Abstract
Fungal diseases, such as powdery mildew and rusts, significantly affect the quality and yield of wheat. Pyramiding diverse types of resistance genes into cultivars represents the preferred strategy to combat these diseases. Moreover, achieving collaborative improvement between diseases resistance, abiotic stress, quality, and agronomic and yield traits is difficult in genetic breeding. In this study, the wheat cultivar, Guinong 29 (GN29), showed high resistance to powdery mildew and stripe rust at both seedling and adult plant stages, and was susceptible to leaf rust at the seedling stage but slow resistance at the adult-plant stage. Meanwhile, it has elite agronomic and yield traits, indicating promising coordination ability among multiple diseases resistance and other key breeding traits. To determine the genetic basis of these elite traits, GN29 was tested with 113 molecular markers for 98 genes associated with diseases resistance, stress tolerance, quality, and adaptability. The results indicated that two powdery mildew resistance (Pm) genes, Pm2 and Pm21, confirmed the outstanding resistance to powdery mildew through genetic analysis, marker detection, genomic in situ hybridization (GISH), non-denaturing fluorescence in situ hybridization (ND-FISH), and homology-based cloning; the stripe rust resistance (Yr) gene Yr26 and leaf rust resistance (Lr) genes Lr1 and Lr46 conferred the stripe rust and slow leaf rust resistance in GN29, respectively. Meanwhile, GN29 carries dwarfing genes Rht-B1b and Rht-D1a, vernalization genes vrn-A1, vrn-B1, vrn-D1, and vrn-B3, which were consistent with the phenotypic traits in dwarf characteristic and semi-winter property; carries genes Dreb1 and Ta-CRT for stress tolerance to drought, salinity, low temperature, and abscisic acid (ABA), suggesting that GN29 may also have elite stress-tolerance ability; and carries two low-molecular-weight glutenin subunit genes Glu-B3b and Glu-B3bef which contributed to high baking quality. This study not only elucidated the genetic basis of the elite traits in GN29 but also verified the capability for harmonious improvement in both multiple diseases resistance and other comprehensive traits, offering valuable information for breeding breakthrough-resistant cultivars.
Similar content being viewed by others
Introduction
Common wheat (Triticum aestivum L.), offering a staple food grain for 35% of the global population and supplying 20% of the caloric intake worldwide, stands as a pivotal agricultural crop in ensuring food supply and security. Multiple diseases seriously affected wheat safety production, including powdery mildew, stripe rust, and leaf rust2,3,4. To combat these diseases, the strategies encompassing cultivation management, chemical prevention, and host resistance have been employed, and the latter is preferred means owe to its high efficiency and environment-friendly characteristics5,6. Meanwhile, a systematic evaluation of multiple diseases resistance in those released wheat cultivars at the molecular and genetic levels is also important for their rational distribution in production and application in breeding7.
Among wheat diseases, powdery mildew caused by the biotrophic fungus Blumeria graminis f. sp. tritici (Bgt), stripe rust caused by Puccinia striiformis f. sp. tritici (Pst) and leaf rust caused by the Puccinia triticina Eriks. are the most widespread and damaging fungal diseases threatening wheat production worldwide2,3,4. To date, more than 100 formally designated Pm genes/alleles at 64 loci (Pm1-Pm69, Pm8 = Pm17, Pm18 = Pm1c, Pm22 = Pm1e, Pm23 = Pm4c, Pm31 = Pm21)8, 83 Yr genes6. If the target genes have not been cloned, closely linked markers can also facilitate the detection of these genes14. Recently, many elite genes have been identified and confirmed in a large number of wheat genotypes through marker detection, including Rht-B1b and Rht-D1b15, Yr1516, Pm217, Pm2118, Pm1218, PmV18, Pm242, and Vp1-B19,20. Based on these information, the genetic basis of these genotypes were clarified, and better cooperative models between different genes were revealed.
Guinong 29 (GN29) is a wheat cultivar with collaborative improvement between high resistance to multiple wheat diseases and elite comprehensive performance. To dissect the genetic basis of multiple diseases resistance and other key breeding traits and discuss their cooperative improvement capability, the following aspects were carried out in the present study: (i) evaluate its powdery mildew, stripe rust, and leaf rust resistance at both seedling and adult plant stages; (ii) investigate its agronomic and yield performance at different wheat production regions; and (iii) dissect the genetic basis for the multiple diseases resistance and other key breeding traits using genetic analysis, molecular detection, and/or homology-based cloning.
Materials and methods
Plant materials
The wheat cultivar GN29 was developed from a cross between the wheat cultivar Guinong 13 (GN13) and the wheat breeding line Guinong 21 (GN21) using marker-assisted selection (MAS) by Guizhou University and Guizhou Sub-center of the National Wheat Improvement Center and released in 2014. The wheat cultivar **’an 9 (PA9) was susceptible to powdery mildew and stripe rust and used as the susceptible parent to cross with GN29 to obtain the F1, F2, and F2:3 populations for genetic and lineage analysis of Pm and Yr genes in GN29. The wheat cultivar Mingxian 169 (MX169) was susceptible to powdery mildew, stripe rust, and leaf rust and used as a susceptible check in multiple diseases resistance evaluation. Nineteen and fourteen wheat donors carrying known Pm genes and Yr genes, respectively, served as positive controls in the molecular marker detection and/or homology-based cloning experiments (Supplementary Table S1). Donors of these Pm and Yr genes were provided by Prof. Hongxing Xu, Henan University, Kaifeng, China and Prof. Caixia Lan, Huazhong Agricultural University, Wuhan, China, respectively.
Resistance assessment to multiple wheat diseases
To assess the powdery mildew resistance, 31 single-spore-derived Bgt isolates with different virulence spectra, provided by Prof. Yilin Zhou, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, and Prof. Hongxing Xu, Henan University, were used to test the seedling reaction patterns of GN29 using MX169 as the susceptible control and nine resistant donors with known Pm genes as the resistant controls. Five seeds of each genotype were planted in trays (54 × 28 × 4.2 cm) with 128 cells (3.2 × 3.2 × 4.2 cm). When the seedlings grown to the two-leaf stage, they were inoculated with the fresh conidiospores previously developed on the MX169 seedlings. Seedlings in different trays were inoculated with the 31 Bgt isolates separately, and each tray was covered with a glass shroud to avoid cross-infection between different isolates. Several resistance stocks with documented Pm genes in production or with high resistance were used as resistant controls and MX169 was used as susceptible control. When the pustules were fully developed on the first leaves of MX169 seedlings, approximately 14–15 days after inoculation, infection types (ITs) for each plant were scored based on a 0–4 scale standard, of which ITs 0–2 were regarded as resistant and ITs 3 and 4 as susceptible21. All tests were repeated thrice to ensure data reliability.
At the adult stage, GN29 was inoculated with a mixture of all the Bgt isolates used in the seedling stage for three consecutive years (2018 to 2020) at Yantai University, Yantai City, Shandong Province, China (121.39′ E, 37.52′ N). Sowing and inoculation methods were referred to our proven technique system22. Disease reaction at the adult stage was scored on a 0–9 scale, of which 0–4 was considered as resistant and 5–9 as susceptible21. Each plant was assessed twice.
