200 family in loss- and gain-of-function settings, showing a role of miR-200 in neuronal maturation. This function is mediated by targeting the transcription factor Zeb2. Interestingly, so far functional interaction between miR-200 and Zeb2 has been exclusively reported in cancer or cultured stem cells. Our data demonstrate that this regulatory interaction is also active during normal neurogenesis.
Similar content being viewed by others
Introduction
Fine-tuning of gene expression is a fundamental requirement for the control of developmental processes. This is particularly evident during nervous system development, where stem cell populations generate a multitude of neuronal and glial cell types in a temporally and quantitatively perfectly orchestrated manner. After their generation, precursors migrate to their respective target structures and form functional connections with their environment. Neurogenesis continues into postnatal and adult stages in defined regions of the mammalian brain, making the control and stabilization of regulatory processes a lifelong requirement1. It is evident that complex molecular networks, superposed levels of control and tight interactions between regulatory mechanisms guard induction and maintenance of neurogenesis. MicroRNAs (microRNAs) represent one key control level providing the needed flexibility and stability2.
Dicer mutant mouse lines have been widely used to show the general involvement of the microRNA pathway in brain development and function3,4,5,6,7. Specific microRNAs have been implicated in the control of neurogenesis at different levels. First, they act at the level of initiation of differentiation and the progression of progenitors towards a differentiated state. For example, miR-124 and the miR-9/miR-9* duplex inhibit the expression of molecular components that oppose neuronal differentiation8,37,38,39,40. Moreover, several specific microRNAs were shown to regulate neurogenesis in vivo. For example miR-9 and miR-124 are general regulators of the neurogenic process12,41 whereas other microRNAs were shown to regulate specific steps during neurogenesis (for review: refs 2 and 42). Finally few microRNAs have been shown to regulate neuronal fate decisions, as exemplified by miR-7a and miR-17-3p13,14,43. However, our understanding of the in vivo role of the microRNA pathway in neurogenesis is still limited.
This limitation is mostly due to technical issues, since the particular molecular structure of microRNAs renders traditional approaches difficult. For example, the lack of polyA-tails prevents the use of classical linear amplification protocols and therefore genomic analyses with limited amounts of material. Moreover, the small size and the strong sequence homology between microRNA molecules makes in situ hybridization experiments more problematic. Therefore, a global representation of dynamic microRNA expression along neuron differentiation from neural stem cell to mature neurons has, to the best of our knowledge, not been reported.
Here we use the unique features of postnatal OB neurogenesis to investigate the expression of microRNAs during neurogenesis at high resolution. Indeed, during postnatal neurogenesis in the OB, the main neurogenic stages are spatially distinct and can be physically isolated: the VZ-SVZ region contains mitotic progenitors, post-mitotic neuroblasts migrate in the RMS and young neurons terminally differentiate and integrate into the OB circuitry18.
Our deep sequencing approach described the dynamics and regionalization of all known microRNAs during the different phases of the forebrain neurogenic process. Among the microRNAs expressed in the system some appeared stably expressed, whereas other are tightly regulated. We focused our functional analysis on the miR-200 family. All members of this family, despite being coded by two independent loci, are induced and function at late stages of the OB neurogenic process.
miR-200 family members are major regulators of tumorigenesis, notably through the capacity to inhibit the transcription factors Zeb1 and Zeb2, two major factors controlling epithelial–mesenchymal-transition44,45,46,47. Our immunohistological analyses and in situ data (the Allen Brain project) demonstrate strong Zeb2 expression in the forebrain neurogenic system. Moreover, both, Zeb2 loss-of-function and miR-200 gain-of-function led to a comparable phenotype, the premature expression of the late neuronal subtype marker calretinin.