For stripe rust resistance assessment, three Pst isolates, CYR32, CYR33, and CYR34 with different virulence spectra were used to test the seedling reaction patterns of GN29 using MX169 as the susceptible control. Assessments of adult plant stripe rust responses were conducted at Yantai University using a mixture of CYR32, CYR33, and CYR34. The sowing and inoculation methods were referred to the reported procedure23. The ITs at the seedling and adult plant stages were both scored based on a scale of 0–9, of which ITs ranging from 0 to 6 were classified as resistant, whereas ITs 7–9 as susceptible24.
For the leaf rust resistance assessment, mixed P. triticina pathotypes collected in Yantai, China were used to test the seedling and adult-stage reaction patterns of GN29 using MX169 as the susceptible control. The sowing and inoculation methods were referred to the reported procedure18. Infection types were scored according to the Stakman scale with moderate modification25.
Assessment of agronomic and yield performance
GN29 were planted at Guiyang city, Guizhou Province (26.57 N, 106.71 E), China, area available for popularization and Langfang City, Hebei Province (39.53 N, 116.72 E), China, in a randomized complete block design with three replicates. The wheat cultivars Guinong 19 (GN19) and Yannong 999 (YN999) were used as controls in Guizhou and Langfang, respectively. Each cultivar was planted as a plot with three rows (length: 1.5 m; distance between rows: 0.25 m) and 30 seeds per row. Three plants in the middle of the two internal rows were sampled to evaluate the plant height (PH), spike numbers per plant (SNPP), spikelet numbers per spike (SNS), sterile spikelet numbers per spike (SSNS), kernel numbers per spike (KNS), and thousand-kernel weight (TKW).
Genetic analysis, molecular marker detection, and homology-based cloning of the Pm genes
To determine the inheritance of powdery mildew resistance in GN29 at the seedling stage, two Bgt isolates E09 (prevalent) and E18 (hypertoxic) were used to inoculate GN29 and PA9 and their F1 hybrids, F2 population, and F2:3 families (30 seeds per F2:3 family were selected) at the one-leaf stage. After phenoty**, goodness-of-fit was analyzed using the chi-square (χ2) test to investigate deviations in the observed phenotypic data of the F2 populations and F2:3 families from the theoretically expected segregation ratios.
Total genomic DNA was isolated using the cetyltrimethylammonium bromide (CTAB) method from young leaves of the wheat seedlings27. To detect the Pm genes in GN29, 39 diagnostic/linked markers of 31 known Pm genes were used to genotype GN29, PA9, and 19 wheat donors with known Pm genes (Supplementary Tables S1 and S2). Polymorphic markers were genotyped in the corresponding F2:3 families.
PCR amplification and visualization were performed as described in our lab27. PCR amplification was carried out in a 10 μL volume system, including 5 μL 2 × Taq Master Mix (Vazyme, China), 1 μL 50 ng/μL template DNA and 0.5 μL 10 μM/μL primers. The PCR amplification condition was set as follows: pre-denaturation at 94 °C for 5 min followed by 36 cycles of 94 °C for 30 s, 50–65 °C (depending on the specific primers) for 40 s, 72 °C for 40–120 s (depending on the target bands), finally, extension at 72 °C for 10 min and preservation at 25 °C. The PCR products were then separated on 8% non-denaturing polyacrylamide gels with a 29:1 ratio of acrylamide to bis-acrylamide or 1.5% agarose gel based on the size of the target bands28,29.
After confirming the presence of Pm genes in GN29, total RNA from the young leaves of GN29 were extracted using the Spectrum Plant Total RNA kit (Sigma-Aldrich, Shanghai, China) following the manufacturer’s recommendations. The RNA samples were quantified by measuring the absorbance at 260 and 280 nm using a NanoDrop 1000 spectrophotometer (Thermo Scientific, Shanghai, China). High-quality RNA was treated with Promega DNase I for cDNA synthesis using Invitrogen SuperScript II reverse transcriptase, according to the manufacturer’s guidelines. Based on the reports of the cloning of Pm230 and Pm2131, the full length of the homologous sequences of Pm2 and Pm21 were isolated. After obtaining the coding sequence (CDS) of Pm2 and Pm21, they were sequenced using Sanger sequencing and compared with those of Pm230 and Pm2131.
Cytogenetic analysis
Genomic in situ hybridization (GISH) was firstly performed to detect D. villosum chromatin in the GN29. Mitotic chromosomes of the root tip cells of GN29 were prepared and observed, as previously described32. The genomic DNA of D. villosum was labeled with fluorescein-12-dUTP as a probe to detect chromosomal fragments for GISH. After hybridization with probes, the chromosomes were counterstained with propidium iodide (PI) and mounted on Vectashield (Roche Co., Burlingame, CA, USA). Signals were examined under an Olympus BX60 epifluorescence microscope (Olympus Co., Tokyo, Japan).
To clearly determine the chromosome composition of GN29, we also performed non-denaturing fluorescence in situ hybridization (ND-FISH) to analyze mitotic chromosomes of the root tip cells. The probes in this study were Oligo-pSc199.2-1 (green) and Oligo-pTa535-1 (red), and they were distributed as 5′ end-labeled with 6-carboxyfluorescein (FAM) and 6-carboxytetramethylrhodamine (TAMRA).
Molecular marker detection of the Yr and Lr genes
To determine the presence of Yr genes in GN29, 17 diagnostic/linked markers for known Yr genes were used to test GN29 and PA9, using 14 resistant donors with known Yr genes as controls (Supplementary Table S2). If the polymorphic band(s) of one Yr gene were detected in GN29 and not PA9, this Yr gene would most likely exist in GN29. To confirm this result, the polymorphic markers were also used to genotype the segregated populations of GN29 and PA9.
To detect the presence of Lr genes in GN29, a similar but simplified procedure was performed using 12 diagnostic/linked markers of known Lr genes by comparing the polymorphic band(s) in GN29 and PA9 but not by genoty** the segregation population (Supplementary Table S2). PCR amplification and product visualization were performed as described above27,28,29.
Molecular marker detection of other key breeding traits
To determine the presence of other key breeding traits in GN29, 45 diagnostic/linked markers closely linked to wheat adaptability, PH, stress tolerance, and quality were used to test GN29, including seven markers for seven vernalization genes (Vrn-A1c, vrn-A1, Vrn-B1, vrn-B1, Vrn-D1, vrn-D1, Vrn-B3, and vrn-B3), five markers for five dwarfing genes (Rht-B1b, Rht-B1a, Rht-D1a, Rht-D1b, and Rh8), two markers for two drought tolerance genes (Dreb1 and Ta-CRT), and four markers for seven pre-harvest sprouting resistance genes (TaAFP-Bb, TaAFP-Ba, Vp-1Ba, Vp-1Bb, Vp-1Bc, Vp-1Bf and TaPHS1) (Supplementary Table S2). PCR amplification and product visualization were performed as described above27,28,29.
Results
Agronomic and yield performance of GN29
When GN29 was planted in Guizhou Province, it showed comprehensively excellent performance for the investigated traits, including PH, SNPP, SNS, SSNS, KNS, and TKW (Table 1), no obvious disadvantages were detected. Compared with the famous wheat cultivar GN19, GN29 still has significant advantages in terms of SNPP, SSNS, and TKW. To investigate the adaptation of GN29 in other wheat production region, GN29 was also surveyed in Langfang City, Hebei Province. Compared to the performance in Guizhou Province, GN29 significantly increased SNPP but decreased TKW. Significant decrease in the PH was also observed (Fig. 1, Table 1). Although significant changes in agronomic and yield performance occurred in other distinct agroecological area, GN29 still showed better adaptation. Even compared to the famous wheat cultivar YN999, GN29 still showed satisfactory agronomic and yield performance in the Langfang region (Fig. 1, Table 1).
Evaluation and inheritance of the powdery mildew resistance in GN29
When tested with the Bgt isolate E09, GN29 showed hypersensitivity on the first leaves and can be regarded as immune with IT 0;, whereas PA9 showed abundant sporulation with > 80% of the first leaves covered with aerial hyphae and hence as highly susceptible with IT 4. The F1 plants of GN29 × PA9 showed a similar reaction pattern to E09 as that to GN29 with an IT 0;, suggesting that the Pm gene(s) in GN29 displayed dominant inheritance. The F2 population fitted the theoretical ratio of 15:1 for the segregation model of the two dominant genes (Table 2).
Another Bgt isolate E18 was also used to inoculate GN29, PA9, and their derived F1 plants, F2 populations, and F2:3 families. Interestingly, a dominant monogenic segregation model was clearly observed using this highly virulent Bgt isolate, suggesting that one of the two Pm genes in GN29 was defeated by this Bgt isolate and the remaining Pm gene displayed dominant monogenic inheritance (Table 2).
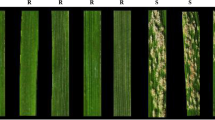
To further assess seedling resistance to powdery mildew in GN29, it was inoculated with 31 Bgt isolates using several Pm gene donors in production or with a high breeding value as controls. The results showed that GN29 was immune to all the tested Bgt isolates (Fig. 2, Table 3). Compared to the other tested Pm genes, GN29 had broad spectrum resistance. At the adult stage, GN29 was also immune to the Bgt mixture in three consecutive years. Therefore, GN29 showed elite powdery mildew resistance at the whole growth stage.
Molecular detection and homology-based cloning of the Pm genes in GN29
To identify the Pm genes in GN29, 39 diagnostic/linked markers for 31 known Pm genes were selected to test GN29 and PA9 (Table 4). Only the diagnostic markers Pm2b-map-3 for Pm2 and MBH1 for Pm21 amplified the targeted bands of Pm2 and Pm21, respectively, in GN29 but not in PA9. To further verify the presence of Pm2 and Pm21, the diagnostic marker MBH1 of Pm21 was used to genotype the F2:3 families phenotyped by Bgt isolate E18. As expected, MBH1 is co-segregated with the phenotype, suggesting the presence of Pm21 (Fig. 3). To confirm the existence pattern of Pm21, GISH analysis was carried out and showed that GN29 had a pair of alien chromosome arms of D. villosum (Fig. 4A). ND-FISH analysis further showed that the alien chromosome arms were 6VS in GN29 (Fig. 4B). The diagnostic marker Pm2b-map-3 of Pm2 was also used to genotype the susceptible plants of the F2 population phenotyped by the Bgt isolate E09, and it co-segregated with the phenotype, suggesting the presence of Pm2. To clarify the allelic types of Pm2 and Pm21 in GN29, we cloned their homologous sequences. Following sequence alignment, the haplotypes in GN29 were confirmed as Pm2a and Pm21.
Amplification pattern of the diagnostic marker mbh1 of Pm21 in genoty** Guinong 29 (GN29), **’an 9 (PA9) and random selected F2:3 families of GN29 × PA9 at the seedling stage. Lane M, pUC19 Msp I; lanes 1–2: parents GN29 and PA9; lanes 3–7: homozygous resistant F2:3 families; lanes 8–12, homozygous susceptible F2:3 families; lanes 13–17: heterozygous F2:3 families. The white arrows indicate the polymorphic bands in GN29.
Genomic in situ hybridization (GISH) (A) and nondenaturing FISH (ND-FISH) (B) analysis of Guinong 29 (GN29). (A) GISH analysis on the chromosome constitution in GN29 shows light blue hybridization signals evenly distributed by using Dasypyrum villosum genomic DNA as a probe, and the wheat chromosomes were counterstained with 4, 6-diamidino-2-phenylindole (DAPI) (dark blue). (B) ND-FISH analysis on the chromosome constitution of Guinong 29 shows no specific hybridization bands on chromosomes of the D. villosum in GN29. White arrows note the pair of added 6VS chromosomes.
Evaluation and identification of stripe rust and leaf rust resistance in GN29
When tested with Pst isolates CYR32, CYR33, and CYR34 at the seedling stage, GN29 showed high resistance with IT 1. When tested with a mixture of these three Pst isolates at the adult plant stage, GN29 also showed high resistance ranging from IT1-2. For the leaf rust resistance assessment, GN29 was highly susceptible to the mixed Pt races; however, it showed slow reactance at the adult plant stage, suggesting that the slow leaf rust resistance gene may be involved in GN29.
To investigate the Yr gene(s) in GN29, 17 diagnostic/linked molecular markers for 14 known Yr genes were used to test GN29 (Table 4). The results showed that only the Yr26-linked marker WE173 could amplify the target band, suggesting that GN29 was most likely to carry Yr26. To further verify the presence of Yr26, WE173 was used to genotype the F2:3 families phenotyped by isolate CRY34. The result showed that WE173 was co-segregated with phenotypes, further confirming the presence of Yr26 (Fig. 5). A simple and similar procedure was also carried out to detect the Lr gene(s) in GN29 by analyzing the polymorphic band(s) in GN29 but not by genoty** the segregation population, suggesting that Lr1 and Lr46 may exist in GN29 (Supplementary Table S2).
Amplification pattern of WE173 linked to Yr26 in genoty** Guinong 29 (GN29), **’an 9 (PA9) and random selected F2:3 families of GN29 × PA9 at the seedling stage. Lane M, pUC19 Msp I; lanes 1–2: parents GN29 and PA9; lanes 3–7: homozygous resistant F2:3 families; lanes 8–12, homozygous susceptible F2:3 families; lanes 13–17: heterozygous F2:3 families. The white arrows indicate the polymorphic bands in GN29.
Molecular identification of drought tolerance and preharvest sprouting resistance in GN29
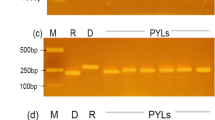
To investigate the drought tolerance and preharvest sprouting resistance genes in GN29, six markers linked to two main drought tolerance genes and three markers linked to three main preharvest sprouting resistance genes were used to detect GN29 (Table 4). The results indicated that markers P18, P20, P21, P22, and P25 associated with the drought tolerance gene Dreb1, and DF/DR linked to Ta-CRT, successfully amplified target bands, suggesting that GN29 was most likely to carry Dreb1 and Ta-CRT, implying the potential drought tolerance of GN29 (Supplementary Table S2). However, marker detection results also indicated that GN29 had none of the three main preharvest sprouting resistance genes, TaAFP-Bb/TaAFP-Ba, Vp-1Bb/Vp-1Ba/Vp-1Bc, and Vp-1Ba/Vp-1Bf, implying potential risks in pre-harvest sprouting (Supplementary Table S2).