Two alternative explanations can be proposed for this observation. First, repression of Zeb2 by miR-200 microRNAs has a direct impact on differentiation of at least a subfraction of neuronal progenitors. Indeed, in the develo** cortex conditional deletion of Zeb2 induced premature neuronal and glial differentiation48. Such a role for Zeb2 in the differentiation process would account for the appearance of calretinin positive cells in the RMS in the context of miR-200 overexpression. It would also explain the lack of differentiation, as measured by decreased NeuN staining, when miR-200 is inhibited. Second, induction of calretinin expression in the RMS might be a consequence of slowed neuronal migration. Indeed, it has been shown that interfering with migration through knockdown of DCX leads to the appearance of calretinin positive cells in the RMS49. Moreover, direct roles of Zeb2 in in migration of metastatic cancer cells30,50 and cortical interneurons36 have been shown.
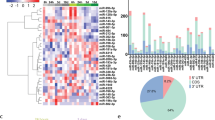
Another question concerns the observation that only a small fraction of neuronal precursors shows altered differentiation after interference with miR-200 expression. The quantity of NeuN negative cells in the OB after miR-200 knockdown increases by only 5%, while premature expression of the family induces calretinin in less than 10% of all transfected cells. It should be noted that such minor alterations, typical for the fine-tuning function of microRNAs, would likely be missed in the analysis of other neurogenic processes, which do not permit the same high-resolution analysis. The limited effects might be due to the fact that only a subfraction of the transfected cells are responsive to either inhibition or increase of miR-200 microRNAs. Our finding that the miR-200 promoter fragment that we used to drive GFP expression is only active in a small fraction of the transfected neurons supports this potential lack of competence. Alternatively, it is possible that other microRNAs have redundant functions in the system. In line with this idea, we found that the miR-183/96/182 cluster, a group of microRNAs that has been implicated in the maintenance of retinal neuron integrity51,52 and shares common predicted targets53, appears as nearest neighbors with the miR-200 family in our heat map representation (Fig. 2a,b). Functional studies using tools targeting both groups of microRNAs in the forebrain compartments will be necessary to address this issue.
Material and Methods
Mouse lines
Mice carrying floxed Zeb2 alleles (Zeb2fl/fl)54 were crossed to the RCE reporter mice55. Resulting progeny was subsequently crossed with Gsh2-Cre mice56 to generate Gsh2-Cre; RCE; Zeb2fl/fl mutant mice or Gsh2-Cre; RCE; Zeb2fl/wt control mice. Animal experiments were carried out in accordance to European Communities Council Directive and approved by French ethical committees (Comité d’Ethique pour l′expérimentation animale n°14; permission number: 62-12112012).
Plasmid constructs
The pCX-Cre and pCX-GFP vectors are derived from pCX-MCS257. To generate the vector expressing gfp under the control of the human miR-200b/miR-200a/miR-429 regulatory sequence we subcloned gfp from pCX-GFP into the pGL3-1574/ + 120 vector obtained from addgene. The miR-200 expression vector (miR-200-gof) was generated by PCR amplification of both miR-200 clusters from CD1 mouse genomic DNA and sub-cloning of amplified fragments into pCX-MCS2. The sponge construct was designed according to58 with 4 repetitions of 2 oligonucleotides (5′-GACACATCGTTACTCTCAGTGTTAGACACGGCATTACTCTCAGTATTA and 5′-GACTTCATCATTACTCCCAGTATTAGACCCATCTTTACTCTCAGTGTTA) partially complementary to any member of the miR-200 family were placed behind a destabilized GFP gene in pCX-d2-GFP plasmid.
Zeb2 3′UTR was PCR-amplified from CD1 mouse brain cDNA and cloned into the pMir-Glo vector (Promega) to generate the 3′UTR-Zeb2 pmiRGlo vector.
RNA extraction and deep sequencing
Total RNA was extracted from CD1 mice using the miRNeasy kit (Qiagen). RNA was extracted from dorsal or lateral VZ-SVZ at P1 and P6, from RMS at P15 and P28 and from OB at P15 and P28. All samples were dissected in triplicate. Deep sequencing analysis were performed on the 15–50 bp RNA molecules using the Applied Biosystems SOLiD™ System. For each sample, results were normalized for each microRNA as number of reads per million. Results were submitted to GEO (GSE60817).