Molecular identification of vernalization and dwarfing genes in GN29
To investigate the vernalization and dwarfing genes in GN29, nine markers linked to eight vernalization genes and five markers linked to five dwarfing genes were used to detect GN29. The results indicated that the markers BF-WR1 linked to Rht-B1b, DF-MR2 linked to Rht-D1a and gwm261 linked to Rht8 could amplify the target bands, suggesting that GN29 most likely carries the dwarfing genes Rht-B1b, Rht-D1a, and Rht8 (Table 4, Supplementary Table S2). The PH of GN29 planted in Guizhou Province did not correlate with the presence of Rht-B1b, Rht-D1a, and Rht8; however, the PH in Hebei Province was mostly affected by their presence. For the detection of vernalization genes, GN29 had four of the seven tested vernalization genes vrn-A1, Vrn-B1, vrn-D1, and Vrn-B3, hence, GN29 could be considered a semi-winter cultivar. The better adaptation in Guizhou and Hebei Provinces may be related to this factor.
Molecular identification of quality-related genes in GN29
To investigate the quality-related genes in GN29, 22 markers linked to 22 glutenin subunit genes were used to detect GN29. The result indicated that the markers SB2 linked to Glu-B3b and SB10 linked to Glu-B3bef could amplify the target bands, suggesting that GN29 was most likely to carry two low-molecular-weight glutenin subunit genes, Glu-B3b and Glu-B3bef, which contribute to the malleability of the dough and food processing quality (Table 4, Supplementary Table S2). We noticed that the elite high-molecular-weight glutenin subunit genes Dx5 and Dx10 were absent in GN29, which may affect the gluten strength of GN29 (Table 4, Supplementary Table S2).
Discussion
Wheat breeding involves the integration of elite traits from diverse donors into a unified genetic background. In this process, harmonious improvement between different traits is critical, particularly between multiple diseases resistance and other comprehensive traits13. For instance, it is difficult to pyramid large mumble of elite traits between multiple diseases resistance and stress tolerance, adaptation, quality, and high yield and better express all of them in a harmonious pattern36,37,38. A similar situation occurs for Pm21, which is currently the most effective Pm gene and has been used in at least 20 cultivars39. Given the high value of Pm2 and Pm21, extending their operation lifespan during wheat production is imperative. Gene pyramiding is a promising strategy for develo** durable resistance. Fortunately, Pm2 and Pm21 are pyramided in GN29 and achieve better collaboration, which is expected to be interdependent and can build durable resistance against continuous Bgt variations. In addition to powdery mildew resistance, GN29 also pyramided one Yr gene, Yr26, and two different kinds of Lr genes, Lr1 and Lr46. Among them, Yr26 is an elite resistance gene with high- and broad-spectrum resistance to stripe rust throughout the whole growth stage and has been used in production for many years40; Lr1 is a frequently used resistance gene with leaf rust resistance throughout the whole growth stage41; Lr46 is a resistance gene with slow resistance to leaf rust, and this locus is also resistant to stripe rust (Yr29)42, powdery mildew (Pm39)43, and stem rust (Sr58)44 as a multiple resistant locus. Therefore, it is rare to pyramid so many resistance genes into a single cultivar, and meanwhile realize better collaboration.
Beyond the five different resistance genes, GN29 also pyramided two genes conferring drought tolerance, which implies its potential drought resistance ability and may be suitable for extension and application in arid and water-scarce regions. The Dreb gene is an important gene involved in abiotic stress tolerance in wheat production, including tolerance to drought, salinity, low temperature, and ABA45. The combination of Dreb and the five resistance genes further suggests that GN29 may possess a cooperative ability to improve both biotic and abiotic stresses.
GN29 also pyramided two low-molecular-weight glutenin subunit genes, Glu-B3b and Glu-B3bef, but no high-molecular-weight glutenin subunit genes, particularly the elite Dx5, were detected in GN29. Therefore, GN29 is regarded as a low-gluten cultivar and valuable for producing low-gluten flour, which is mainly used to make biscuits.
Given the extensive pyramiding of genes related to biotic and abiotic stress, as well as quality in GN29, an important question arises regarding the impact of these genes on agricultural and yield performance. From the agricultural and yield analyses, we found that GN29 maintained elite agricultural and yield performance and no obvious defects were observed. Meanwhile, GN29 can be considered a semi-winter cultivar based on the detection of vernalization genes46,47,48, suggesting that it is suitable for extension and application in southwestern wheat production regions, such as Guizhou, Yunnan, and Sichuang provinces of China, and in the south-central region of the northern winter wheat region. Agricultural and yield performances in distinct regions also indicated that GN29 has better adaptability, whether in Guizhou Province in the south or Hebei Province in the north. In addition, differences in plant height were obvious in different regions. Although three dwarfing genes were identified in GN29, they could not be adequately displayed in the Guizhou Province where GN29 has been selected and popularized. In Hebei Province, the three dwarfing genes were fully displayed, which may be related to different ecoclimatic conditions.
Conclusion
The wheat cultivar GN29 showed promising coordination between multiple diseases resistance and other key breeding traits. To determine the genetic foundation of these elite traits, GN29 was tested with 113 molecular markers for 98 target genes associated with diseases resistance, stress tolerance, quality, and adaptability. Several key genes were confirmed using genetic analysis, marker detection, and/or homology-based cloning. This study not only dissect the genetic basis of GN29 but also verified the harmonious improvement ability across multiple diseases resistance and other key breeding traits, which can provide elite gene resources and references for gene pyramiding during breeding practices.
Data availability
All data generated or analyzed during this study are included in this published article. We declare that the plant materials used for our study were provided by Chinese Crop Germplasm Resource Bank (Bei**g, China) as seed materials and could be also purchased by commercial channel. We did not use endangered plant species for the experiments. Experimental research and field studies on plants (either cultivated or wild), including the collection of plant material, complied with relevant institutional, national, and international guidelines and legislation.
References
Hickey, L. T. et al. Breeding crops to feed 10 billion. Nat. Biotechnol. 37, 744–754. https://doi.org/10.1038/s41587-019-0152-9 (2009).
Lu, P. et al. A rare gain of function mutation in a wheat tandem kinase confers resistance to powdery mildew. Nat. Commun. 11, 680. https://doi.org/10.1038/s41467-020-14294-0 (2020).
Yu, G. T. et al. The wheat stem rust resistance gene Sr43 encodes an unusual protein kinase. Nat. Genet. 55, 921–926. https://doi.org/10.1038/s41588-023-01402-1 (2023).
Wang, Y. J. et al. An unusual tandem kinase fusion protein confers leaf rust resistance in wheat. Nat. Genet. 55, 914–920. https://doi.org/10.1038/s41588-023-01401-2 (2023).
Wu, Y. N. et al. Characterization of PmDGM conferring powdery mildew resistance in Chinese wheat landrace Duanganmang. Plant Dis. 105, 3127–3133. https://doi.org/10.1094/pdis-12-20-2719-re (2021).
Zhang, P. P. et al. Genome-wide association map** of leaf rust and stripe rust resistance in wheat accessions using the 90K SNP array. Theor. Appl. Genet. 134, 1233–1251. https://doi.org/10.1007/s00122-021-03769-3 (2021).
Miedaner, T. et al. Molecular tracking of multiple disease resistance in a winter wheat diversity panel. Theor. Appl. Genet. 133, 419–431. https://doi.org/10.1007/s00122-019-03472-4 (2020).
Li, Y. H. et al. Dissection of a rapidly evolving wheat resistance gene cluster by long-read genome sequencing accelerated the cloning of Pm69. Plant Commun. 5, 100646. https://doi.org/10.1016/j.xplc.2023.100646 (2023).
McIntosh, R. A., Dubcovsky, J., Rogers, W. J., **a, X. C. & Raupp, W. J. Catalogue of gene symbols for wheat 2020 supplement. https://wheat.pw.usda.gov/GG3/WGC (2020)
Kumar, S. et al. Lr80: A new and widely effective source of leaf rust resistance of wheat for enhancing diversity of resistance among modern cultivars. Theor. Appl. Genet. 134, 849–858. https://doi.org/10.1007/s00122-020-03735-5 (2021).
Zhu, S. Y. et al. Pm12 and Pm21 are orthologous genes originated from different wild relatives of wheat and show evolutionary conservation but divergent powdery mildew resistance. Plant Commun. 4, 100472. https://doi.org/10.1016/j.xplc.2022.100472 (2023).
Koller, T., Brunner, S., Herren, G., Hurni, S. & Keller, B. Pyramiding of transgenic Pm3 alleles in wheat results in improved powdery mildew resistance in the field. Theor. Appl. Genet. 131, 861–871. https://doi.org/10.1007/s00122-017-3043-9 (2018).
Han, G. H. et al. Development and identification of a wheat-rye breeding line for harmonious improvement between powdery mildew resistance and high yield potential. Plant Dis. https://doi.org/10.1094/pdis-12-22-2817-re (2023).
Wu, L. R. et al. Genetic dissection of the powdery mildew resistance in wheat breeding line LS5082 using BSR-Seq. Crop J. 10, 1120–1130. https://doi.org/10.1016/j.cj.2021.12.008 (2022).
Ellis, M., Spielmeyer, W., Gale, K., Rebetzke, G. & Richards, R. “Perfect” markers for the Rht-B1b and Rht-D1b dwarfing genes in wheat. Theor. Appl. Genet. 105, 1038–1042. https://doi.org/10.1007/s00122-002-1048-4 (2002).
Peng, J. H. et al. High-density molecular map of chromosome region harboring stripe-rust resistance genes YrH52 and Yr15 derived from wild emmer wheat Triticum dicoccoides. Genetica 109, 199–210. https://doi.org/10.1023/a:1017573726512 (2000).
**, Y. L. et al. Identification of resistant germplasm and detection of genes for resistance to powdery mildew and leaf rust from 2,978 wheat accessions. Plant Dis. 105, 3900–3908. https://doi.org/10.1094/pdis-03-21-0532-re (2021).
Zhang, X. et al. Diagnostic KASP markers of wheat broad-spectrum powdery mildew resistance genes Pm21, PmV, and Pm12 developed for high throughput marker-assisted selection. Plant Dis. 105, 2844–2850. https://doi.org/10.1094/pdis-02-21-0308-re (2021).
Yang, Y. et al. Development and validation of a Viviparous-1 STS marker for pre-harvest sprouting tolerance in Chinese wheats. Theor. Appl. Genet. 115, 971–980. https://doi.org/10.1007/s00122-007-0624-z (2007).
Chang, C. et al. Validating a novel allele of viviparous-1 (Vp-1Bf) associated with high seed dormancy of Chinese wheat landrace Wanxianbaimaizi. Mol. Breed. 25, 517–525. https://doi.org/10.1007/s11032-009-9350-3 (2010).
Si, Q. M., Zhang, X. X., Duan, X. Y. & Sheng, B. Q. Identification of isolates of Blumeria graminis f. sp. tritici. Sci. Agric. Sin. 20, 64–70 (1987).
Mu, Y. J. et al. Identification of the powdery mildew resistance gene in wheat breeding line Yannong 99102–06188 via bulked segregant exome capture sequencing. Front. Plant Sci. 13, 1005627. https://doi.org/10.3389/fpls.2022.1005627 (2022).
**, H. L. et al. Identification of a suppressor for the wheat stripe rust resistance gene Yr81 in Chinese wheat landrace Dahongpao. Theor. Appl. Genet. 136, 67. https://doi.org/10.1007/s00122-023-04347-5 (2023).
Line, R. F. & Qayoum, A. Virulence, aggressiveness, evolution, and distribution of races of Puccinia striiformis (the cause of stripe rust of wheat) in North America, 1968–1987. US Dep. Agric. Tech. Bull. 1788. https://api.semanticscholar.org/CorpusID:83215121 (1992).
Roelfs, A. P., Singh, R. P. & Saari, E. E. Rust diseases of wheat: concepts and methods of disease management. CIMMYT, Mexico, D. F. (1992).
Sharp, P. J., Kreis, M., Shewry, P. R. & Gale, M. D. Location of β-amylase sequences in wheat and its relatives. Theor. Appl. Genet. 75, 286–290. https://doi.org/10.1007/BF00303966 (1998).
Ma, P. T. et al. Characterization of a powdery mildew resistance gene in wheat breeding line 10V–2 and its application in marker-assisted selection. Plant Dis. 102, 925–931. https://doi.org/10.1094/pdis-02-17-0199-re (2018).
Santos, F. R., Pena, S. D. & Epplen, J. T. Genetic and population study of a Y-linked tetranucleotide repeat DNA polymorphism with a simple non-isotopic technique. Hum. Genet. 90, 655–656. https://doi.org/10.1007/bf00202486 (1993).
Gebrewahid, T. W., Zhang, P. P., Yao, Z. J., Li, Z. F. & Liu, D. Q. Identification of leaf rust resistance genes in bread wheat cultivars from Ethiopia. Plant Dis. 104, 2354–2361. https://doi.org/10.1094/pdis-12-19-2606-re (2020).
Sánchez-Martín, J. et al. Rapid gene isolation in barley and wheat by mutant chromosome sequencing. Genome Biol. 17, 221. https://doi.org/10.1186/s13059-016-1082-1 (2016).
He, H. G. et al. Pm21, Encoding a typical CC-NBS-LRR protein, confers broad-spectrum resistance to wheat powdery mildew disease. Mol. Plant 11, 879–882. https://doi.org/10.1016/j.molp.2018.03.004 (2018).
Wang, C. M. et al. Molecular cytogenetic characterization of a new T2BL·1RS wheat-rye chromosome translocation line resistant to stripe rust and powdery mildew. Plant Dis. 21, 419–432. https://doi.org/10.1094/PDIS-93-2-0124 (2009).