Cell sorting (FACS, MACS), qRT-PCR and luciferase assay
For isolation of OB interneurons from GAD67-GFP knock-in mice59, whole bulbs of P30 animals were dissected and dissociated by Trypsin/DNAse digestion. GFP cells were purified using MoFlow (Beckman-Coulter) flow cytometer. For Zeb2 mutant analyses, lateral SVZ of P2 control brains were dissected and dissociated by Papain (Sigma)/DNAse digestion. GFP Cells (provided by the RCE locus) were sorted using a FACSAriaI (BD Biosciences).
To separate neuronal from glial cells by MACS, OB from P30 CD1 mice were subjected to Trypsin/DNAse dissociation. Both neuronal and glial enriched fractions were recovered from single cell suspension containing approximately 1 × 106 cells using the “Neuron Isolation Kit” (Miltenyi).
RNAs were extracted from sorted cells using the mRNeasy or miRNeasy kit (Qiagen). cDNAs were prepared using superscript-III (Life-Technologies) and qPCR was performed using SYBR-GreenER qPCR SuperMix (Life-Technologies), except for Zeb2KO sorted cells which were performed on a LightCycler 480 Instrument (Roche) using SYBR Green PCR Master Mix (Roche). Beta-Actin was used as reference gene. Primers sequences are given in SI. MicroRNA qPCR was performed using LNA-qRT-PCR system from Exiqon and using U6 as reference gene.
Luciferase assay was performed on HeLa cells 48 h after Lipofectamine 2000 (Life-Technologies) mediated transfection using the Dual-Luciferase Reporter Assay (Promega) and a Luminometer (Berthold Technologies).
Immunohistochemistry and Image analysis
Brain sections and staining experiments were performed as in19 except those performed on Gsh2-Cre; RCE; Zeb2fl/fl mice brains processed as in48. Primary antibodies used are: Calretinin (rabbit, Swant, 1/1000), GFP (chicken, Aves, 1/500), mouse IgG1 anti-NeuN (Millipore, 1:100), rat Igg2a anti-BrdU (AbD Serotec (Oxford B), 1/1000). Images were taken using a fluorescence microscope (Axiolmager Z1, ApoTome system, Zeiss) except for Gsh2-Cre; RCE; Zeb2fl/fl sections (Leica DMR microscope) and for spine density measurement (laser confocal scanning microscope, LSM510, Zeiss - magnification: 63x). Data in graphics are presented as mean ± s.e.m of values obtained on n samples (*P < 0,05. **P < 0,01, ***P < 0,001). For BrdU incorporation analysis, animals at 2 dpe were injected once with a BrdU solution (50 μg/g body weight, Sigma, Saint-Louis MO) 2 hours before perfusion. BrdU staining was performed after 15 min incubation at 37° in 2N HCl-0.5%. In Fig. 4c, Zeb2 expression level per transfected cell was assessed as follows. Transfected cells in the RMS were identified based on GFP expression. Quantification of Zeb2 staining was performed using ImageJ software on a single z-plan focused on the nucleus (chosen using DAPI staining). A ROI was subsequently drawn inside the nucleus area and the mean intensity of Zeb2 staining signal was then measured across the ROI.
Additional Information
How to cite this article: Beclin, C. et al. miR-200 family controls late steps of postnatal forebrain neurogenesis via Zeb2 inhibition. Sci. Rep. 6, 35729; doi: 10.1038/srep35729 (2016).
Change history
21 December 2016
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Lim, D. A. & Alvarez-Buylla, A. Adult neural stem cells stake their ground. Trends Neurosci 37, 563–571 (2014).
Follert, P., Cremer, H. & Beclin, C. MicroRNAs in brain development and function: a matter of flexibility and stability. Front Mol Neurosci 7, 5 (2014).
Kawase-Koga, Y., Otaegi, G. & Sun, T. Different timings of Dicer deletion affect neurogenesis and gliogenesis in the develo** mouse central nervous system. Dev Dyn 238, 2800–2812 (2009).
Davis, T. H. et al. Conditional loss of Dicer disrupts cellular and tissue morphogenesis in the cortex and hippocampus. J Neurosci 28, 4322–4330 (2008).
Cuellar, T. L. et al. Dicer loss in striatal neurons produces behavioral and neuroanatomical phenotypes in the absence of neurodegeneration. Proc Natl Acad Sci USA 105, 5614–5619 (2008).
De Pietri Tonelli, D. et al. miRNAs are essential for survival and differentiation of newborn neurons but not for expansion of neural progenitors during early neurogenesis in the mouse embryonic neocortex. Development 135, 3911–3921 (2008).
Schaefer, A. et al. Cerebellar neurodegeneration in the absence of microRNAs. J Exp Med 204, 1553–1558 (2007).
Zhao, C., Sun, G., Li, S. & Shi, Y. A feedback regulatory loop involving microRNA-9 and nuclear receptor TLX in neural stem cell fate determination. Nat Struct Mol Biol 16, 365–371 (2009).
Packer, A. N., **ng, Y., Harper, S. Q., Jones, L. & Davidson, B. L. The bifunctional microRNA miR-9/miR-9* regulates REST and CoREST and is downregulated in Huntington’s disease. J Neurosci 28, 14341–14346 (2008).
Conaco, C., Otto, S., Han, J. J. & Mandel, G. Reciprocal actions of REST and a microRNA promote neuronal identity. Proc Natl Acad Sci USA 103, 2422–2427 (2006).
Yoo, A. S., Staahl, B. T., Chen, L. & Crabtree, G. R. MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 460, 642–646 (2009).
Cheng, L. C., Pastrana, E., Tavazoie, M. & Doetsch, F. miR-124 regulates adult neurogenesis in the subventricular zone stem cell niche. Nat Neurosci 12, 399–408 (2009).
de Chevigny, A. et al. miR-7a regulation of Pax6 controls spatial origin of forebrain dopaminergic neurons. Nat Neurosci 15, 1120–1126 (2012).
Chen, J. A. et al. Mir-17-3p controls spinal neural progenitor patterning by regulating Olig2/Irx3 cross-repressive loop. Neuron 69, 721–735 (2011).
Schratt, G. M. et al. A brain-specific microRNA regulates dendritic spine development. Nature 439, 283–289 (2006).
Fiore, R. et al. Mef2-mediated transcription of the miR379-410 cluster regulates activity-dependent dendritogenesis by fine-tuning Pumilio2 protein levels. EMBO J 28, 697–710 (2009).
Alvarez-Buylla, A. & Garcia-Verdugo, J. M. Neurogenesis in adult subventricular zone. J Neurosci 22, 629–634 (2002).
Lledo, P. M., Merkle, F. T. & Alvarez-Buylla, A. Origin and function of olfactory bulb interneuron diversity. Trends Neurosci 31, 392–400 (2008).
Boutin, C., Diestel, S., Desoeuvre, A., Tiveron, M. C. & Cremer, H. Efficient in vivo electroporation of the postnatal rodent forebrain. PLoS One 3, e1883 (2008).
Sun, A. X., Crabtree, G. R. & Yoo, A. S. MicroRNAs: regulators of neuronal fate. Curr Opin Cell Biol 25, 215–221 (2013).
Song, R. et al. miR-34/449 miRNAs are required for motile ciliogenesis by repressing cp110. Nature 510, 115–120 (2014).
Wu, J. et al. Two miRNA clusters, miR-34b/c and miR-449, are essential for normal brain development, motile ciliogenesis, and spermatogenesis. Proc Natl Acad Sci USA 111, E2851–E2857 (2014).
Marcet, B. et al. Control of vertebrate multiciliogenesis by miR-449 through direct repression of the Delta/Notch pathway. Nat Cell Biol 13, 693–699 (2011).
Spassky, N. et al. Adult ependymal cells are postmitotic and are derived from radial glial cells during embryogenesis. J Neurosci 25, 10–18 (2005).
Pandey, A. et al. Critical role of the miR-200 family in regulating differentiation and proliferation of neurons. J Neurochem 133, 640–652 (2015).
Choi, P. S. et al. Members of the miRNA-200 family regulate olfactory neurogenesis. Neuron 57, 41–55 (2008).