**ao, J. et al. Wheat genomic study for genetic improvement of traits in China. Sci. China Life Sci. 65, 1718–1775. https://doi.org/10.1007/s11427-022-2178-7 (2022).
Pugsley, A. T. & Carter, M. V. The resistance of twelve varieties of Triticum vulgare to Erysiphe graminis tritici. Aust. J. Biol. Sci. 6, 335–346. https://doi.org/10.1071/bi9530335 (1953).
Yu, Z. Y. et al. Mining of wheat Pm2 alleles for goal-oriented marker-assisted breeding. Front. Plant Sci. 13, 912589. https://doi.org/10.3389/fpls.2022.912589 (2022).
Ma, P. T. et al. Molecular map** of a new powdery mildew resistance gene Pm2b in chinese breeding line KM2939. Theor. Appl. Genet. 128, 613–622. https://doi.org/10.1007/s00122-015-2457-5 (2015).
Ma, P. T. et al. The gene PmYB confers broad-spectrum powdery mildew resistance in the multi-allelic Pm2 chromosome region of the Chinese wheat cultivar YingBo 700. Mol. Breed. 35, 124. https://doi.org/10.1007/s11032-015-0320-7 (2015).
Xu, H. X. et al. Molecular tagging of a new broad-spectrum powdery mildew resistance allele Pm2c in Chinese wheat landrace niaomai. Theor. Appl. Genet. 128, 2077–2084. https://doi.org/10.1007/s00122-015-2568-z (2015).
Bie, T. D. et al. Development and characterization of marker MBH1 simultaneously tagging genes Pm21 and PmV conferring resistance to powdery mildew in wheat. Mol. Breed. 35, 1–8. https://doi.org/10.1007/s11032-015-0385-3 (2015).
McIntosh, R., Mu, J. M., Han, D. J. & Kang, Z. S. Wheat stripe rust resistance gene Yr24/Yr26: A retrospective review. Crop J. 6, 3–11. https://doi.org/10.1016/j.cj.2018.02.001 (2018).
Liu, Y. B., Takele, W. G., Zhang, P. P., Li, Z. F. & Liu, D. Q. Identification of leaf rust resistance genes in common wheat varieties from China and foreign countries. J. Integr. Agric. 20, 1302–1313. https://doi.org/10.1016/S2095-3119(20)63371-8 (2021).
William, M., Singh, R. P., Huerta-Espino, J., Islas, S. O. & Hoisington, D. Molecular marker map** of leaf rust resistance gene Lr46 and its association with stripe rust resistance gene Yr29 in wheat. Phytopathology 93, 153–159. https://doi.org/10.1094/phyto.2003.93.2.153 (2003).
Lillemo, M. et al. The adult plant rust resistance loci Lr34/Yr18 and Lr46/Yr29 are important determinants of partial resistance to powdery mildew in bread wheat line Saar. Theor. Appl. Genet. 116, 1155–1166. https://doi.org/10.1007/s00122-008-0743-1 (2008).
Jighly, A. et al. Genomic regions conferring resistance to multiple fungal pathogens in synthetic hexaploid wheat. Mol. Breed. 36, 127. https://doi.org/10.1007/s11032-016-0541-4 (2016).
Wei, B. et al. Dreb1 genes in wheat (Triticum aestivum L.): development of functional markers and gene map** based on SNPs. Mol. Breed. 23, 13–22. https://doi.org/10.1007/s11032-008-9209-z (2009).
Fu, D. L. et al. Large deletions within the first intron in VRN-1 are associated with spring growth habit in barley and wheat. Mol. Genet. Genomics 273, 54–65. https://doi.org/10.1007/s00438-004-1095-4 (2005).
Yan, L. L. et al. Allelic variation at the VRN-1 promoter region in polyploid wheat. Theor. Appl. Genet. 109, 1677–1686. https://doi.org/10.1007/s00122-004-1796-4 (2004).
Yan, L., Fu, D., Li, C. & Dubcovsky, J. The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc. Natl. Acad. Sci. 103, 19581–19586. https://doi.org/10.1073/pnas.0607142103 (2006).
Hewitt, T. et al. A highly differentiated region of wheat chromosome 7AL encodes a Pm1a immune receptor that recognizes its corresponding AvrPm1a effector from Blumeria graminis. New Phytol. 229, 2812–2826. https://doi.org/10.1111/nph.17075 (2021).
Sánchez-Martín, J. et al. Wheat Pm4 resistance to powdery mildew is controlled by alternative splice variants encoding chimeric proteins. Nat. Plants 7, 327–341. https://doi.org/10.1038/s41477-021-00869-2 (2021).
**e, J. Z. et al. A rare single nucleotide variant in Pm5e confers powdery mildew resistance in common wheat. New Phytol. 228, 1011–1026. https://doi.org/10.1111/nph.16762 (2020).
Wan, W. T. et al. Fine map** of wheat powdery mildew resistance gene Pm6 using 2B/2G homoeologous recombinants induced by the ph1b mutant. Theor. Appl. Genet. 133, 1265–1275. https://doi.org/10.1007/s00122-020-03546-8 (2020).
Hurni, S. et al. Rye Pm8 and wheat Pm3 are orthologous genes and show evolutionary conservation of resistance function against powdery mildew. Plant J. 76, 957–969. https://doi.org/10.1111/tpj.12345 (2013).
Chen, X. M. et al. Chromosomal location of powdery mildew resistance gene Pm16 in wheat using ssr marker analysis. Plant Breed. 124, 225–228. https://doi.org/10.1111/j.1439-0523.2005.01094.x (2010).
Mohler, V., Hsam, S. L. K., Zeller, F. J. & Wenzel, G. An STS marker distinguishing the rye-derived powdery mildew resistance alleles at the Pm8/Pm17 locus of common wheat. Plant Breed. 120, 448–450. https://doi.org/10.1046/j.1439-0523.2001.00622.x (2010).
Xue, F. et al. Molecular map** of a powdery mildew resistance gene in common wheat landrace Baihulu and its allelism with Pm24. Theor. Appl. Genet. 125, 1425–1432. https://doi.org/10.1007/s00122-012-1923-6 (2012).
Zhu, Z. D., Zhou, R. H., Kong, X. Y., Dong, Y. C. & Jia, J. Z. Microsatellite markers linked to 2 powdery mildew resistance genes introgressed from Triticum carthlicum accession PS5 into common wheat. Genome 48, 585–590. https://doi.org/10.1139/g05-016 (2005).
Miranda, L. M., Murphy, J. P., Marshall, D. & Leath, S. Pm34: A new powdery mildew resistance gene transferred from Aegilops tauschii Coss. to common wheat (Triticum aestivum L). Theor. Appl. Genet. 113, 1497–1504. https://doi.org/10.1007/s00122-006-0397-9 (2006).
Miranda, L. M., Murphy, J. P., Marshall, D., Cowger, C. & Leath, S. Chromosomal location of Pm35, a novel Aegilops tauschii derived powdery mildew resistance gene introgressed into common wheat (Triticum aestivum L.). Theor. Appl. Genet. 14, 1451–1456. https://doi.org/10.1007/s00122-007-0530-4 (2007).