Acuna-Goycolea, C., Tamamaki, N., Yanagawa, Y., Obata, K. & van den Pol, A. N. Mechanisms of neuropeptide Y, peptide YY, and pancreatic polypeptide inhibition of identified green fluorescent protein-expressing GABA neurons in the hypothalamic neuroendocrine arcuate nucleus. J Neurosci 25, 7406–7419 (2005).
Shepherd, G. M. The synaptic organization of the brain (Oxford University Press, Oxford, 2004).
Bracken, C. P. et al. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res 68, 7846–7854 (2008).
Brabletz, S. & Brabletz, T. The ZEB/miR-200 feedback loop–a motor of cellular plasticity in development and cancer? EMBO Rep 11, 670–677 (2010).
Price, J. L. & Powell, T. P. The mitral and short axon cells of the olfactory bulb. J Cell Sci 7, 631–651 (1970).
Kosaka, K. et al. Chemically defined neuron groups and their subpopulations in the glomerular layer of the rat main olfactory bulb. Neurosci Res 23, 73–88 (1995).
Merkle, F. T., Mirzadeh, Z. & Alvarez-Buylla, A. Mosaic organization of neural stem cells in the adult brain. Science 317, 381–384 (2007).
Merkle, F. T. et al. Adult neural stem cells in distinct microdomains generate previously unknown interneuron types. Nat Neurosci 17, 207–214 (2014).
Lopez-Juarez, A. et al. Gsx2 controls region-specific activation of neural stem cells and injury-induced neurogenesis in the adult subventricular zone. Genes Dev 27, 1272–1287 (2013).
van den Berghe, V. et al. Directed migration of cortical interneurons depends on the cell-autonomous action of Sip1. Neuron 77, 70–82 (2013).
Shao, N. Y. et al. Comprehensive survey of human brain microRNA by deep sequencing. BMC Genomics 11, 409 (2010).
Landgraf, P. et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129, 1401–1414 (2007).
Krichevsky, A. M., King, K. S., Donahue, C. P., Khrapko, K. & Kosik, K. S. A microRNA array reveals extensive regulation of microRNAs during brain development. RNA 9, 1274–1281 (2003).
Yao, M. J. et al. Transcriptome analysis of microRNAs in develo** cerebral cortex of rat. BMC Genomics 13, 232 (2012).
Shibata, M., Nakao, H., Kiyonari, H., Abe, T. & Aizawa, S. MicroRNA-9 regulates neurogenesis in mouse telencephalon by targeting multiple transcription factors. J Neurosci 31, 3407–3422 (2011).
Barca-Mayo, O. & De Pietri Tonelli, D. Convergent microRNA actions coordinate neocortical development. Cell Mol Life Sci 71, 2975–2995 (2014).
de Chevigny, A., Cremer, H. & Core, N. [Micro-RNA miR-7a controls the production of dopaminergic neurons in the mouse forebrain]. Med Sci (Paris) 29, 153–155 (2013).
Korpal, M., Lee, E. S., Hu, G. & Kang, Y. The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 283, 14910–14914 (2008).
Park, S. M., Gaur, A. B., Lengyel, E. & Peter, M. E. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 22, 894–907 (2008).
Gregory, P. A. et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 10, 593–601 (2008).
Burk, U. et al. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep 9, 582–589 (2008).
Seuntjens, E. et al. Sip1 regulates sequential fate decisions by feedback signaling from postmitotic neurons to progenitors. Nat Neurosci 12, 1373–1380 (2009).
Belvindrah, R., Nissant, A. & Lledo, P. M. Abnormal neuronal migration changes the fate of develo** neurons in the postnatal olfactory bulb. J Neurosci 31, 7551–7562 (2011).
Comijn, J. et al. The two-handed E box binding zinc finger protein SIP1 downregulates E-cadherin and induces invasion. Mol Cell 7, 1267–1278 (2001).
Busskamp, V. et al. miRNAs 182 and 183 are necessary to maintain adult cone photoreceptor outer segments and visual function. Neuron 83, 586–600 (2014).
Lumayag, S. et al. Inactivation of the microRNA-183/96/182 cluster results in syndromic retinal degeneration. Proc Natl Acad Sci USA 110, E507–E516 (2013).
Kundu, S. T. et al. The miR-200 family and the miR-183~96~182 cluster target Foxf2 to inhibit invasion and metastasis in lung cancers. Oncogene 35, 173–186 (2016).
Higashi, Y. et al. Generation of the floxed allele of the SIP1 (Smad-interacting protein 1) gene for Cre-mediated conditional knockout in the mouse. Genesis 32, 82–84 (2002).
Sousa, V. H., Miyoshi, G., Hjerling-Leffler, J., Karayannis, T. & Fishell, G. Characterization of Nkx6-2-derived neocortical interneuron lineages. Cereb Cortex 19 Suppl 1, i1–10 (2009).
Kessaris, N. et al. Competing waves of oligodendrocytes in the forebrain and postnatal elimination of an embryonic lineage. Nat Neurosci 9, 173–179 (2006).
Morin, X., Jaouen, F. & Durbec, P. Control of planar divisions by the G-protein regulator LGN maintains progenitors in the chick neuroepithelium. Nat Neurosci 10, 1440–1448 (2007).
Ebert, M. S., Neilson, J. R. & Sharp, P. A. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat Methods 4, 721–726 (2007).
Tamamaki, N. et al. Green fluorescent protein expression and colocalization with calretinin, parvalbumin, and somatostatin in the GAD67-GFP knock-in mouse. J Comp Neurol 467, 60–79 (2003).
Saeed, A. I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003).
Acknowledgements
We thank Aude Barani from the Precym platform of Centre d’Océanologie de Marseille for FACS, Xavier Morin for plasmids, Lucie Castelein for technical help and members of the Cremer and Barbry groups for discussion and support. The expertise of members of the Functional Genomics Platform of Nice Sophia Antipolis, supported by the Program “Investissements d’Avenir” managed by the Agence National de la Recherche (ANR-10-09-INBS), is greatly acknowledged. We thank the PiCSL-FBI platform and the animal facilities (IBDM, AMU-Marseille). This work was supported by funds from the Agence National de la Recherche (ANR, FORDOPA, AtMir), Fondation pour la Recherche Médicale (Label Equipe FRM), Fondation de France and the European Commission (Marie-Curie: ITN AXREGEN and IAPP DopaNew) to HC and AB, Association France Parkinson to PF. The work of the DH-ES team was supported by KU Leuven (GOA-11/012), Fund for Scientific Research-Flanders (FWO-V G.0954.11 and G.0782.14), Interuniversity Attraction Poles from the Belgian Science Policy Office (Belspo IUAP-VII/07), InfraMouse Hercules Foundation (ZW09-03) and Erasmus MC.
Author information
Authors and Affiliations
Contributions
C.B. designed and performed experiments, analyzed data, wrote the paper; P.F. designed and performed experiments; Eve Seuntjens designed and performed experiments; S.B., C.N., A.d.C., V.M. and K.L. performed experiments; U.B., A.B., D.H. and P.B. provided unpublished tools and techniques; Elke Stappers provided unpublished tools, designed and performed experiments; H.C. designed experiments, analyzed data, wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Beclin, C., Follert, P., Stappers, E. et al. miR-200 family controls late steps of postnatal forebrain neurogenesis via Zeb2 inhibition. Sci Rep 6, 35729 (2016). https://doi.org/10.1038/srep35729
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35729
- Springer Nature Limited
This article is cited by
-
The genomic response of human granulosa cells (KGN) to melatonin and specific agonists/antagonists to the melatonin receptors
Scientific Reports (2022)
-
Analyses of the possible anti-tumor effect of yokukansan
Journal of Natural Medicines (2019)
-
FOXG1 Regulates PRKAR2B Transcriptionally and Posttranscriptionally via miR200 in the Adult Hippocampus
Molecular Neurobiology (2019)
-
Comparative studies of vertebrate iduronate 2-sulfatase (IDS) genes and proteins: evolution of A mammalian X-linked gene
3 Biotech (2017)