Lagudah, E. S. et al. Molecular genetic characterization of the Lr34/Yr18 slow rusting resistance gene region in wheat. Theor. Appl. Genet. 114, 21–30. https://doi.org/10.1007/s00122-006-0406-z (2006).
Cobo, N., Wanjugi, H., Lagudah, E. & Dubcovsky, J. A high-resolution map of wheat QYr.ucw-1BL, an adult plant stripe rust resistance locus in the same chromosomal region as Yr29. Plant Genome 12, 180055. https://doi.org/10.3835/plantgenome2018.08.0055 (2019).
Li, M. M. et al. A CNL protein in wild emmer wheat confers powdery mildew resistance. New Phytol. 228, 1027–1037. https://doi.org/10.1111/nph.16761 (2020).
Hua, W. et al. Identification and genetic map** of Pm42, a new recessive wheat powdery resistance gene derived from wild emmer (Triticum turgidum var. discoccoides). Theor. Appl. Genet. 119, 223–230. https://doi.org/10.1007/s00122-009-1031-4 (2009).
Ma, H. Q. et al. Identification and map** of a new powdery mildew resistance gene on chromosome 6D of common wheat. Theor. Appl. Genet. 123, 1099–1106. https://doi.org/10.1007/s00122-011-1651-3 (2011).
**ao, M. G. et al. Identification of the gene Pm47 on chromosome 7BS conferring resistance to powdery mildew in the Chinese wheat landrace Hongyanglazi. Theor. Appl. Genet. 126, 1397–1403. https://doi.org/10.1007/s00122-013-2060-6 (2013).
Mohler, V., Bauer, C., Schweizer, G., Kempf, H. & Hartl, L. Pm50: A new powdery mildew resistance gene in common wheat derived from cultivated emmer. J. Appl. Genet. 54, 259–263. https://doi.org/10.1007/s13353-013-0158-9 (2013).
Wu, P. P. et al. Fine map** of the wheat powdery mildew resistance gene Pm52 using comparative genomics analysis and the Chinese Spring reference genomic sequence. Theor. Appl. Genet. 132, 1451–1461. https://doi.org/10.1007/s00122-019-03291-7 (2019).
Zhang, R. Q. et al. Pm55, a developmental-stage and tissue-specific powdery mildew resistance gene introgressed from Dasypyrum villosum into common wheat. Theor. Appl. Genet. 129, 1975–1984. https://doi.org/10.1007/s00122-016-2753-8 (2016).
Hao, M. et al. Introgression of powdery mildew resistance gene Pm56 on rye chromosome arm 6RS into wheat. Front. Plant Sci. 9, 1040. https://doi.org/10.3389/fpls.2018.01040 (2018).
Xue, S. L. et al. Fine map** of Pm58 from Aegilops tauschii conferring powdery mildew resistance. Theor. Appl. Genet. 135, 1657–1669. https://doi.org/10.1007/s00122-022-04061-8 (2022).
Tan, C. C., Li, G. Q., Cowger, C., Carver, B. F. & Xu, X. Y. Characterization of Pm59, a novel powdery mildew resistance gene in Afghanistan wheat landrace PI 181356. Theor. Appl. Genet. 131, 1145–1152. https://doi.org/10.1007/s00122-018-3067-9 (2018).
Zou, S. H., Wang, H., Li, Y. W., Kong, Z. S. & Tang, D. Z. The NB-LRR gene Pm60 confers powdery mildew resistance in wheat. New Phytol. 218, 298–309. https://doi.org/10.1111/nph.14964 (2018).
Sun, H. et al. Pm61: a recessive gene for resistance to powdery mildew in wheat landrace Xuxusanyuehuang identified by comparative genomics analysis. Theor. Appl. Genet. 131, 2085–2097. https://doi.org/10.1007/s00122-018-3135-1 (2018).
Zhang, D. Y. et al. Wheat powdery mildew resistance gene Pm64 derived from wild emmer (Triticumturgidum var. dicoccoides) is tightly linked in repulsion with stripe rust resistance gene Yr5. Crop J. 7, 761–770. https://doi.org/10.1016/j.cj.2019.03.003 (2019).
Li, G. Q., Cowger, C., Wang, X. W., Carver, B. F. & Xu, X. Y. Characterization of Pm65, a new powdery mildew resistance gene on chromosome 2AL of a facultative wheat cultivar. Theor. Appl. Genet. 132, 2625–2632. https://doi.org/10.1007/s00122-019-03377-2 (2019).
Qiu, J. W. et al. Physical map** and identification of a candidate for the leaf rust resistance gene Lr1 of wheat. Theor. Appl. Genet. 115, 159–168. https://doi.org/10.1007/s00122-007-0551-z (2007).
Schachermayr, G. et al. Identification and localization of molecular markers linked to the Lr9 leaf rust resistance gene of wheat. Theor. Appl. Genet. 88, 110–115. https://doi.org/10.1007/bf00222402 (1994).
Schachermayr, G., Feuillet, C. & Keller, B. Molecular markers for the detection of the wheat leaf rust resistance gene Lr10 in diverse genetic backgrounds. Mol. Breed. 3, 65–74. https://doi.org/10.1023/A:1009619905909 (1997).
Gupta, S. K., Charpe, A., Koul, S., Prabhu, K. V. & Haq, Q. M. R. Development and validation of molecular markers linked to an Aegilops umbellulata-derived leaf-rust-resistance gene, Lr9, for marker-assisted selection in bread wheat. Genome 48, 823–830. https://doi.org/10.1139/g05-051 (2005).
Neu, C., Stein, N. & Keller, B. Genetic map** of the Lr20-Pm1 resistance locus reveals suppressed recombination on chromosome arm 7AL in hexaploid wheat. Genome 45, 737–744. https://doi.org/10.1139/g02-040 (2022).
Schachermayr, G. M. et al. Identification of molecular markers linked to the Agropyron elongatum-derived leaf rust resistance gene Lr24 in wheat. Theor. Appl. Genet. 90, 982–990. https://doi.org/10.1007/bf00222911 (1995).
Chai, J. F., Zhou, R. H., Jia, J. Z. & Liu, X. Development and application of a new co-dominant PCR marker for detecting 1BL·1RS wheat-rye chromosome translocations. Plant Breed. 125, 302–304. https://doi.org/10.1111/j.1439-0523.2006.01186.x (2006).
Helguera, M. et al. PCR assays for the Lr37-Yr17-Sr38 cluster of rust resistance genes and their use to develop isogenic hard red spring wheat lines. Crop Sci. 43, 1839–1847. https://doi.org/10.2135/cropsci2003.1839 (2003).
Liu, Y. et al. Analysis on the rust resistance genes of a new wheat germplasm YW243 by molecular markers. Sci. Agric. Sin. 2, 295–299. https://doi.org/10.3864/j.issn.0578-1752.at-2005-5891 (2006).
Chen, X. M., Soria, M. A., Yan, G. P., Sun, J. & Dubcovsky, J. Development of sequence tagged site and cleaved amplified polymorphic sequence markers for wheat stripe rust resistance gene Yr5. Crop Sci. 43, 2058–2064. https://doi.org/10.2135/cropsci2003.2058 (2003).
Liu, C. et al. Isolation of a new repetitive DNA sequence from Secale africanum enables targeting of Secale chromatin in wheat background. Euphytica 159, 249–258. https://doi.org/10.1007/s10681-007-9484-5 (2008).
Shao, Y. T. et al. Identification of an AFLP marker linked to the stripe rust resistance gene Yr10 in wheat. Chin. Sci. Bull. 46, 1466–1468. https://doi.org/10.1007/BF03187033 (2001).
Klymiuk, V. et al. Cloning of the wheat Yr15 resistance gene sheds light on the plant tandem kinase-pseudokinase family. Nat. Commun. 9, 3735. https://doi.org/10.1038/s41467-018-06138-9 (2018).
Jia, J. Q., Lei, M. P., Liu, C., Li, G. R. & Yang, Z. J. Exploitation and application of a new SCAR marker linked to strip rust resistance gene Yr17 in wheat. J. Triticeae Crops 30, 11–16. https://doi.org/10.7606/j.issn.1009-1041.2010.01.003 (2010).
McIntosh, R. et al. Wheat stripe rust resistance gene Yr24/Yr26: A retrospective review. Crop J. 6, 321–329. https://doi.org/10.1016/j.cj.2018.02.001 (2018).
Wang, C. M. et al. SSR and STS markers for wheat stripe rust resistance gene Yr26. Euphytica 159, 359–366. https://doi.org/10.1007/s10681-007-9524-1 (2008).
Hayden, M. J., Kuchel, H. & Chalmers, K. J. Sequence tagged microsatellites for the Xgwm533 locus provide new diagnostic markers to select for the presence of stem rust resistance gene Sr2 in bread wheat (Triticum aestivum L.). Theor. Appl. Genet. 109, 1641–1647. https://doi.org/10.1007/s00122-004-1787-5 (2004).
Luo, P. G. et al. Allelic analysis of stripe rust resistance genes on wheat chromosome 2BS. Genome 51, 922–927. https://doi.org/10.1139/g08-079 (2008).
Xu, H. X. et al. Development and validation of molecular markers closely linked to the wheat stripe rust resistance gene YrC591 for marker-assisted selection. Euphytica 198, 317–323. https://doi.org/10.1007/s10681-014-1108-2 (2014).
Feng, J. Y. et al. Molecular map** of YrSP and its relationship with other genes for stripe rust resistance in wheat chromosome 2BL. Phytopathology 105, 1206–1213. https://doi.org/10.1094/phyto-03-15-0060-r (2015).
Korzun, V., Röder, M. S., Ganal, M. W., Worland, A. J. & Law, C. N. Genetic analysis of the dwarfing gene (Rht8) in wheat. Part I. Molecular map** of Rht8 on the short arm of chromosome 2D of bread wheat (Triticum aestivum L.). Theor. Appl. Genet. 96, 1104–1109. https://doi.org/10.1007/s001220050845 (1998).
Wang, J. P., Li, R. Z., Mao, X. G. & **g, R. L. Functional analysis and marker development of TaCRT-D gene in common wheat (Triticum aestivum L.). Front. Plant Sci. 8, 1557. https://doi.org/10.3389/fpls.2017.01557 (2017).
Feng, Y. M. et al. A 4-bp deletion in the 5’UTR of TaAFP-B is associated with seed dormancy in common wheat (Triticum aestivum L.). BMC Plant Biol. 19, 349. https://doi.org/10.1186/s12870-019-1950-4 (2019).
Liu, S. B., Cai, S. B., Grayboossch, R., Chen, C. X. & Bai, G. H. Quantitative trait loci for resistance to pre-harvest sprouting in US hard white winter wheat Rio Blanco. Theor. Appl. Genet. 117, 691–699. https://doi.org/10.1007/s00122-008-0810-7 (2008).
Lei, Z. S. et al. Y-type gene specific markers for enhanced discrimination of high-molecular weight glutenin alleles at the Glu-B1 locus in hexaploid wheat. J. Cereal Sci. 43, 94–101. https://doi.org/10.1016/j.jcs.2005.08.003 (2006).
Xu, Q. et al. PCR-based markers for identification of HWM-GS at Glu-B1x loci in common wheat. J. Cereal Sci. 47, 394–398. https://doi.org/10.1016/j.jcs.2007.05.002 (2008).
Ma, W., Zhang, W. & Gale, K. R. Multiplex-PCR ty** of high molecular weight glutenin alleles in wheat. Euphytica 134, 51–60. https://doi.org/10.1023/A:1026191918704 (2003).
Smith, R. L., Schweder, M. E. & Barnett, R. D. Identification of glutenin alleles in wheat and triticale using PCR-generated DNA markers. Crop Sci. 34, 1373–1378. https://doi.org/10.2135/cropsci1994.0011183X003400050042x (1994).
Wang, L. H., Li, G. Y., Peña, R. J., **a, X. C. & He, Z. H. Development of STS markers and establishment of multiplex PCR for Glu-A3 alleles in common wheat (Triticum aestivum L.). J. Cereal Sci. 51, 305–312. https://doi.org/10.1016/j.jcs.2010.01.005 (2010).
Wang, L. H. et al. Characterization of low-molecular-weight glutenin subunit Glu-B3 genes and development of STS markers in common wheat (Triticum aestivum L.). Theor. Appl. Genet. 118, 525–539. https://doi.org/10.1007/s00122-008-0918-9 (2009).
Acknowledgements
We are grateful to Prof. Yilin Zhou, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Bei**g, China, Prof. Hongxing Xu, Henan University, Kaifeng, China and Prof. Caixia Lan, Huazhong Agricultural University, Wuhan, China, for providing the donors with the known Pm and Yr genes and/or Blumeria graminis f. sp. tritici isolates.
Funding
This research was financially supported by the National Natural Science Foundation of China (32072053), the Key Research and Development Project of Shandong Province (2020CXGC010805), Science and Technology Innovation Project of Hebei Academy of Agricultural and Forestry Sciences (2024KJCXZX-LYS-15), Key Research and Development Project of Yantai City (2022XCZX092), Key R&D Plan of Shandong Province (2022LZG002-4) and Wheat Industry Technology System of Shandong Province (SDAIT-01-01).
Author information
Authors and Affiliations
Contributions
Conceptualization, P.M., F.W., and C.L.; Methodology, P.M., B.X., Y.J., N.Y., and X.W.; Validation, B.X., J.W., Z.Q., Y.Z., and Y.Q.; Formal analysis, N.S., L.L., T.Y., and W.L.; Investigation, P.M., Y.J. and C.L.; Resources, P.M., Y.J., and C.L.; Writing–original draft preparation, Y.J., and P.M.; Writing–review and editing, P.M. and C.L.; Visualization, P.M., F.W., and C.L.; Funding acquisition, P.M., and C.L. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
**ao, B., Qie, Y., **, Y. et al. Genetic basis of an elite wheat cultivar Guinong 29 with harmonious improvement between multiple diseases resistance and other comprehensive traits. Sci Rep 14, 14336 (2024). https://doi.org/10.1038/s41598-024-64998-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-64998-2
- Springer Nature Limited